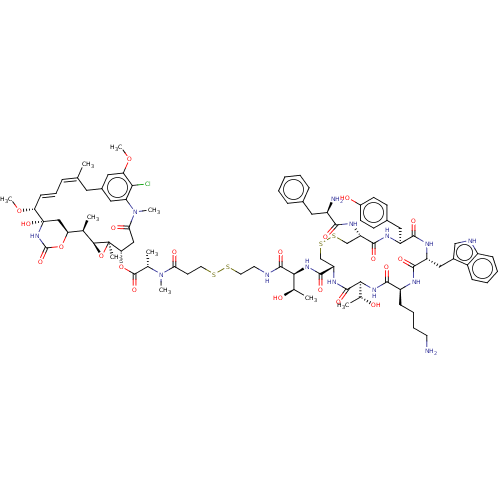
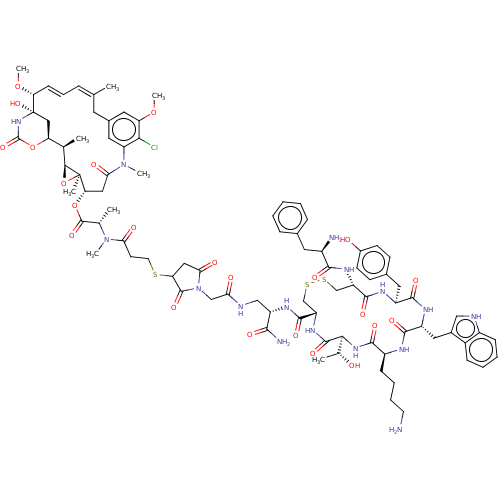
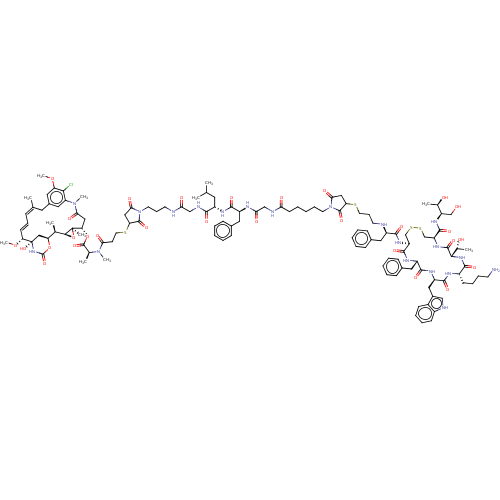
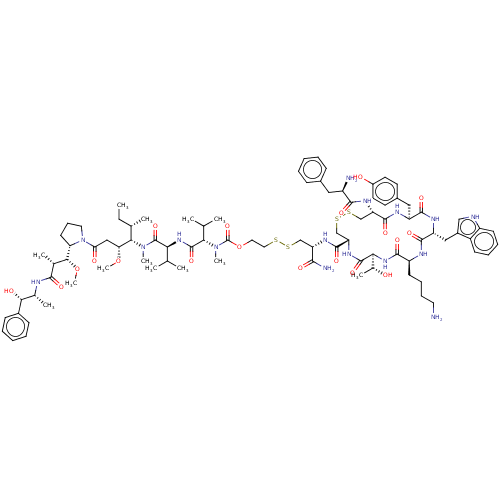
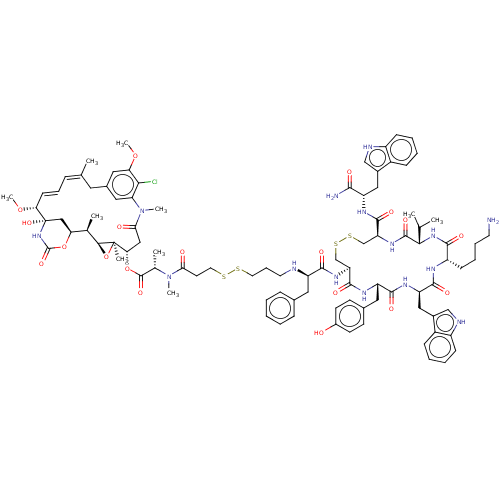
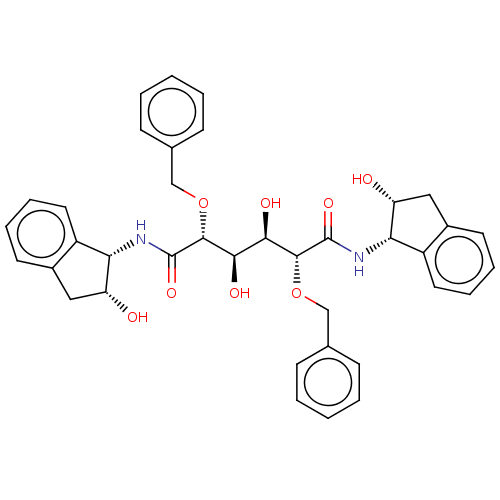
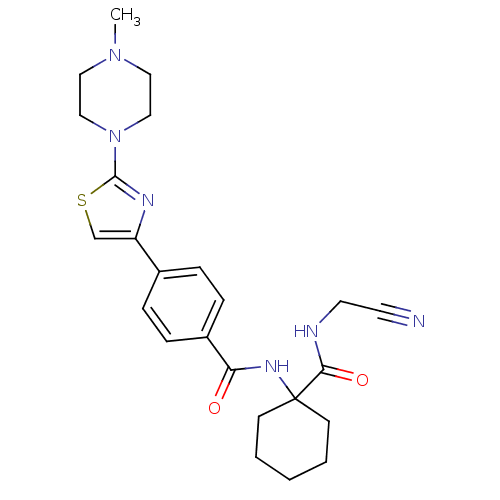
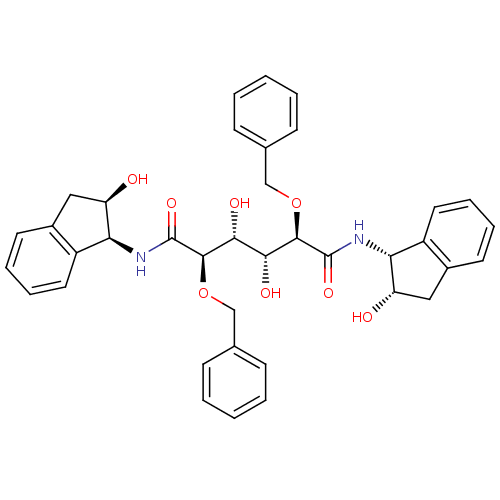
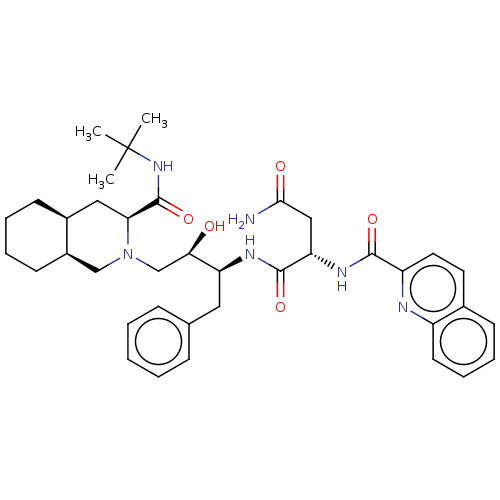
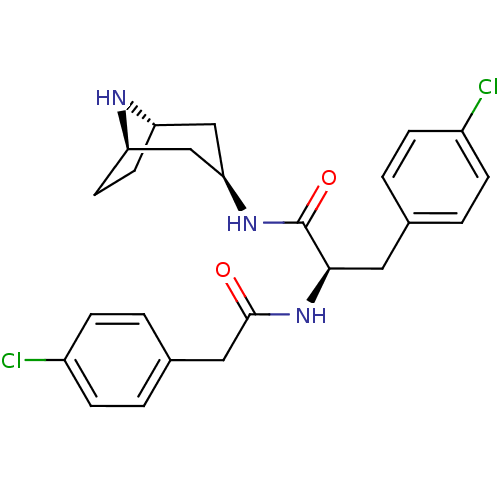
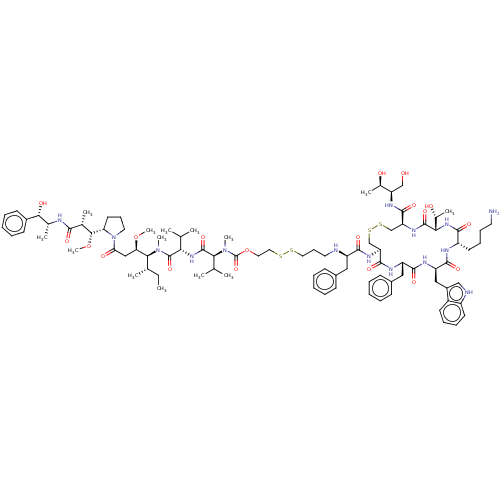
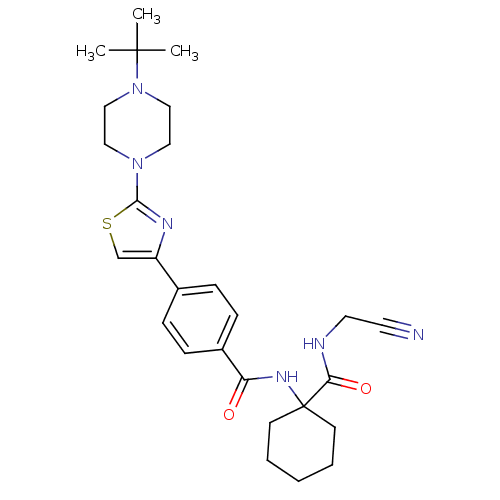
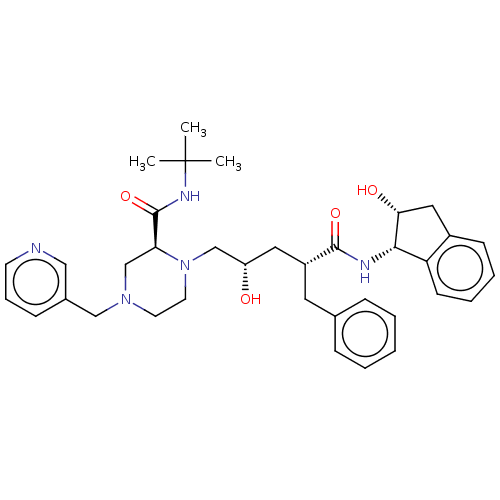
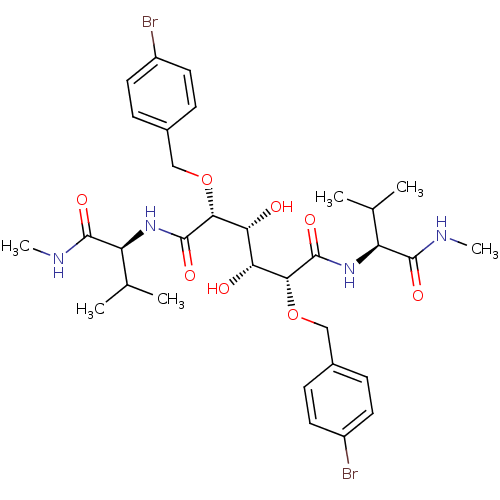
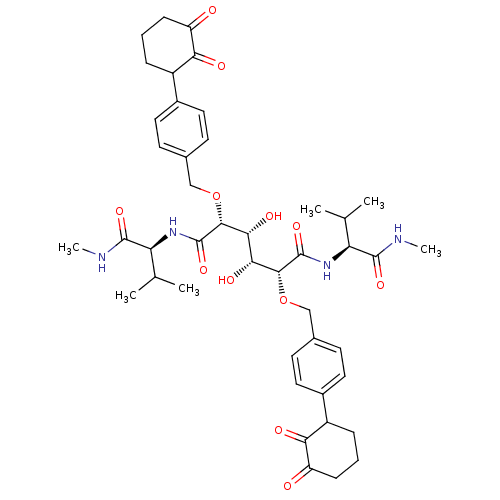
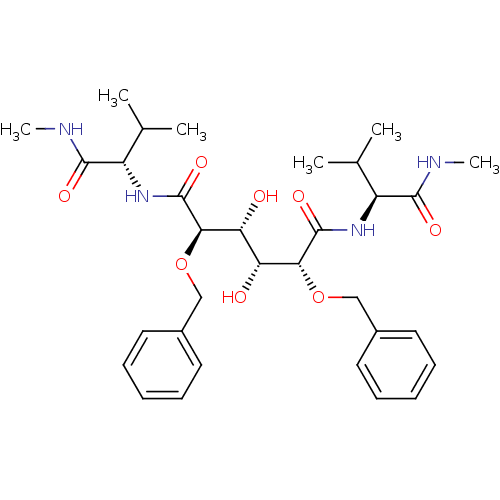
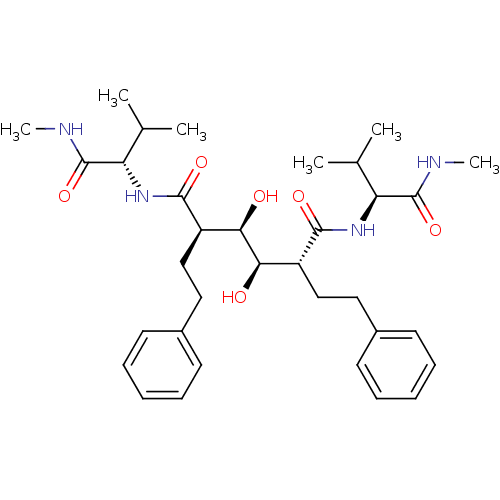
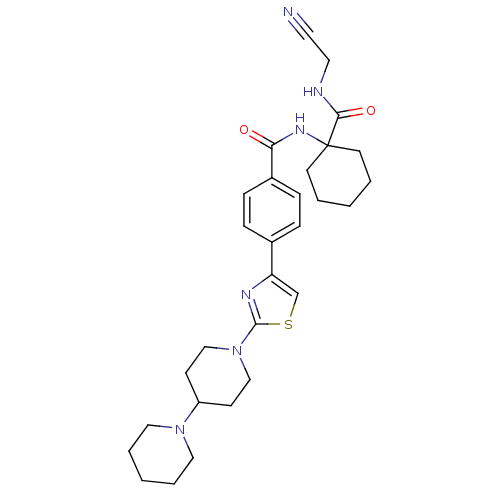
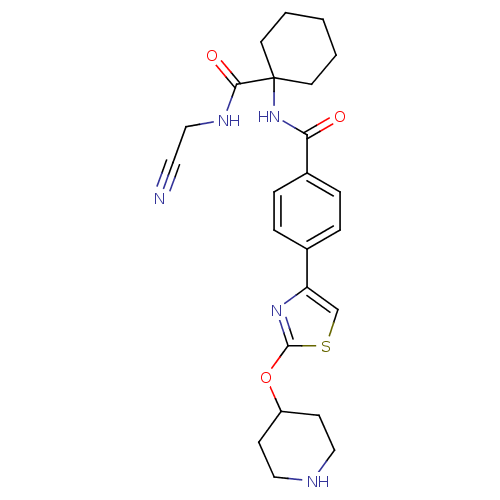
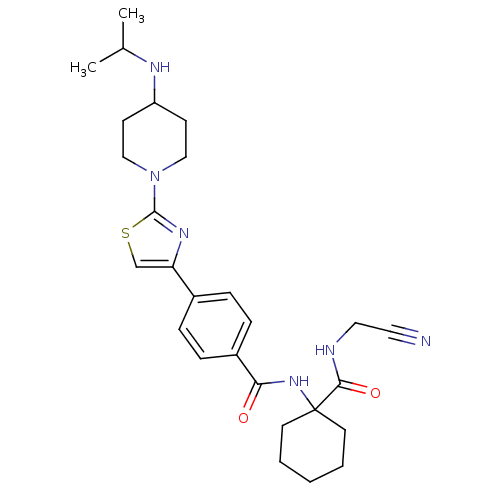
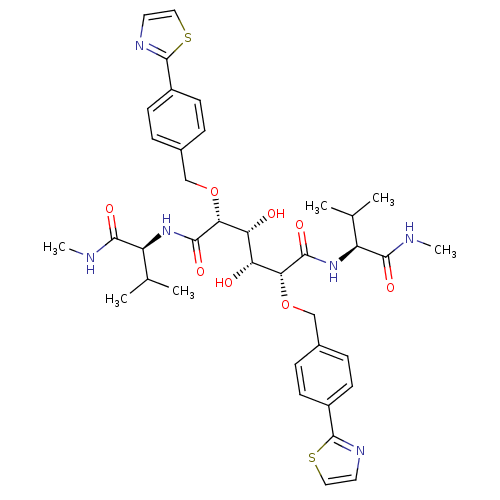
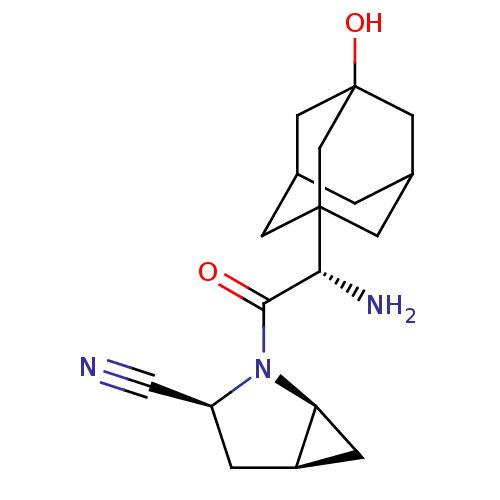
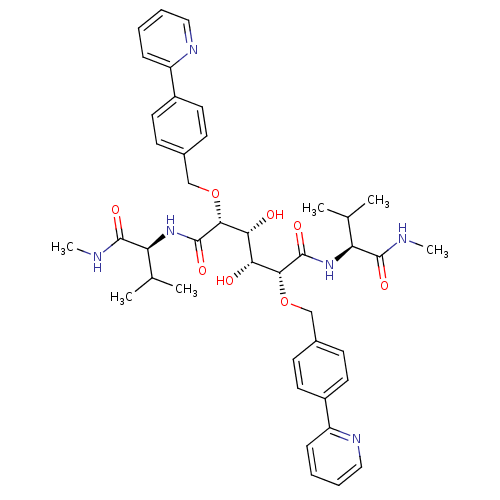
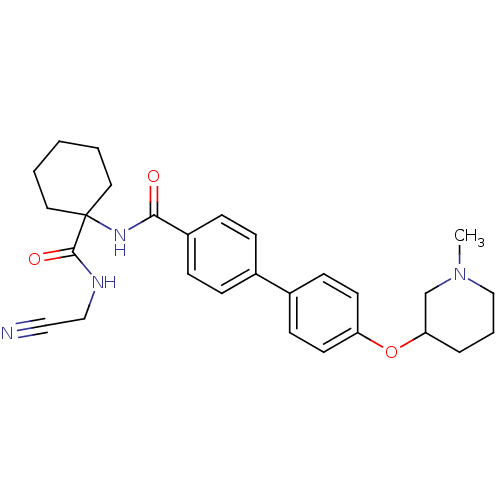
Affinity DataKi: 0.00500nMAssay Description:Displacement of [125I]somatostatin from human SSTR2 expressed in CHO-K1 cell membranes after 240 minsMore data for this Ligand-Target Pair
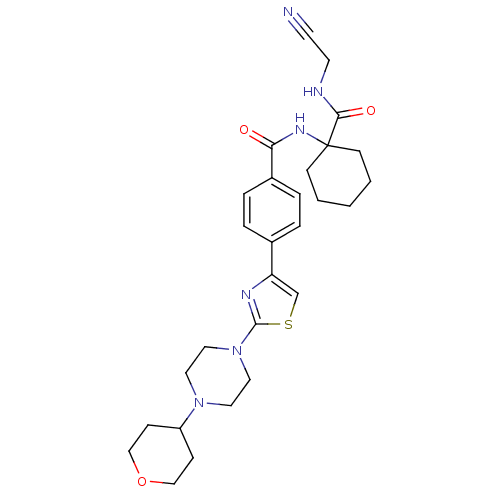
Affinity DataKi: 0.0130nMAssay Description:Displacement of [125I]somatostatin from human SSTR2 expressed in CHO-K1 cell membranes after 240 minsMore data for this Ligand-Target Pair
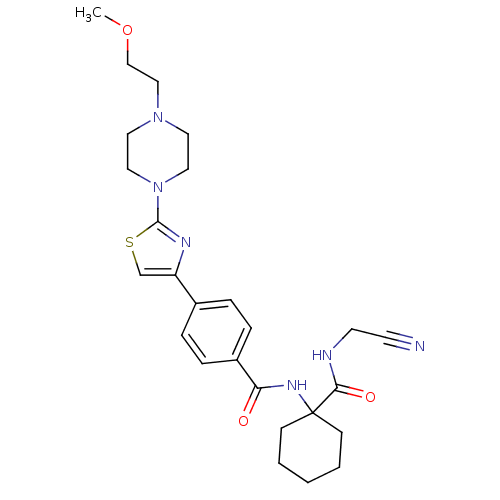
Affinity DataKi: 0.0150nMAssay Description:Displacement of [125I]somatostatin from human SSTR2 expressed in CHO-K1 cell membranes after 240 minsMore data for this Ligand-Target Pair
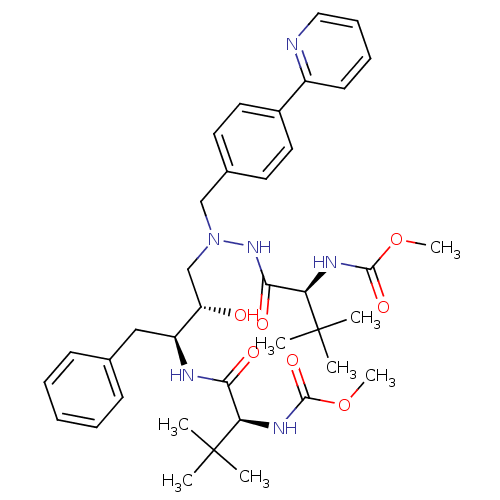
Affinity DataKi: 0.0150nMAssay Description:Displacement of [125I]somatostatin from human SSTR2 expressed in CHO-K1 cell membranes after 240 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.0150nMAssay Description:Displacement of [125I]somatostatin from human SSTR2 expressed in CHO-K1 cell membranes after 240 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.0180nMAssay Description:Displacement of [125I]somatostatin from human SSTR2 expressed in CHO-K1 cell membranes after 240 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.0220nMAssay Description:Displacement of [125I]somatostatin from human SSTR2 expressed in CHO-K1 cell membranes after 240 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.0610nMAssay Description:Displacement of [125I]somatostatin from human SSTR2 expressed in CHO-K1 cell membranes after 240 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.0750nMAssay Description:Displacement of [125I]somatostatin from human SSTR2 expressed in CHO-K1 cell membranes after 240 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.0900nM ΔG°: -58.3kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nMAssay Description:Displacement of [125I]somatostatin from human SSTR2 expressed in CHO-K1 cell membranes after 240 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.100nM ΔG°: -58.0kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 0.100nM ΔG°: -58.0kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 0.120nMAssay Description:Displacement of [125I]somatostatin from human SSTR2 expressed in CHO-K1 cell membranes after 240 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.120nMAssay Description:Displacement of [125I]somatostatin from human SSTR2 expressed in CHO-K1 cell membranes after 240 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.140nMAssay Description:Displacement of [125I]somatostatin from human SSTR2 expressed in CHO-K1 cell membranes after 240 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.140nMAssay Description:Displacement of [125I]somatostatin from human SSTR2 expressed in CHO-K1 cell membranes after 240 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.180nMAssay Description:Displacement of [125I]somatostatin from human SSTR2 expressed in CHO-K1 cell membranes after 240 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.190nMAssay Description:Displacement of [125I]somatostatin from human SSTR2 expressed in CHO-K1 cell membranes after 240 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.200nM ΔG°: -56.3kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
LinköPing University
Curated by ChEMBL
LinköPing University
Curated by ChEMBL
Affinity DataKi: 0.200nMAssay Description:Inhibitory activity against purified HIV-1 protease expressed in E. coli in sensitive fluorometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli by fluorometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.234nMAssay Description:Displacement of [3H]AVP from human V1A receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.25nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.260nMAssay Description:Displacement of [125I]somatostatin from human SSTR2 expressed in CHO-K1 cell membranes after 240 minsMore data for this Ligand-Target Pair
Affinity DataKi: 0.290nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli by fluorometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.300nM ΔG°: -55.3kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nM ΔG°: -55.3kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair
Affinity DataKi: 0.300nM ΔG°: -55.3kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair
Affinity DataKi: 0.400nM ΔG°: -54.5kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair
Affinity DataKi: 0.400nM ΔG°: -54.5kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
TargetGag-Pol polyprotein [489-587](Human immunodeficiency virus type 1)
LinköPing University
Curated by ChEMBL
LinköPing University
Curated by ChEMBL
Affinity DataKi: 0.400nMAssay Description:Inhibitory activity against purified HIV-1 protease expressed in E. coli in sensitive fluorometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.400nM ΔG°: -54.5kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 0.450nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.470nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.480nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coli by fluorometric assayMore data for this Ligand-Target Pair
Affinity DataKi: 0.520nMAssay Description:Inhibition of HIV1 protease expressed in Escherichia coliMore data for this Ligand-Target Pair
Affinity DataKi: 0.562nMAssay Description:Displacement of [3H]AVP from human V1A receptor expressed in CHO cellsMore data for this Ligand-Target Pair
Affinity DataKi: 0.570nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.590nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.600nM ΔG°: -53.5kJ/molepH: 5.0 T: 2°CAssay Description:A fluorimetric assay was used to determine the effects of the inhibitors on HIV-1 protease. This assay used an internally quenched fluorescent peptid...More data for this Ligand-Target Pair
Affinity DataKi: 0.600nM ΔG°: -53.5kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair
TargetDipeptidyl peptidase 4(Homo sapiens (Human))
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Bristol-Myers Squibb Research And Development
Curated by ChEMBL
Affinity DataKi: 0.600nMAssay Description:Inhibition of human DPP4More data for this Ligand-Target Pair
Affinity DataKi: 0.600nM ΔG°: -53.5kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair
Affinity DataKi: 0.670nMAssay Description:Inhibitory constant against rabbit cathepsin K using Z-Phe-Arg-AMC substrateMore data for this Ligand-Target Pair
Affinity DataKi: 0.700nM ΔG°: -53.1kJ/molepH: 5.0 T: 2°CAssay Description:Ki values were determined by using a fluorescent substrate (DABCYL-gamma-Abu-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-EDANS). All incubations were performed a...More data for this Ligand-Target Pair

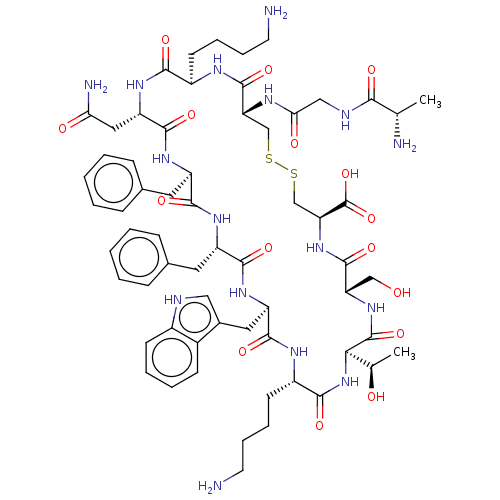
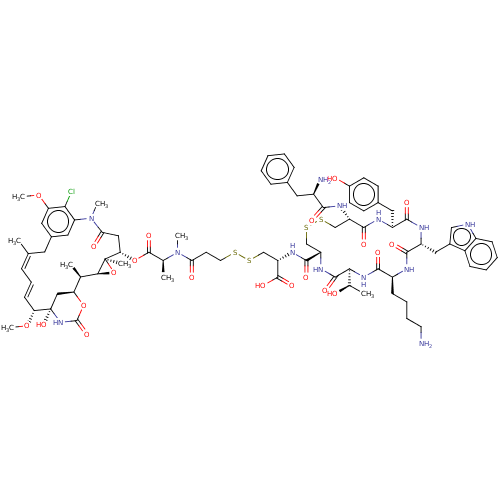
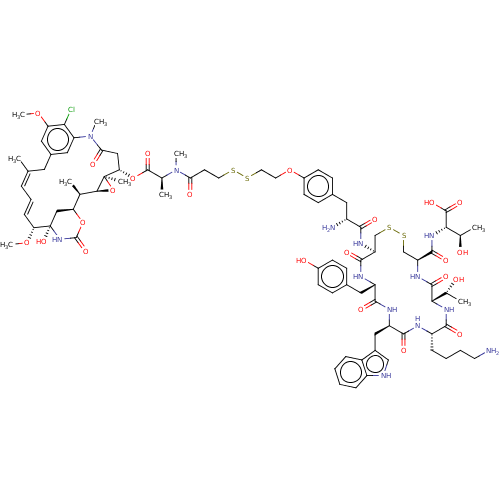
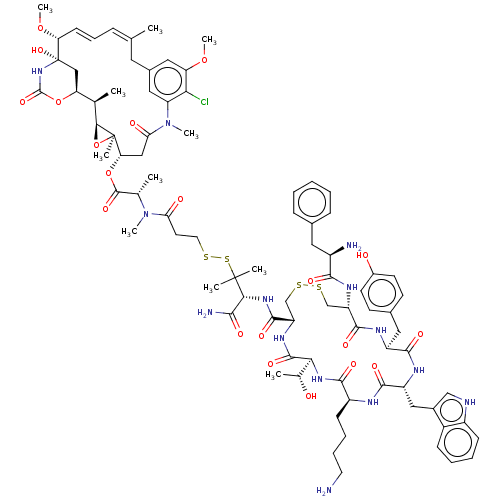
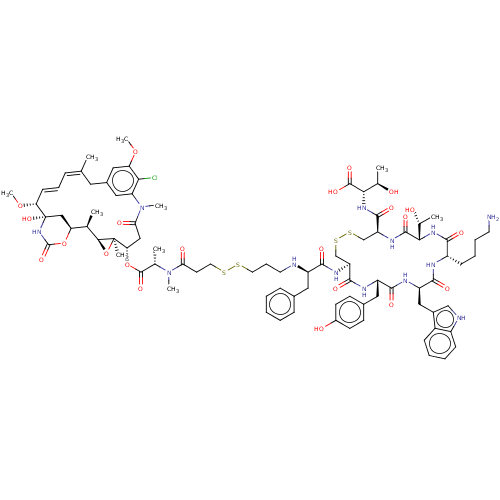
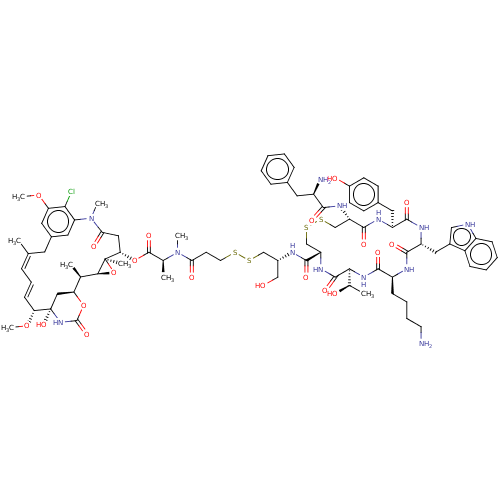
 3D Structure (crystal)
3D Structure (crystal)