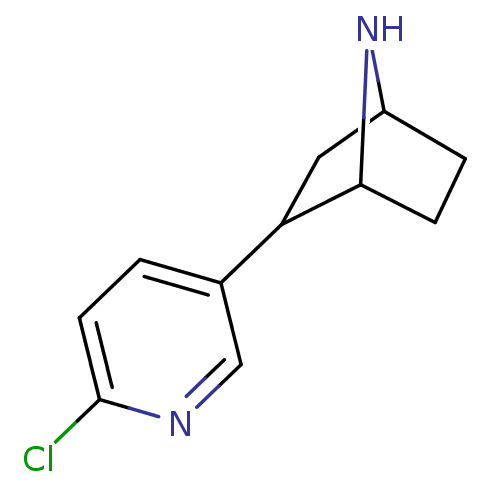
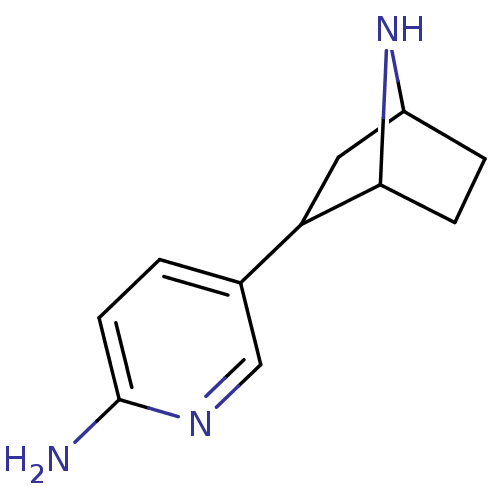
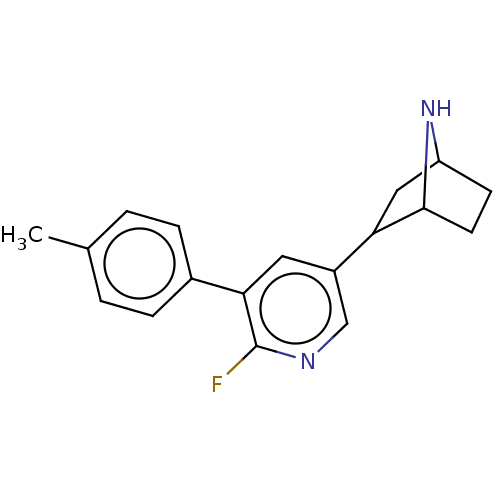
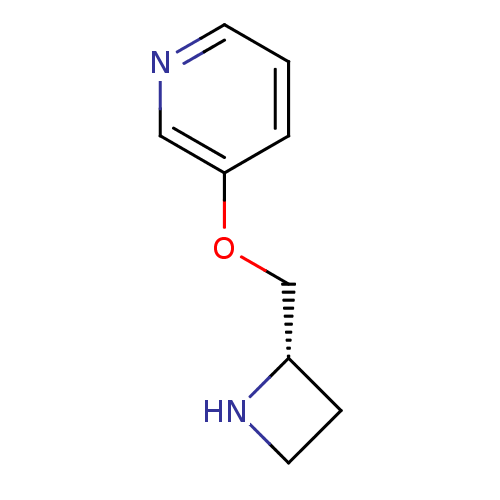
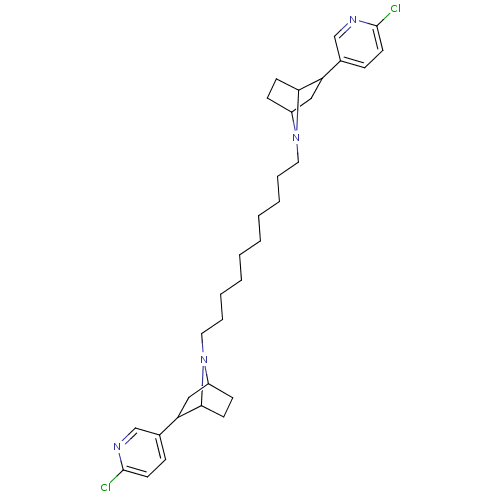
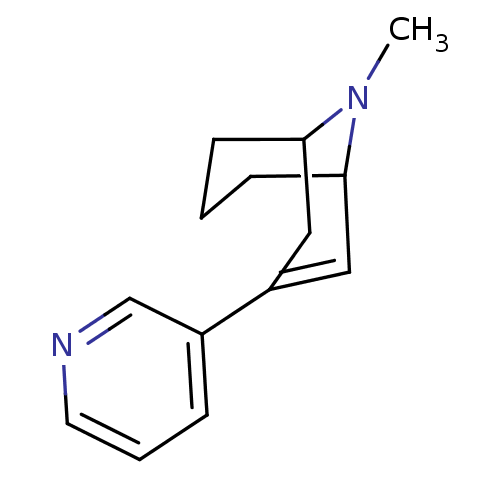
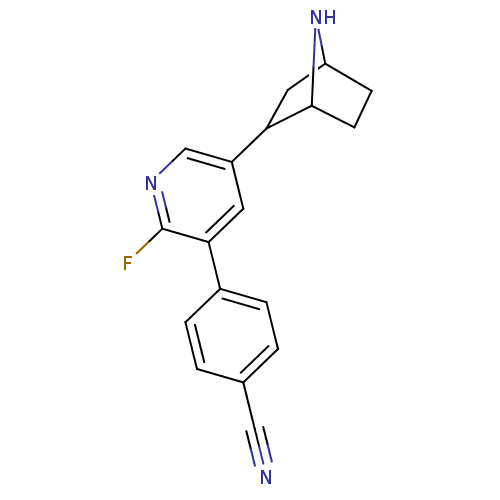
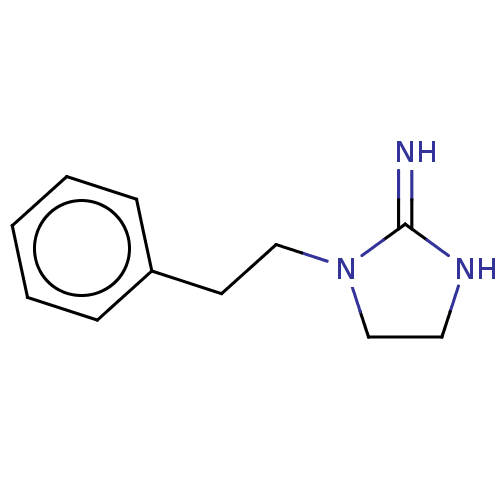
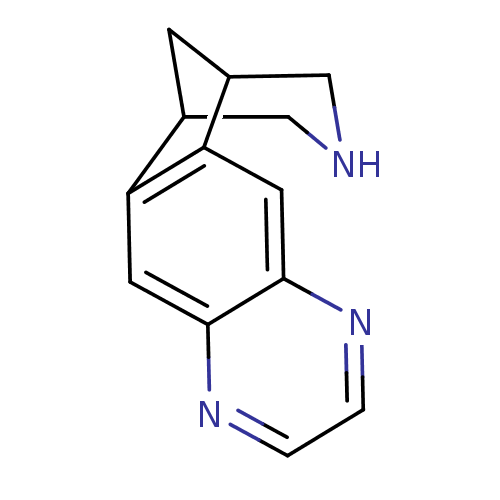
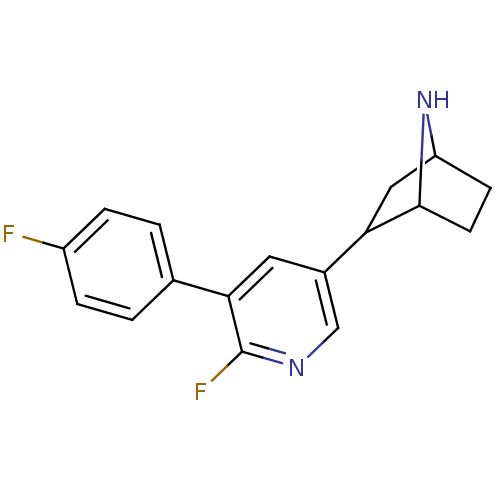
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 0.150nMAssay Description:Displacement of [3H]epibatidine from rat alpha3beta4 nAChR expressed in HEK292 cells after 3 hrsMore data for this Ligand-Target Pair
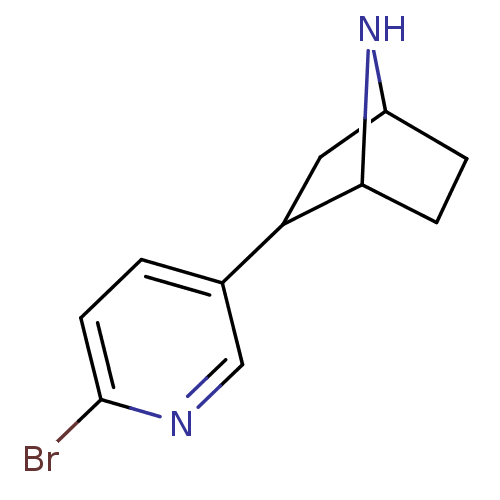
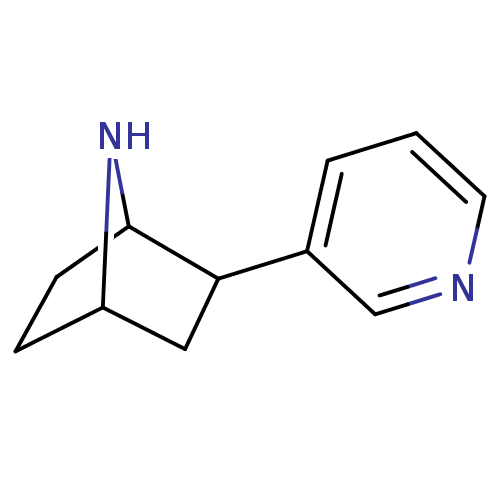
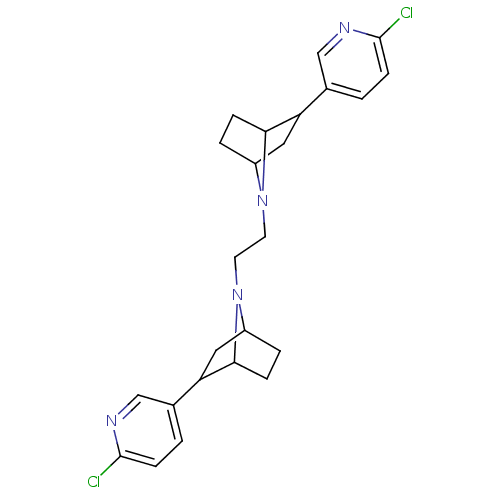
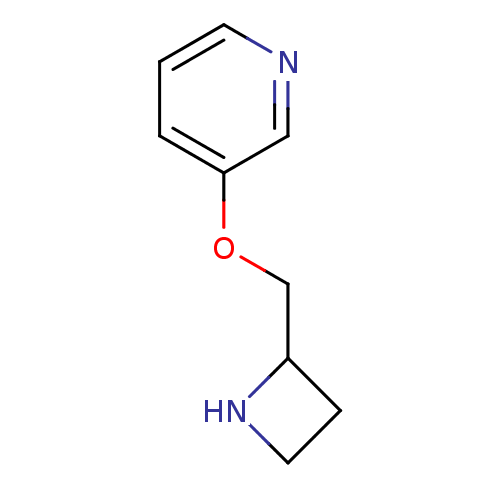
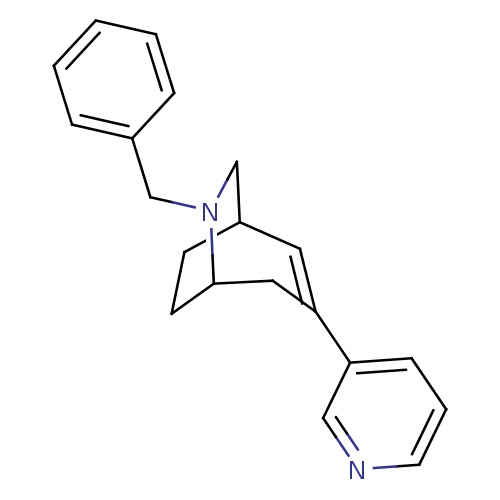
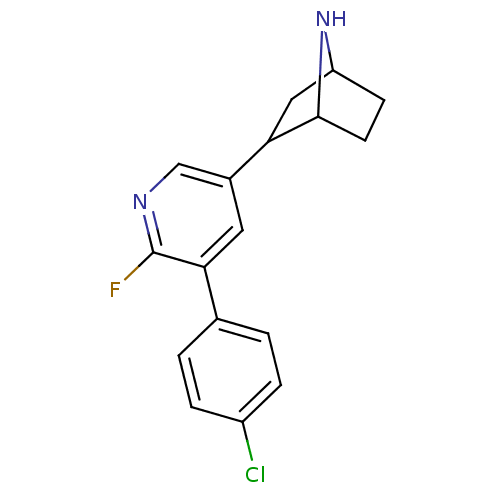
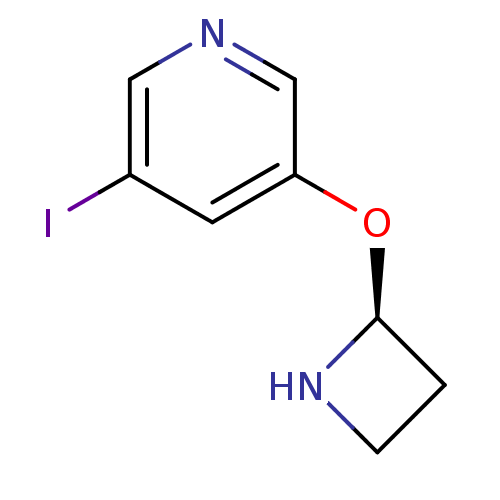
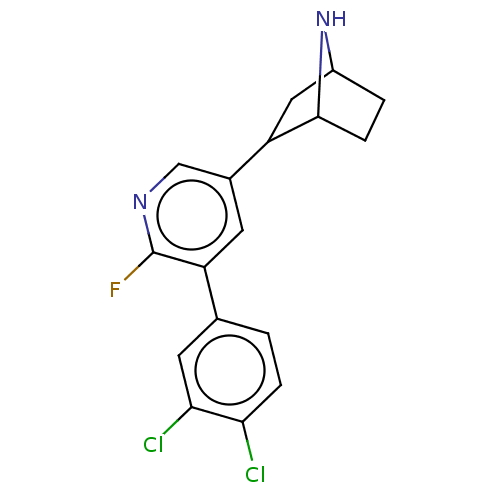
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 0.220nMAssay Description:Inhibition of [3H]-nicotine binding to alpha4-beta2 nACh receptor from rat membranes from ref 18More data for this Ligand-Target Pair
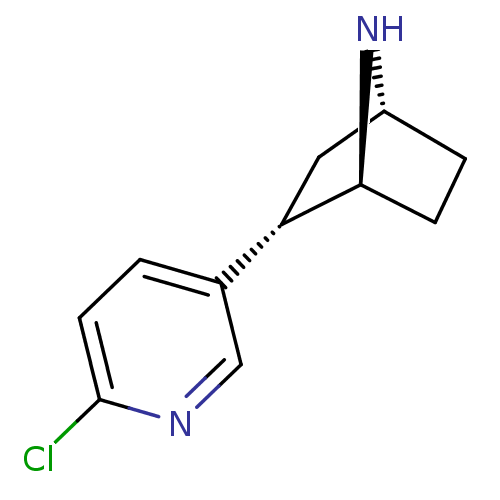
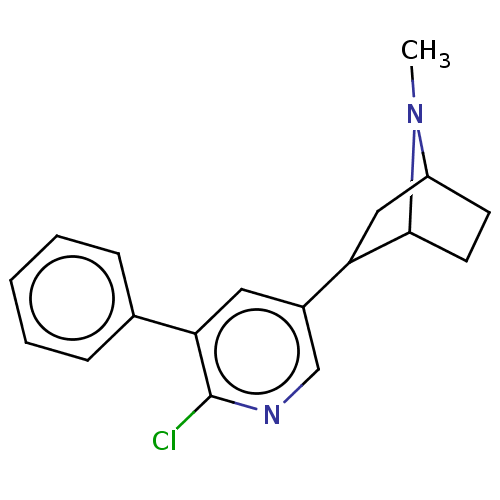
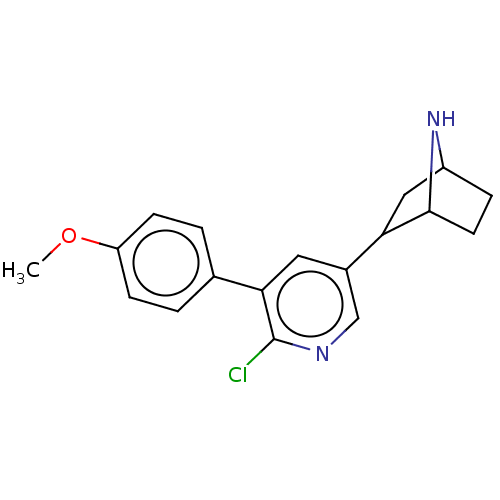
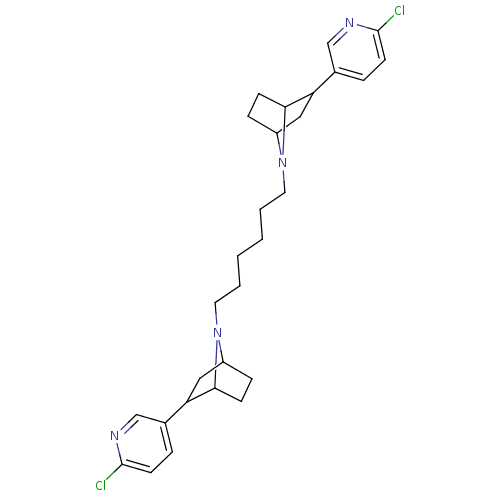
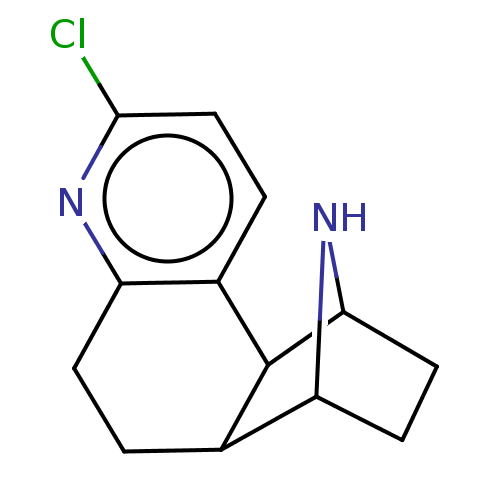
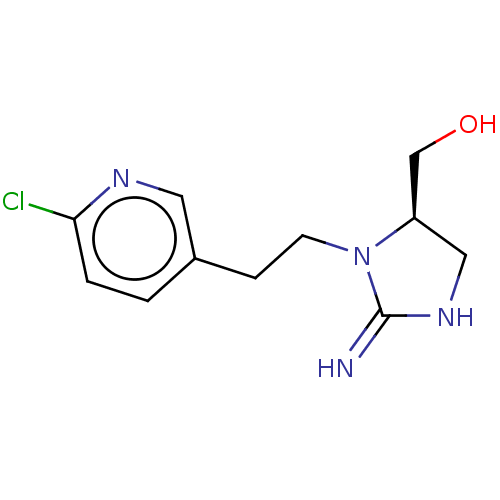
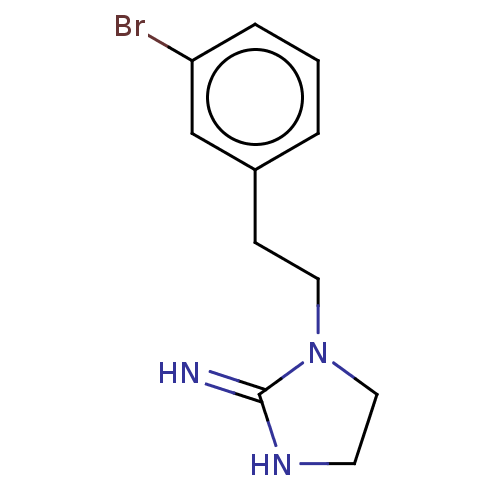
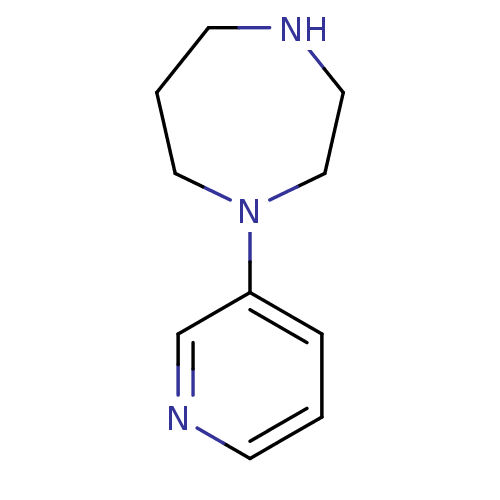
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 0.300nMAssay Description:Binding affinity towards rat Nicotinic acetylcholine receptor alpha4-beta2 expressed in Xenopus oocytes using [3H]-epibatidine as radioligandMore data for this Ligand-Target Pair
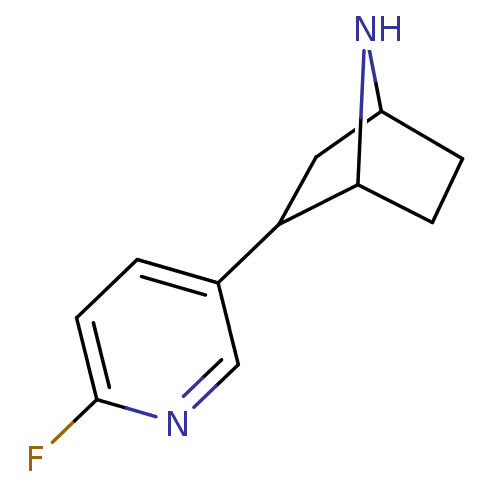
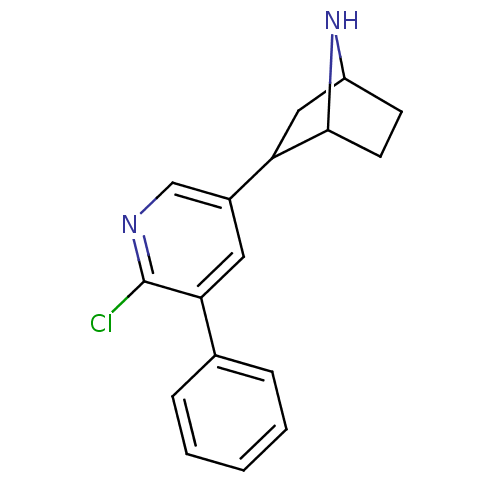
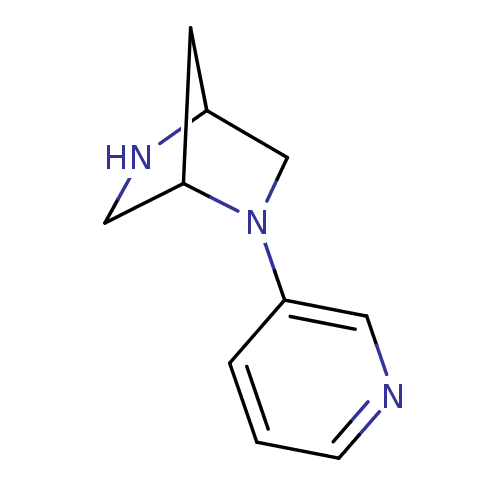
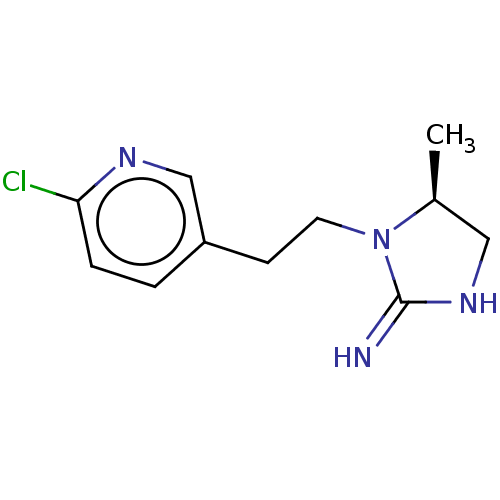
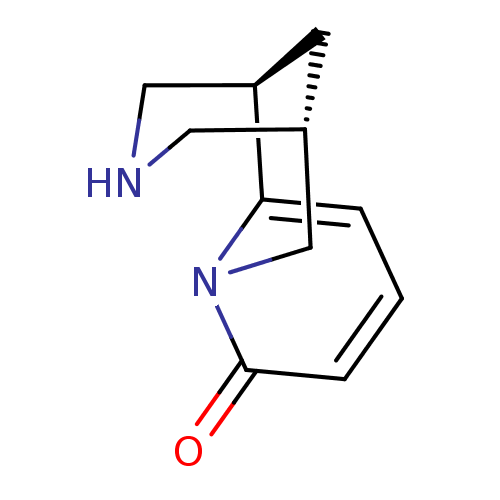
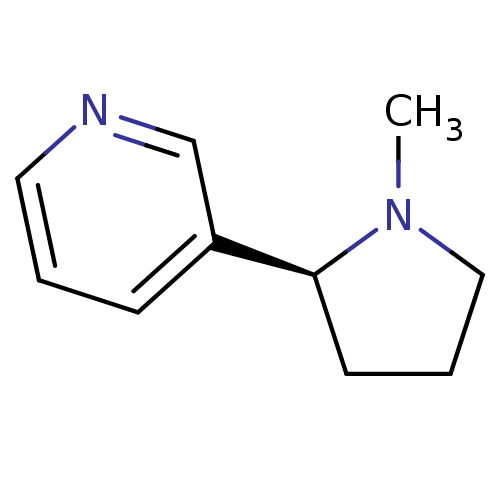
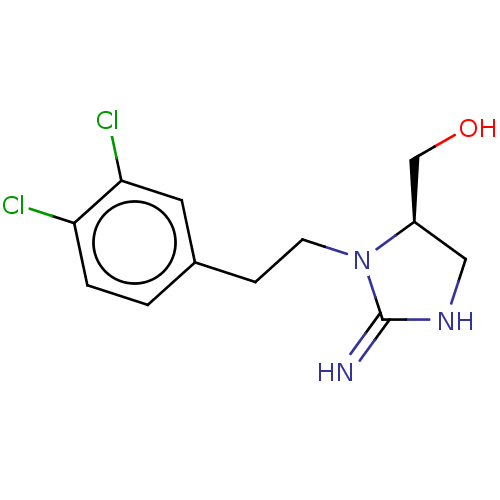
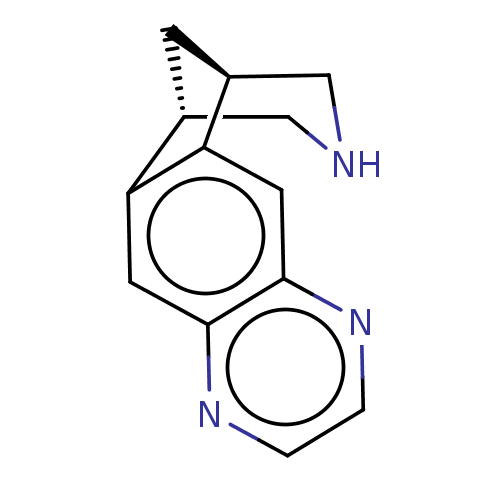
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 0.380nMAssay Description:Displacement of [3H]epibatidine from Alpha3 Beta4 Nicotinic acetylcholine receptor of rat brain homogenatesMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 0.380nMAssay Description:Displacement of [3H]epibatidine from Alpha3 Beta4 Nicotinic acetylcholine receptor of rat brain homogenatesMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 0.480nMAssay Description:Binding affinity towards rat Nicotinic acetylcholine receptor alpha4-beta4 expressed in HEK293 cells using [3H]EB as radioligandMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 0.480nMAssay Description:Binding affinity towards rat Nicotinic acetylcholine receptor alpha3-beta2 expressed in HEK293 cells using [3H]EB as radioligandMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 0.565nMAssay Description:Binding affinity towards Nicotinic acetylcholine receptor alpha4-beta4 using [3H]epibatidineMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 0.565nMAssay Description:Tested for binding affinity against Nicotinic acetylcholine receptor alpha3-beta4More data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 0.570nMAssay Description:Inhibition of [3H]epibatidine binding to rat Nicotinic acetylcholine receptor alpha3-beta4More data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 0.570nMAssay Description:In vitro binding affinity towards rat Nicotinic acetylcholine receptor alpha3-beta4 using 0.5 nM [3H]epibatidineMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 0.570nMAssay Description:Displacement of [3H]epibatidine from rat alpha3beta4 nACHR expressed in human HEK293 cellsMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 2.10nMAssay Description:Binding affinity towards rat Nicotinic acetylcholine receptor alpha3-beta4 expressed in Xenopus oocytes using [3H]-epibatidine as radioligandMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 7.20nMAssay Description:Inhibition of [3H]-epibatidine binding to Nicotinic acetylcholine receptor alpha4-beta2 from rat membranesMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 11nMAssay Description:Inhibition of [3H]-nicotine binding to Nicotinic acetylcholine receptor alpha4-beta2 from rat membranes from ref 15More data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 15nMAssay Description:In vitro binding affinity towards rat Nicotinic acetylcholine receptor alpha3-beta4 using 0.5 nM [3H]epibatidineMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 24nMAssay Description:In vitro binding affinity towards rat Nicotinic acetylcholine receptor alpha3-beta4 using 0.5 nM [3H]epibatidineMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 29nMAssay Description:In vitro binding affinity towards rat Nicotinic acetylcholine receptor alpha3-beta4 using 0.5 nM [3H]epibatidineMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 48nMAssay Description:Binding affinity towards nicotinic acetylcholine receptor alpha2-beta2 using [3H]epibatidineMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 55nMAssay Description:In vitro binding affinity towards rat Nicotinic acetylcholine receptor alpha3-beta4 using 0.5 nM [3H]epibatidineMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 65nMAssay Description:Displacement of [3H]epibatidine from rat alpha3beta4 nAChR by liquid scintillation countingMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 72nMAssay Description:Binding affinity towards rat Nicotinic acetylcholine receptor alpha3-beta2 expressed in HEK293 cells using [3H]EB as radioligandMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 74nMAssay Description:Displacement of [3H]epibatidine from Alpha3 Beta4 Nicotinic acetylcholine receptor of rat brain homogenatesMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 82nMAssay Description:Binding affinity towards nicotinic acetylcholine receptor alpha2-beta2 using [3H]epibatidineMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 95nMAssay Description:Displacement of [3H]epibatidine from rat alpha3beta4 nACHR expressed in human HEK293 cellsMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 112nMAssay Description:Binding affinity towards Nicotinic acetylcholine receptor alpha4-beta4 using [3H]epibatidineMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 125nMAssay Description:Displacement of [3H]epibatidine from rat alpha3beta4 nAChR expressed in HEK cell membranes after 2 hrsMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 138nMAssay Description:Binding affinity towards Nicotinic acetylcholine receptor alpha4-beta2 using [3H]epibatidineMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 170nMAssay Description:Displacement of [3H]epibatidine from rat alpha3beta4 nAChR expressed in HEK292 cells after 3 hrsMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 201nMAssay Description:Tested for binding affinity against Nicotinic acetylcholine receptor alpha3-beta4More data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 203nMAssay Description:Displacement of [3H]epibatidine from rat alpha3beta4 nAChR expressed in HEK292 cells after 3 hrsMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 210nMAssay Description:Binding affinity towards rat Nicotinic acetylcholine receptor alpha4-beta2 expressed in HEK293 cells using [3H]EB as radioligandMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 217nMAssay Description:Displacement of [3H]epibatidine from rat alpha3beta4 nACHR expressed in human HEK293 cellsMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 241nMAssay Description:Displacement of [3H]epibatidine from rat alpha3beta4 nAChR expressed in HEK292 cells after 3 hrsMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 256nMAssay Description:In vitro binding affinity towards rat Nicotinic acetylcholine receptor alpha3-beta4 using 0.5 nM [3H]epibatidineMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 270nMAssay Description:Displacement of [3H]epibatadine from rat alpha3beta4 nAChR expressed in HEK293 cells after 2 hrs by betaplate counting analysisMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 281nMAssay Description:Displacement of [3H]epibatidine from rat alpha3beta4 nAChR expressed in HEK cell membranes after 2 hrsMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 288nMAssay Description:Displacement of [3H]epibatidine from rat neuronal acetylcholine receptor subunit alpha3beta4 expressed in HEK293 cell membranes incubated for 4 hrs b...More data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 289nMAssay Description:In vitro binding affinity towards rat Nicotinic acetylcholine receptor alpha3-beta4 using 0.5 nM [3H]epibatidineMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 290nMAssay Description:Displacement of [3H]epibatidine from rat neuronal acetylcholine receptor subunit alpha3beta4 expressed in HEK293 cell membranes incubated for 4 hrs b...More data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 290nMAssay Description:Binding affinity of the compound was determined against Nicotinic acetylcholine receptor using [3H]-(-)-nicotine radioligandMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 310nMAssay Description:Inhibition of [3H]epibatidine binding to rat Nicotinic acetylcholine receptor alpha3-beta4More data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 314nMAssay Description:Displacement of [3H]epibatidine from rat alpha3beta4 nAChR expressed in HEK cell membranes after 2 hrsMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 321nMAssay Description:Displacement of [3H]epibatidine from rat alpha3beta4 nAChR expressed in HEK cell membranes after 2 hrsMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 325nMAssay Description:Displacement of [3H]epibatidine from rat alpha3beta4 nAChR expressed in HEK cell membranes after 2 hrsMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 334nMAssay Description:In vitro binding affinity towards rat Nicotinic acetylcholine receptor alpha3-beta4 using 0.5 nM [3H]epibatidineMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 366nMAssay Description:Displacement of [3H]epibatidine from rat alpha3beta4 nAChR by liquid scintillation countingMore data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 390nM ΔG°: -8.12kcal/molepH: 7.4 T: 2°CAssay Description:Briefly, cultured cells at >80% confluence were removed from their flasks (80 cm^2) with a disposable cell scraper and placed in 10 mL of 50 mM Tris....More data for this Ligand-Target Pair
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
TargetNeuronal acetylcholine receptor subunit alpha-3/beta-4(Rattus norvegicus (Rat))
Sri International
Curated by ChEMBL
Sri International
Curated by ChEMBL
Affinity DataKi: 405nMAssay Description:In vitro binding affinity towards rat Nicotinic acetylcholine receptor alpha3-beta4 using 0.5 nM [3H]epibatidineMore data for this Ligand-Target Pair