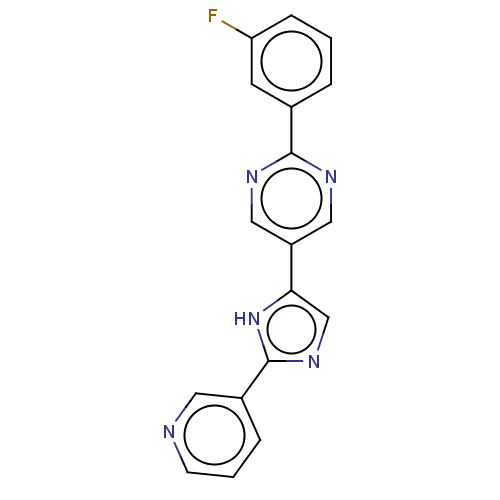
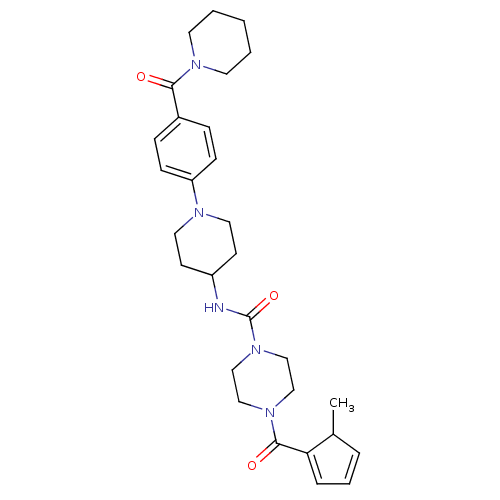
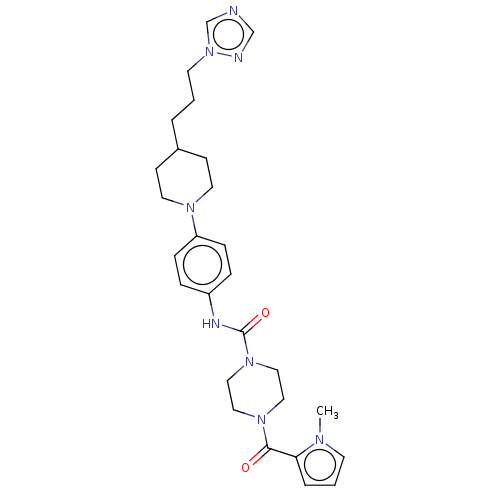
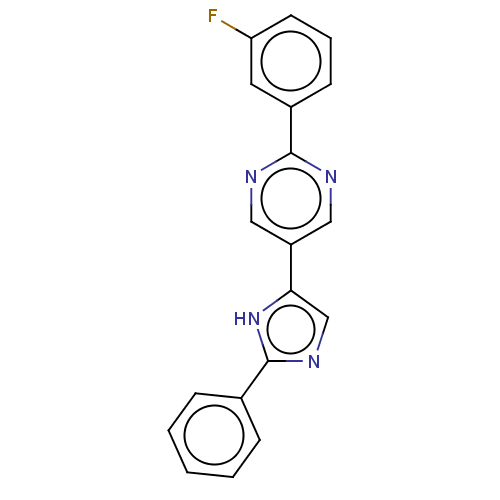
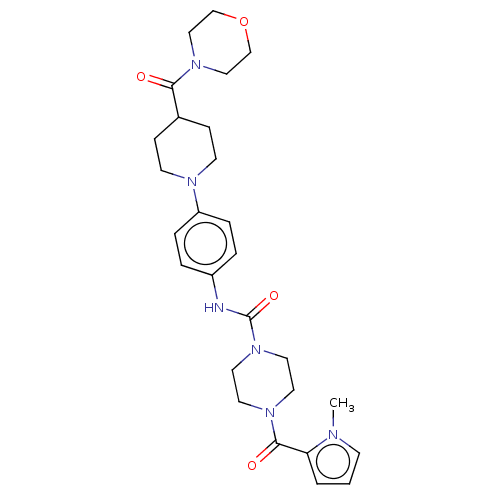
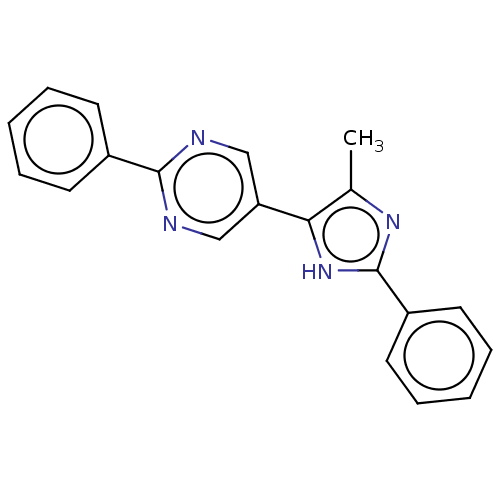
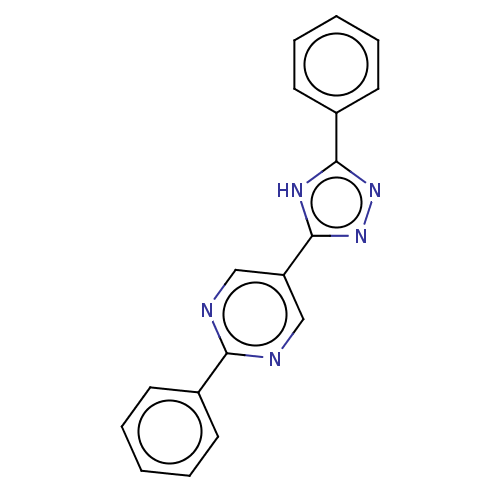
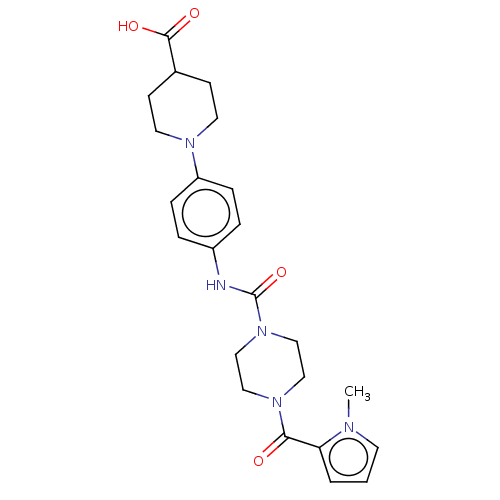
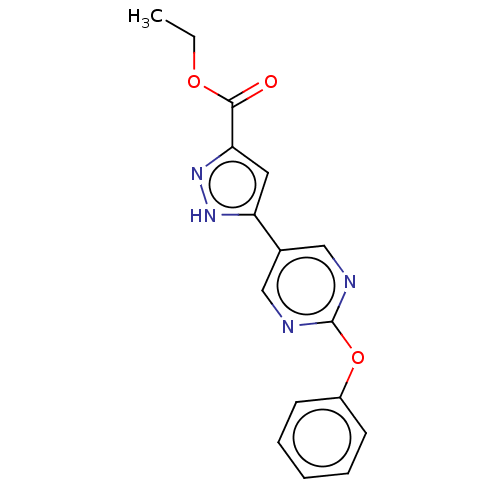
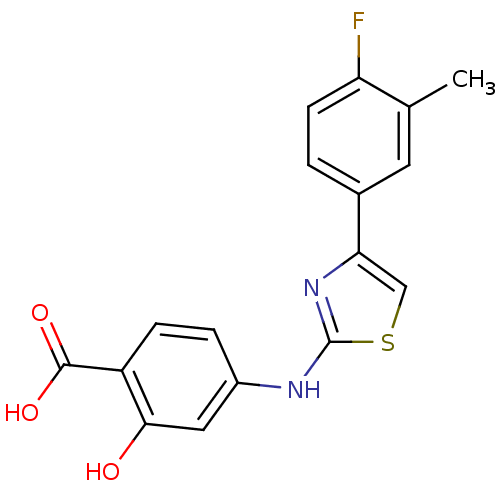
Affinity DataIC50: 4nMpH: 8.0 T: 2°CAssay Description:The assay is carried out by the following steps: 1. Inhibitor screening is performed in 100 mM Tris-HCl, pH 8.0 containing 1 mM GSH, 1 mM MgCl.sub.2,...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMpH: 8.0 T: 2°CAssay Description:The assay is carried out by the following steps: 1. Inhibitor screening is performed in 100 mM Tris-HCl, pH 8.0 containing 1 mM GSH, 1 mM MgCl.sub.2,...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMpH: 7.4Assay Description:I. Assay Solutions a. Preparation of 0.1M K2HPO4/KH2PO4 buffer (pH 7.4) Prepare 0.1 M KH2PO4 from 1M KH2PO4 (Sigma, Cat# P-8709) ...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMpH: 7.4Assay Description:I. Assay Solutions a. Preparation of 0.1M K2HPO4/KH2PO4 buffer (pH 7.4) Prepare 0.1 M KH2PO4 from 1M KH2PO4 (Sigma, Cat# P-8709) ...More data for this Ligand-Target Pair
Affinity DataIC50: 12nMpH: 8.0 T: 2°CAssay Description:The assay is carried out by the following steps: 1. Inhibitor screening is performed in 100 mM Tris-HCl, pH 8.0 containing 1 mM GSH, 1 mM MgCl.sub.2,...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMpH: 8.0Assay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 21nMpH: 7.2 T: 2°CAssay Description:The PGDS glutathione-S-transferase (GST) activity was measured by using MonoChloroBimane (MCB) as a chromogenic substrate. The assay was run at 384-w...More data for this Ligand-Target Pair
Affinity DataIC50: 21nMpH: 8.0 T: 2°CAssay Description:The assay is carried out by the following steps: 1. Inhibitor screening is performed in 100 mM Tris-HCl, pH 8.0 containing 1 mM GSH, 1 mM MgCl.sub.2,...More data for this Ligand-Target Pair
Affinity DataIC50: 23nMpH: 8.0 T: 2°CAssay Description:The assay is carried out by the following steps: 1. Inhibitor screening is performed in 100 mM Tris-HCl, pH 8.0 containing 1 mM GSH, 1 mM MgCl.sub.2,...More data for this Ligand-Target Pair
Affinity DataIC50: 26nMpH: 7.4Assay Description:I. Assay Solutions a. Preparation of 0.1M K2HPO4/KH2PO4 buffer (pH 7.4) Prepare 0.1 M KH2PO4 from 1M KH2PO4 (Sigma, Cat# P-8709) ...More data for this Ligand-Target Pair
Affinity DataIC50: 27nMpH: 8.0Assay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 27nMpH: 8.0Assay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 31nMpH: 8.0Assay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 32nMpH: 8.0 T: 2°CAssay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 37nMpH: 8.0 T: 2°CAssay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 39nMpH: 8.0Assay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 40nMpH: 8.0 T: 2°CAssay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 41nMpH: 8.0 T: 2°CAssay Description:The assay is carried out by the following steps: 1. Inhibitor screening is performed in 100 mM Tris-HCl, pH 8.0 containing 1 mM GSH, 1 mM MgCl.sub.2,...More data for this Ligand-Target Pair
Affinity DataIC50: 41nMpH: 8.0Assay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 43nMpH: 8.0 T: 2°CAssay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 46nMpH: 8.0 T: 2°CAssay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 47nMpH: 8.0 T: 2°CAssay Description:The assay is carried out by the following steps: 1. Inhibitor screening is performed in 100 mM Tris-HCl, pH 8.0 containing 1 mM GSH, 1 mM MgCl.sub.2,...More data for this Ligand-Target Pair
Affinity DataIC50: 52nMpH: 8.0Assay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 53nMpH: 8.0 T: 2°CAssay Description:The assay is carried out by the following steps: 1. Inhibitor screening is performed in 100 mM Tris-HCl, pH 8.0 containing 1 mM GSH, 1 mM MgCl.sub.2,...More data for this Ligand-Target Pair
Affinity DataIC50: 54nMpH: 8.0 T: 2°CAssay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 58nMpH: 8.0 T: 2°CAssay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 59nMpH: 8.0 T: 2°CAssay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 59nMpH: 8.0 T: 2°CAssay Description:The assay is carried out by the following steps: 1. Inhibitor screening is performed in 100 mM Tris-HCl, pH 8.0 containing 1 mM GSH, 1 mM MgCl.sub.2,...More data for this Ligand-Target Pair
Affinity DataIC50: 60nMpH: 8.0 T: 2°CAssay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 60nMpH: 8.0 T: 2°CAssay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 61nMpH: 8.0Assay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 62nMpH: 8.0 T: 2°CAssay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 67nMpH: 8.0Assay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 67nMpH: 8.0 T: 2°CAssay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 68.4nMpH: 8.0Assay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 71nMpH: 8.0 T: 2°CAssay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 75nMpH: 7.2 T: 2°CAssay Description:The PGDS glutathione-S-transferase (GST) activity was measured by using MonoChloroBimane (MCB) as a chromogenic substrate. The assay was run at 384-w...More data for this Ligand-Target Pair
Affinity DataIC50: 75nMpH: 8.0 T: 2°CAssay Description:The assay is carried out by the following steps: 1. Inhibitor screening is performed in 100 mM Tris-HCl, pH 8.0 containing 1 mM GSH, 1 mM MgCl.sub.2,...More data for this Ligand-Target Pair
Affinity DataIC50: 76nMpH: 8.0 T: 2°CAssay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 82nMpH: 8.0 T: 2°CAssay Description:The assay is carried out by the following steps: 1. Inhibitor screening is performed in 100 mM Tris-HCl, pH 8.0 containing 1 mM GSH, 1 mM MgCl.sub.2,...More data for this Ligand-Target Pair
Affinity DataIC50: 84nMpH: 8.0 T: 2°CAssay Description:The assay is carried out by the following steps: 1. Inhibitor screening is performed in 100 mM Tris-HCl, pH 8.0 containing 1 mM GSH, 1 mM MgCl.sub.2,...More data for this Ligand-Target Pair
Affinity DataIC50: 85nMpH: 8.0 T: 2°CAssay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 86nMpH: 8.0 T: 2°CAssay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 86nMpH: 8.0Assay Description:The H-PGDS catalyzed conjugation of GSH and 1-chloro-2,4-dinitrobenzene (CDNB) was used as the biochemical assay for enzyme inhibition. Reactions wer...More data for this Ligand-Target Pair
Affinity DataIC50: 91nMpH: 8.0 T: 2°CAssay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 97nMpH: 8.0 T: 2°CAssay Description:The assay is carried out by the following steps: 1. Inhibitor screening is performed in 100 mM Tris-HCl, pH 8.0 containing 1 mM GSH, 1 mM MgCl.sub.2,...More data for this Ligand-Target Pair
Affinity DataIC50: 106nMpH: 8.0Assay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 106nMpH: 8.0 T: 2°CAssay Description:The test was carried out according to the method of Urade, Y. et al. (J. Biol. Chem., 262, 3820-3825, (1987)). More specifically, the reaction mixtur...More data for this Ligand-Target Pair
Affinity DataIC50: 130nMpH: 8.0 T: 2°CAssay Description:The assay is carried out by the following steps: 1. Inhibitor screening is performed in 100 mM Tris-HCl, pH 8.0 containing 1 mM GSH, 1 mM MgCl.sub.2,...More data for this Ligand-Target Pair
Affinity DataIC50: 138nMpH: 7.2 T: 2°CAssay Description:The PGDS glutathione-S-transferase (GST) activity was measured by using MonoChloroBimane (MCB) as a chromogenic substrate. The assay was run at 384-w...More data for this Ligand-Target Pair

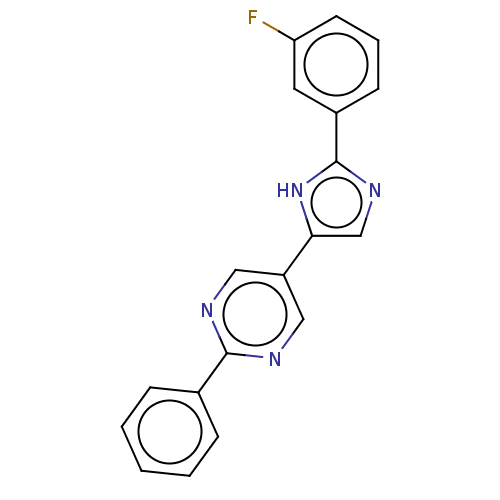
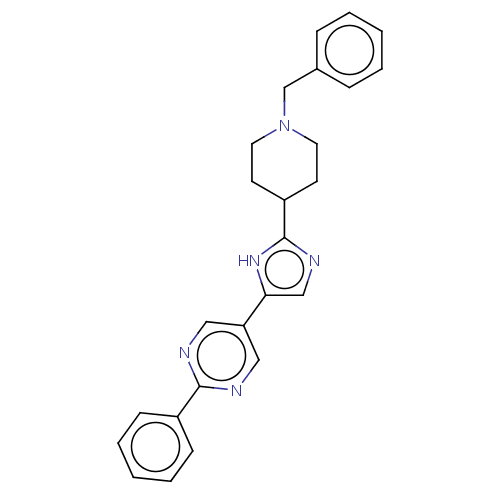
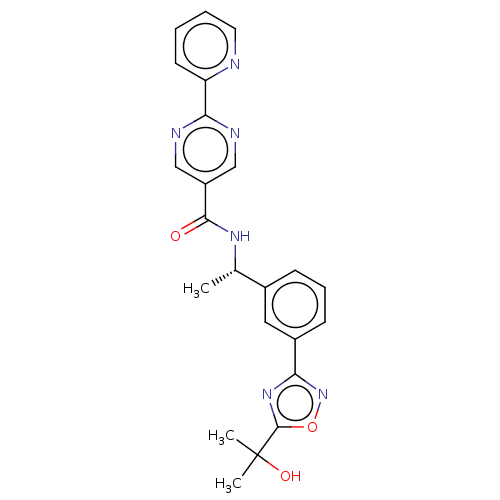
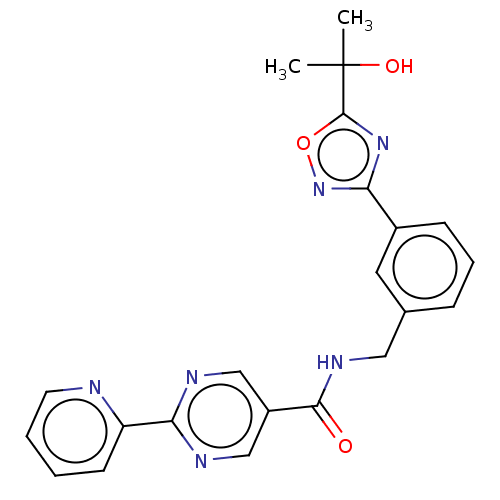
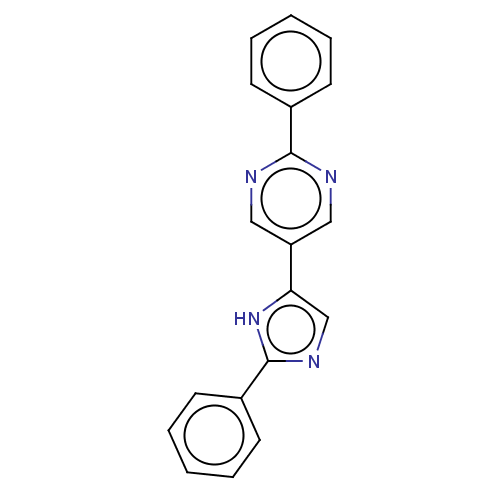
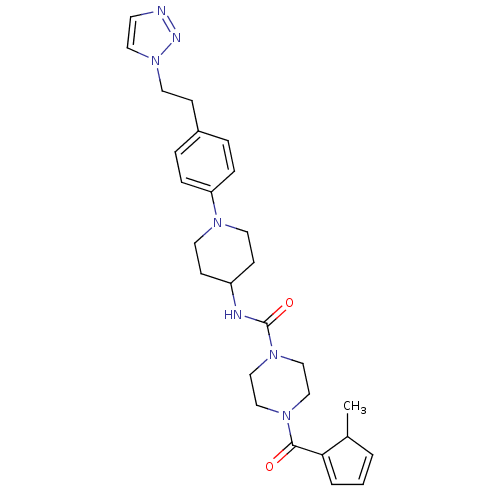
 3D Structure (crystal)
3D Structure (crystal)