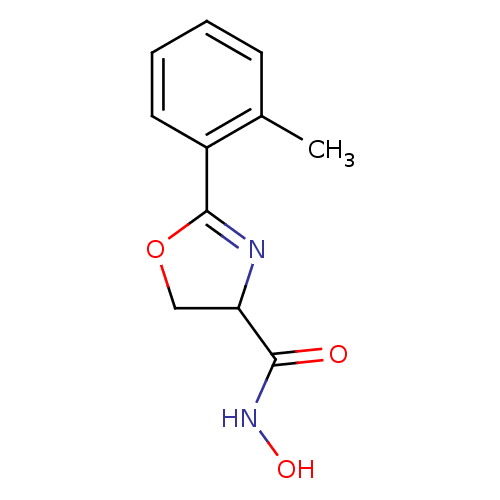
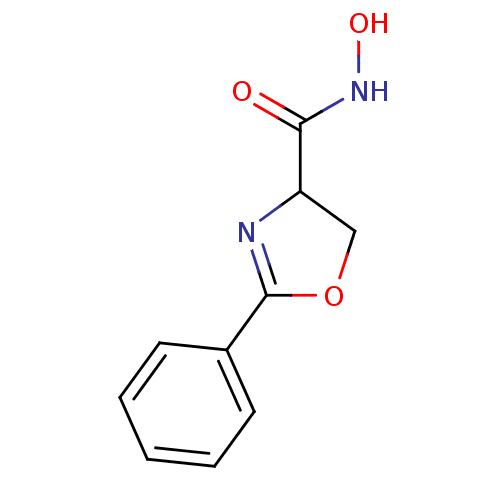
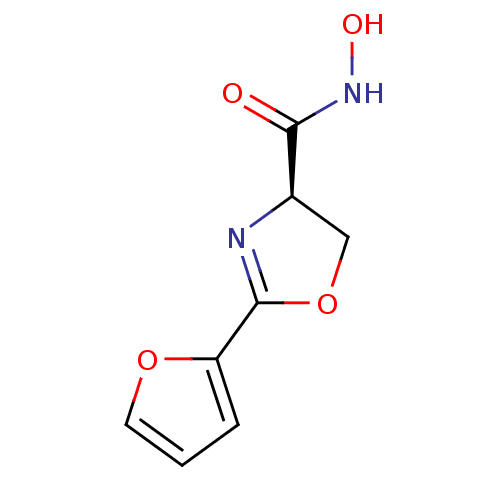
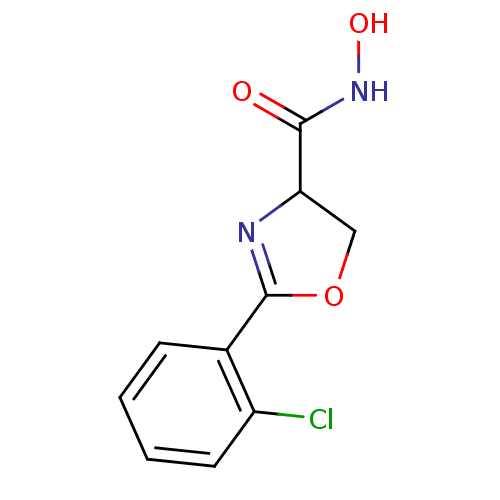
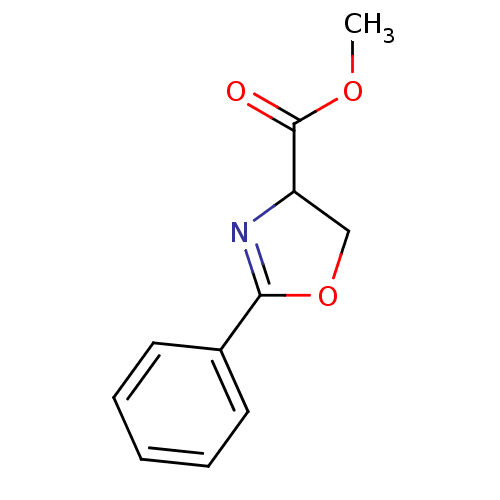
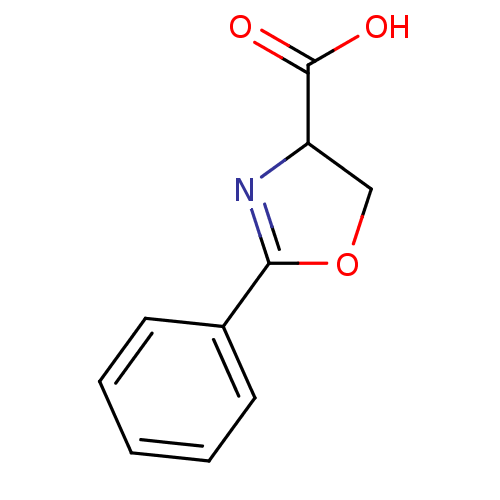
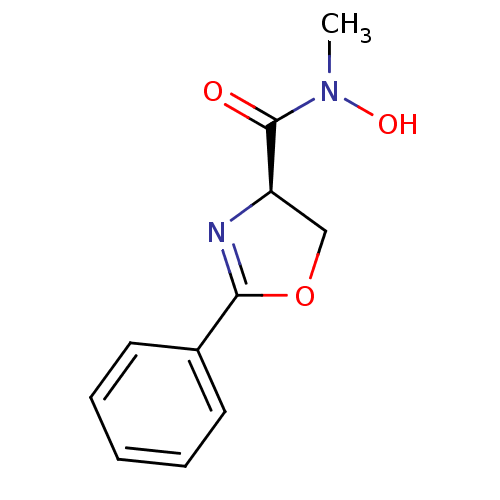
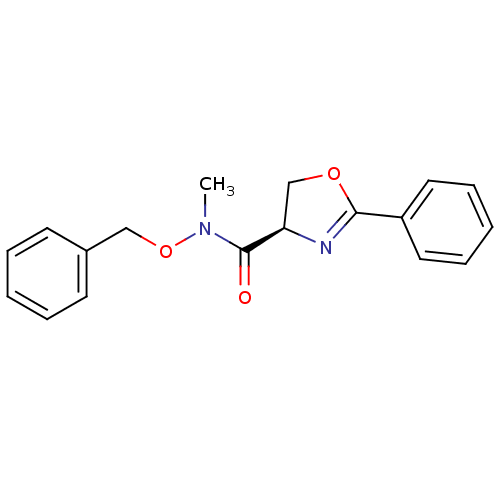
Report error Found 31 Enz. Inhib. hit(s) with all data for entry = 50008576
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
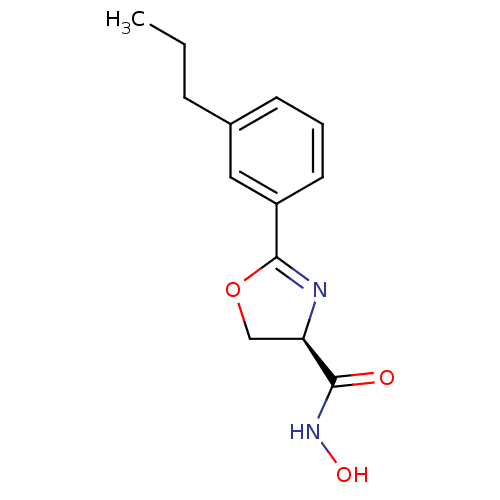
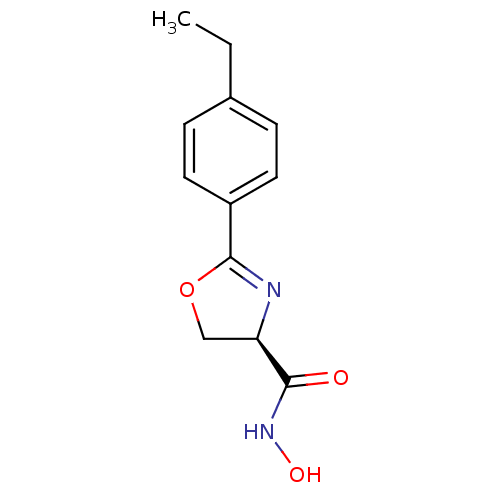
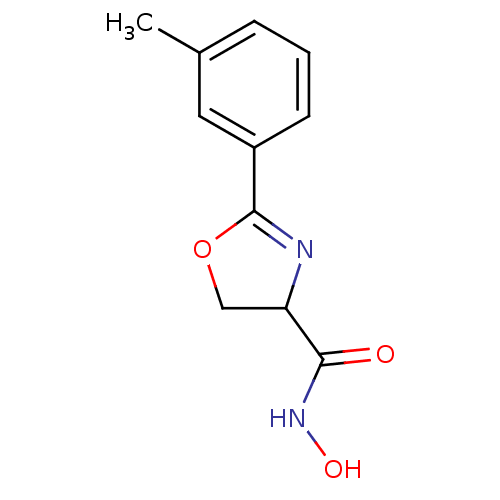
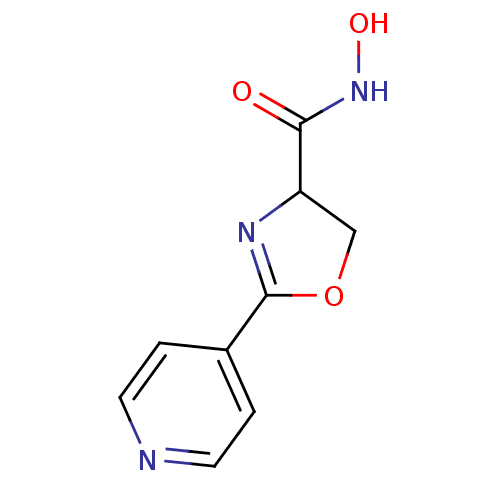
Affinity DataIC50: 20nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
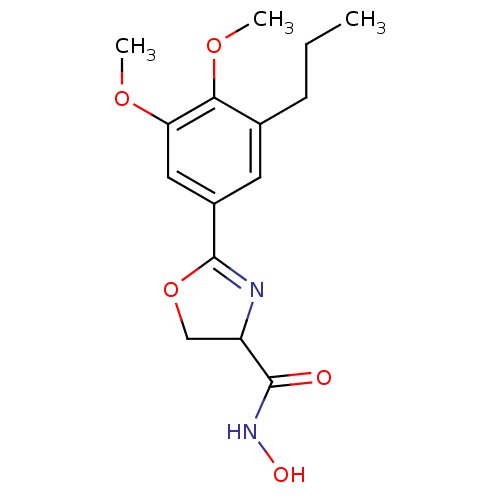
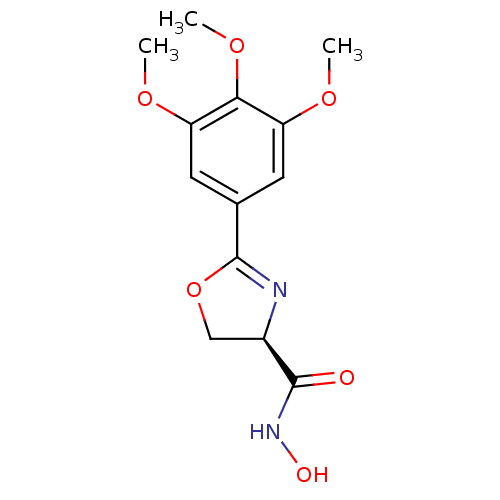
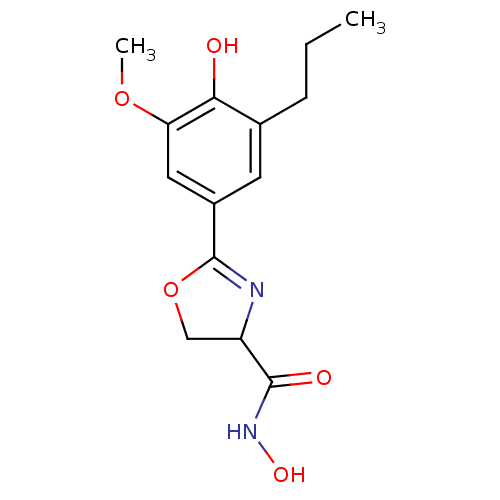
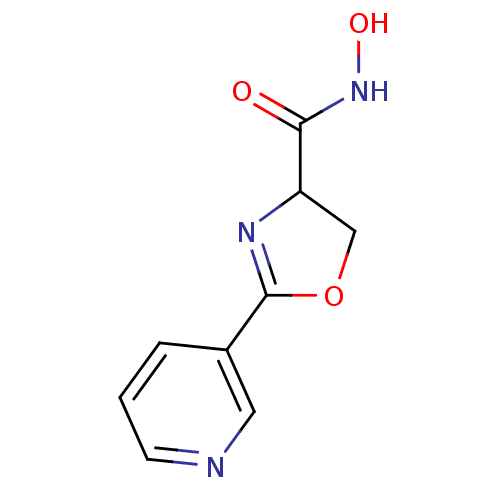
Affinity DataIC50: 30nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
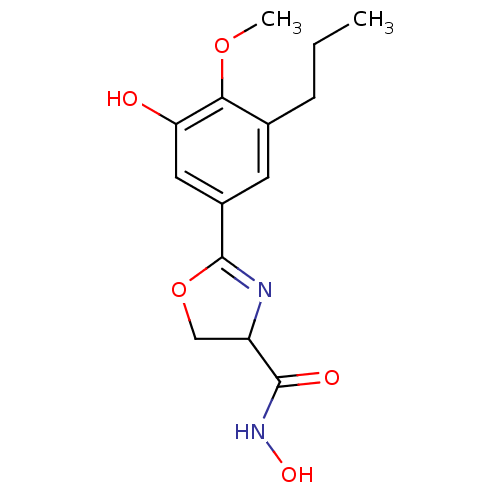
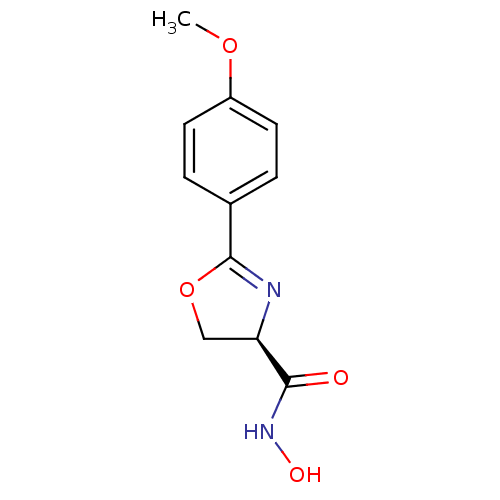
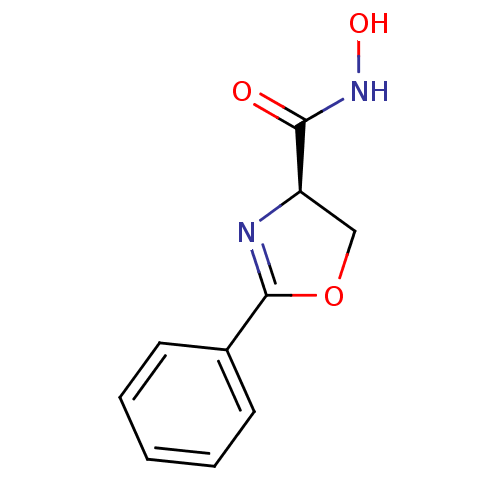
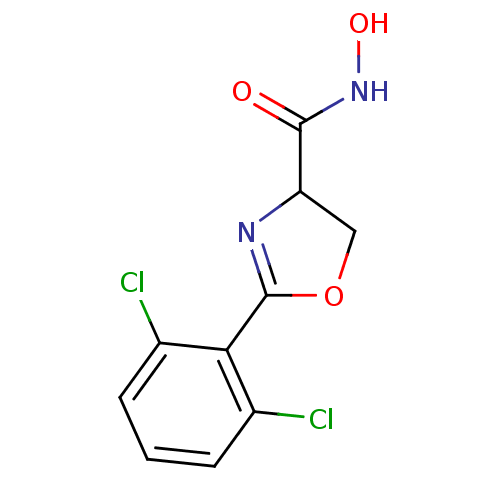
Affinity DataIC50: 50nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
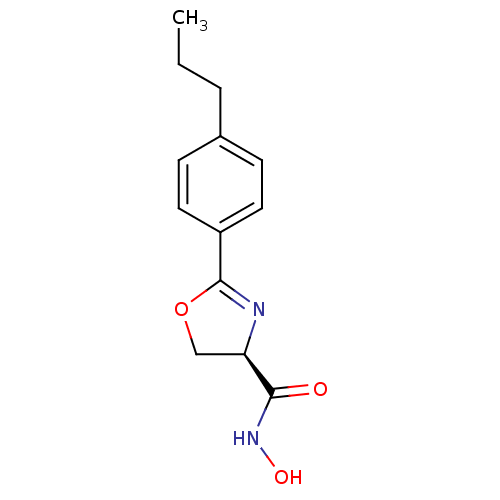
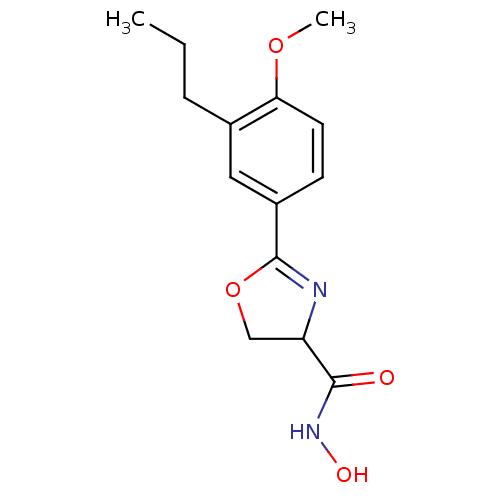
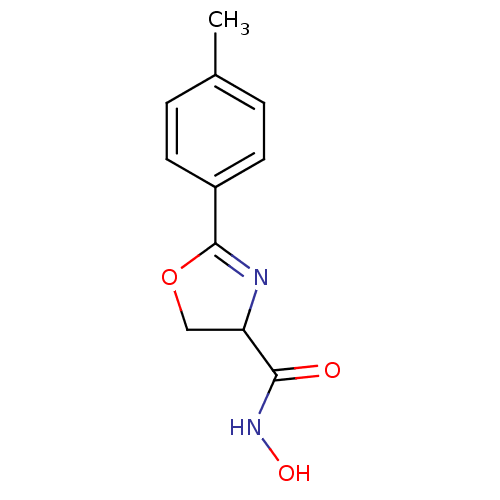
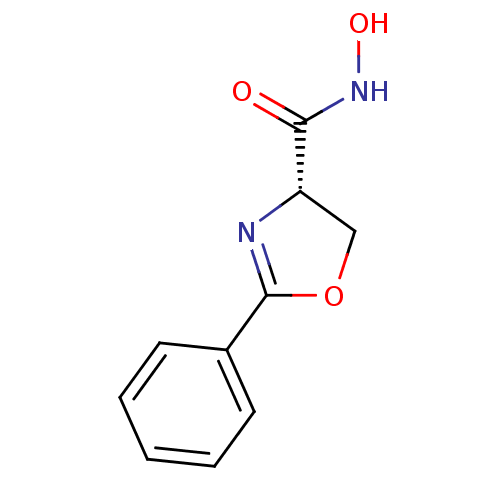
Affinity DataIC50: 50nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using coupled radiochemical assay (WAVE) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 50nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 100nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 100nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 200nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 300nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 300nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using coupled radiochemical assay (WAVE) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 300nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 600nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 1.40E+3nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 1.40E+3nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using coupled radiochemical assay (WAVE) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 2.60E+3nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using coupled radiochemical assay (WAVE) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 9.00E+3nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 3.60E+5nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 4.00E+5nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 4.00E+5nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 4.00E+5nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
TargetUDP-3-O-acyl-N-acetylglucosamine deacetylase(Escherichia coli (strain K12))
Merck Research Laboratories
Curated by ChEMBL
Merck Research Laboratories
Curated by ChEMBL
Affinity DataIC50: 4.00E+5nMAssay Description:Inhibition of UDP-3-O-[R-3-hydroxymyristoyl]-GlcNAc deacetylase using direct deacetylase assay (DEACET) in Escherichia coli strain JB 1104More data for this Ligand-Target Pair
 3D Structure (crystal)
3D Structure (crystal)