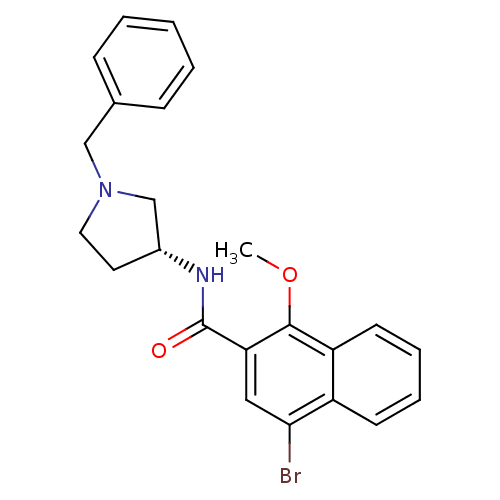
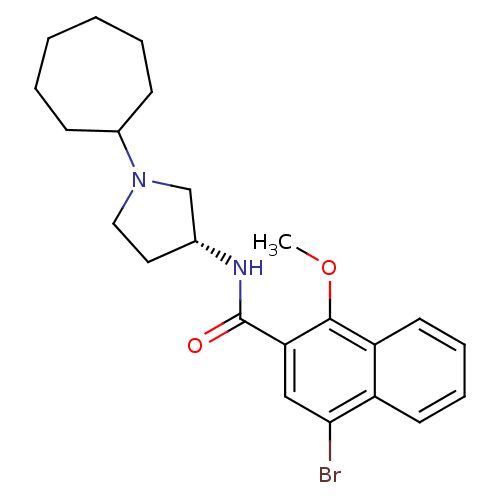
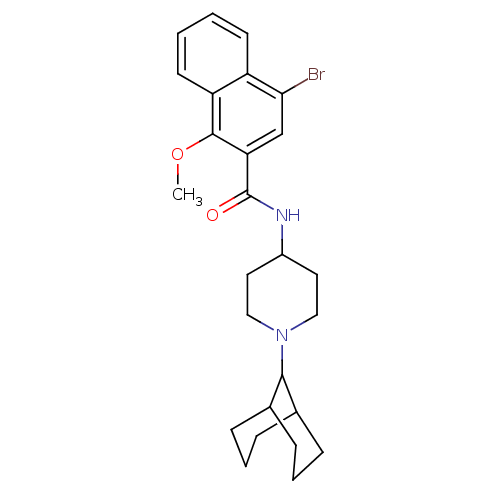
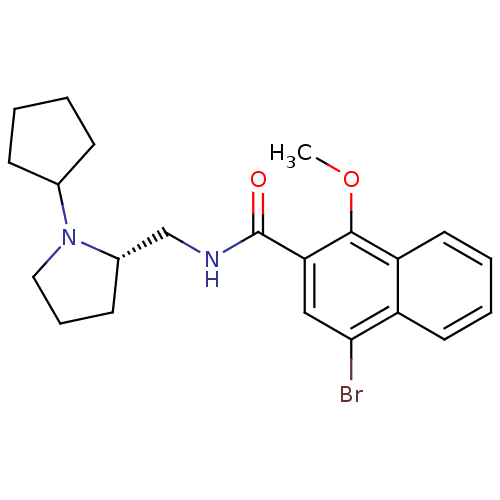
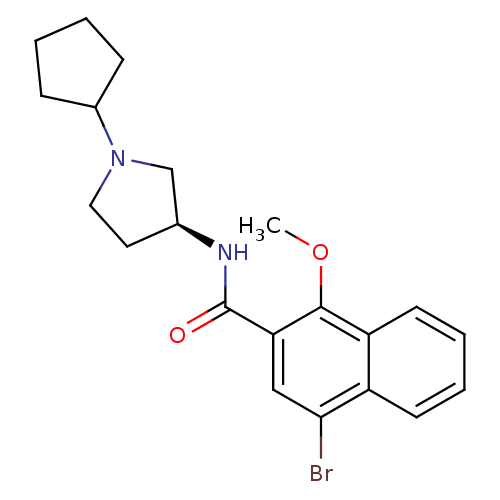
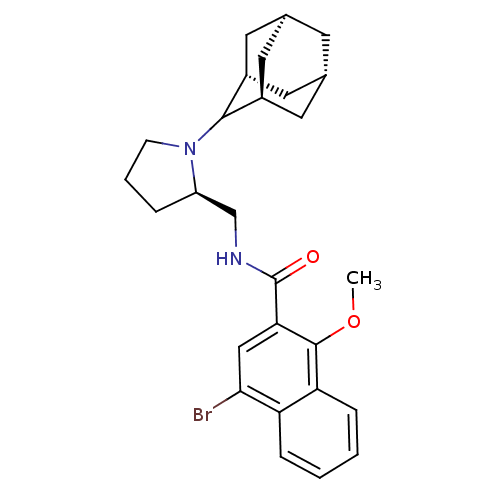
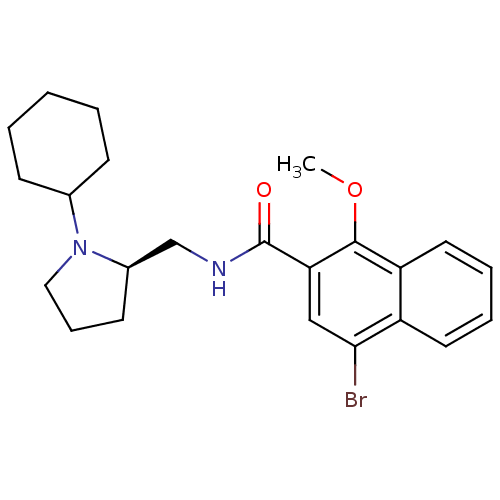
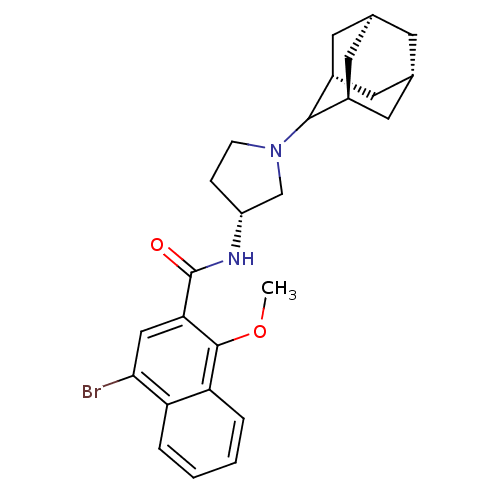
Report error Found 108 Enz. Inhib. hit(s) with all data for entry = 50010891
Affinity DataKi: 0.0400nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 0.0400nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:Binding affinity for dopamine receptor D2 long expressed in Sf9 cells using [125I]IABN radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Binding affinity for dopamine receptor D2 long expressed in Sf9 cells using [125I]IABN radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 0.400nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Human)
Wake Forest University School of Medicine
Curated by ChEMBL
Wake Forest University School of Medicine
Curated by ChEMBL
Affinity DataKi: 0.700nMAssay Description:Binding affinity for sigma-1 receptor measured on guinea pig brain membranes using [3H](+)-pentazocine as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 0.800nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Human)
Wake Forest University School of Medicine
Curated by ChEMBL
Wake Forest University School of Medicine
Curated by ChEMBL
Affinity DataKi: 1.10nMAssay Description:Binding affinity for sigma-1 receptor measured on guinea pig brain membranes using [3H](+)-pentazocine as radioligand.More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Human)
Wake Forest University School of Medicine
Curated by ChEMBL
Wake Forest University School of Medicine
Curated by ChEMBL
Affinity DataKi: 1.10nMAssay Description:Binding affinity for sigma-1 receptor measured on guinea pig brain membranes using [3H](+)-pentazocine as radioligand.More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Human)
Wake Forest University School of Medicine
Curated by ChEMBL
Wake Forest University School of Medicine
Curated by ChEMBL
Affinity DataKi: 1.20nMAssay Description:Binding affinity for sigma-1 receptor measured on guinea pig brain membranes using [3H](+)-pentazocine as radioligand.More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Human)
Wake Forest University School of Medicine
Curated by ChEMBL
Wake Forest University School of Medicine
Curated by ChEMBL
Affinity DataKi: 1.20nMAssay Description:Binding affinity for sigma-1 receptor measured on guinea pig brain membranes using [3H](+)-pentazocine as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Human)
Wake Forest University School of Medicine
Curated by ChEMBL
Wake Forest University School of Medicine
Curated by ChEMBL
Affinity DataKi: 1.5nMAssay Description:Binding affinity for sigma-1 receptor measured on guinea pig brain membranes using [3H](+)-pentazocine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Human)
Wake Forest University School of Medicine
Curated by ChEMBL
Wake Forest University School of Medicine
Curated by ChEMBL
Affinity DataKi: 1.70nMAssay Description:Binding affinity for sigma-1 receptor measured on guinea pig brain membranes using [3H](+)-pentazocine as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 1.80nMAssay Description:Binding affinity for dopamine receptor D2 long expressed in Sf9 cells using [125I]IABN radioligand.More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Human)
Wake Forest University School of Medicine
Curated by ChEMBL
Wake Forest University School of Medicine
Curated by ChEMBL
Affinity DataKi: 1.90nMAssay Description:Binding affinity for sigma-1 receptor measured on guinea pig brain membranes using [3H](+)-pentazocine as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 1.90nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 2.40nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Human)
Wake Forest University School of Medicine
Curated by ChEMBL
Wake Forest University School of Medicine
Curated by ChEMBL
Affinity DataKi: 2.5nMAssay Description:Binding affinity for sigma-1 receptor measured on guinea pig brain membranes using [3H](+)-pentazocine as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 2.5nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 2.60nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 2.90nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Human)
Wake Forest University School of Medicine
Curated by ChEMBL
Wake Forest University School of Medicine
Curated by ChEMBL
Affinity DataKi: 3.20nMAssay Description:Binding affinity for sigma-1 receptor measured on guinea pig brain membranes using [3H](+)-pentazocine as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 3.20nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Human)
Wake Forest University School of Medicine
Curated by ChEMBL
Wake Forest University School of Medicine
Curated by ChEMBL
Affinity DataKi: 3.60nMAssay Description:Binding affinity for sigma-1 receptor measured on guinea pig brain membranes using [3H](+)-pentazocine as radioligand.More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Human)
Wake Forest University School of Medicine
Curated by ChEMBL
Wake Forest University School of Medicine
Curated by ChEMBL
Affinity DataKi: 4nMAssay Description:Binding affinity for sigma-1 receptor measured on guinea pig brain membranes using [3H](+)-pentazocine as radioligand.More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Human)
Wake Forest University School of Medicine
Curated by ChEMBL
Wake Forest University School of Medicine
Curated by ChEMBL
Affinity DataKi: 4.80nMAssay Description:Binding affinity for sigma-1 receptor measured on guinea pig brain membranes using [3H](+)-pentazocine as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 5.5nMAssay Description:Binding affinity for dopamine receptor D2 long expressed in Sf9 cells using [125I]IABN radioligand.More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Human)
Wake Forest University School of Medicine
Curated by ChEMBL
Wake Forest University School of Medicine
Curated by ChEMBL
Affinity DataKi: 5.60nMAssay Description:Binding affinity for sigma-1 receptor measured on guinea pig brain membranes using [3H](+)-pentazocine as radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 5.70nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Human)
Wake Forest University School of Medicine
Curated by ChEMBL
Wake Forest University School of Medicine
Curated by ChEMBL
Affinity DataKi: 5.80nMAssay Description:Binding affinity for sigma-1 receptor measured on guinea pig brain membranes using [3H](+)-pentazocine as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 5.80nMAssay Description:Binding affinity for dopamine receptor D2 long expressed in Sf9 cells using [125I]IABN radioligand.More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Human)
Wake Forest University School of Medicine
Curated by ChEMBL
Wake Forest University School of Medicine
Curated by ChEMBL
Affinity DataKi: 5.90nMAssay Description:Binding affinity for sigma-1 receptor measured on guinea pig brain membranes using [3H](+)-pentazocine as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 5.90nMAssay Description:Binding affinity for dopamine receptor D2 long expressed in Sf9 cells using [125I]IABN radioligand.More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Human)
Wake Forest University School of Medicine
Curated by ChEMBL
Wake Forest University School of Medicine
Curated by ChEMBL
Affinity DataKi: 6.20nMAssay Description:Binding affinity for sigma-1 receptor measured on guinea pig brain membranes using [3H](+)-pentazocine as radioligand.More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Human)
Wake Forest University School of Medicine
Curated by ChEMBL
Wake Forest University School of Medicine
Curated by ChEMBL
Affinity DataKi: 6.20nMAssay Description:Binding affinity for sigma-1 receptor measured on guinea pig brain membranes using [3H](+)-pentazocine as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 7.5nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 7.60nMAssay Description:Binding affinity for dopamine receptor D2 long expressed in Sf9 cells using [125I]IABN radioligandMore data for this Ligand-Target Pair
Affinity DataKi: 9.70nMAssay Description:Binding affinity for dopamine receptor D2 long expressed in Sf9 cells using [125I]IABN radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 11.4nMAssay Description:Binding affinity for dopamine receptor D2 long expressed in Sf9 cells using [125I]IABN radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 11.8nMAssay Description:Binding affinity for dopamine receptor D2 long expressed in Sf9 cells using [125I]IABN radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Binding affinity for dopamine receptor D2 long expressed in Sf9 cells using [125I]IABN radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 12nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
TargetSigma non-opioid intracellular receptor 1(Human)
Wake Forest University School of Medicine
Curated by ChEMBL
Wake Forest University School of Medicine
Curated by ChEMBL
Affinity DataKi: 12.7nMAssay Description:Binding affinity for sigma-1 receptor measured on guinea pig brain membranes using [3H](+)-pentazocine as radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
Affinity DataKi: 13.2nMAssay Description:Binding affinity for dopamine receptor D3 expressed in Sf9 cells using [125I]IABN the radioligand.More data for this Ligand-Target Pair
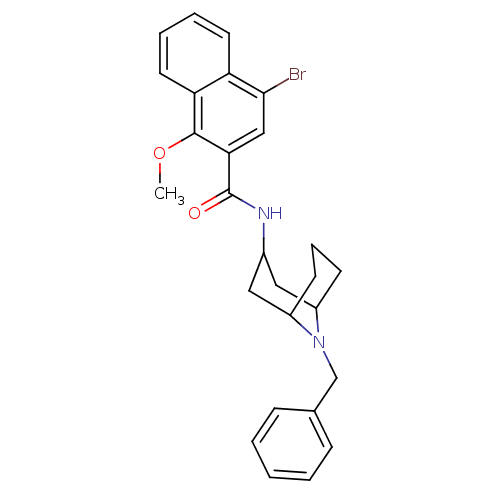

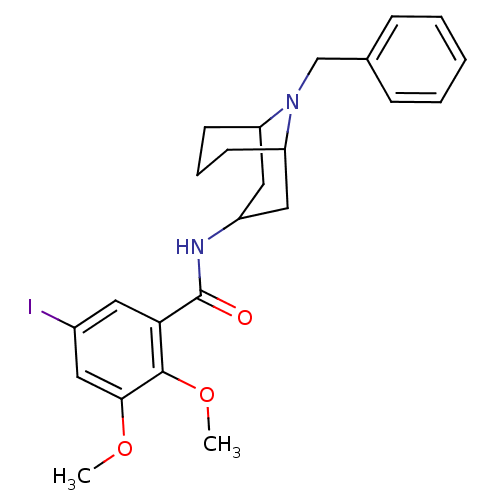
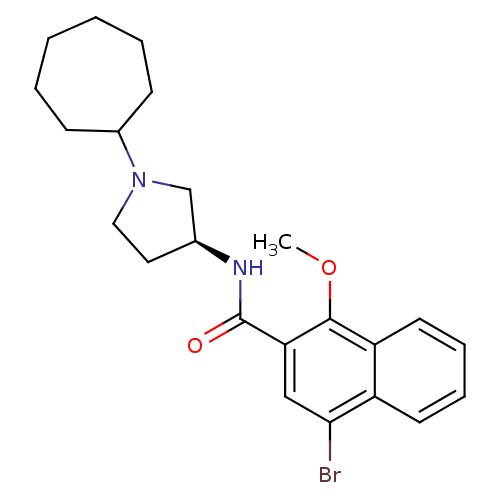
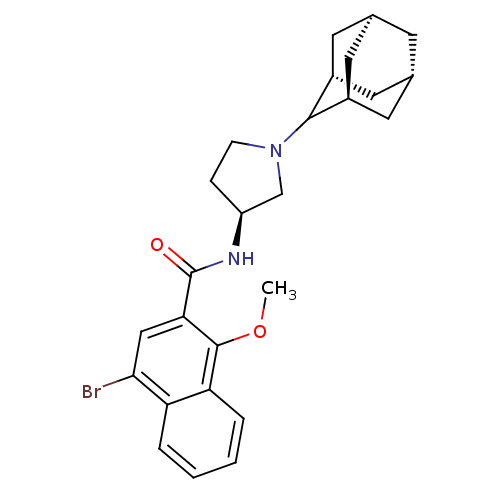
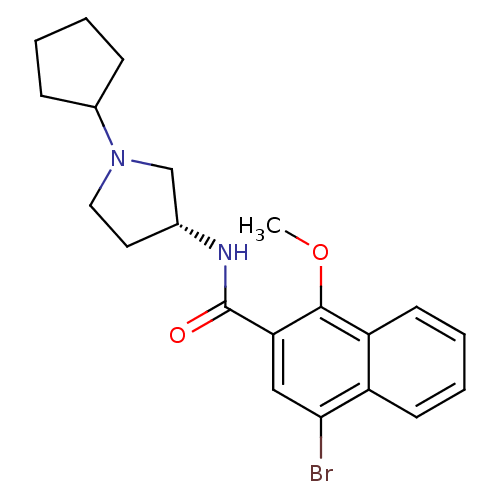
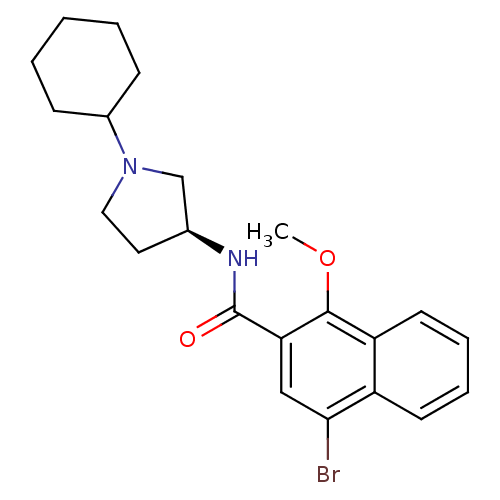
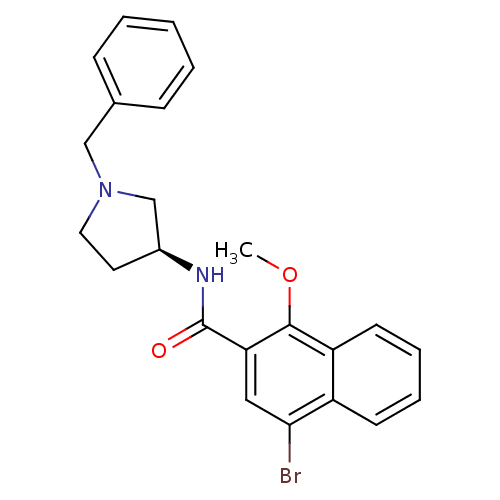
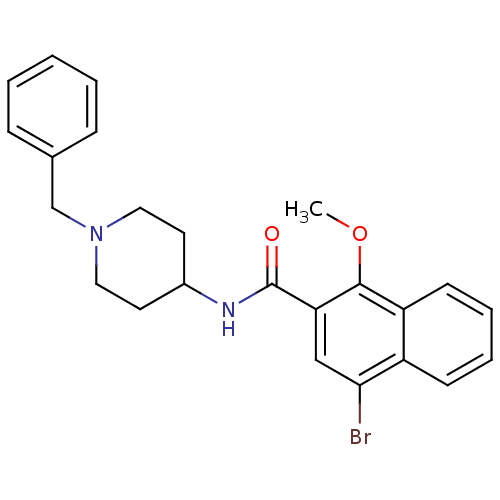
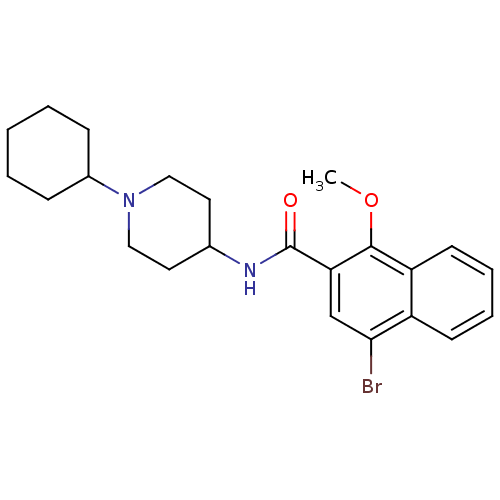
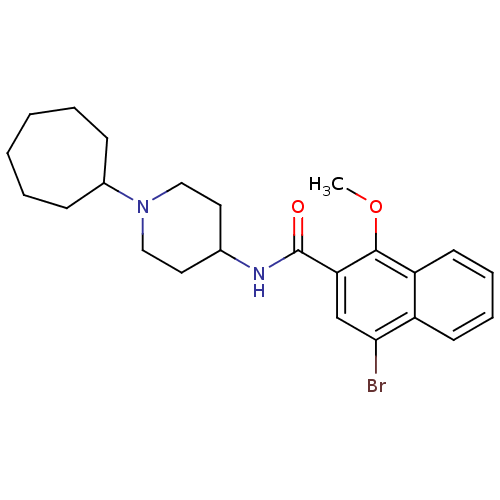
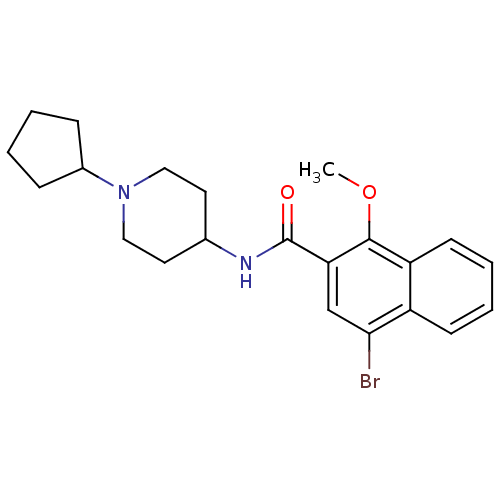
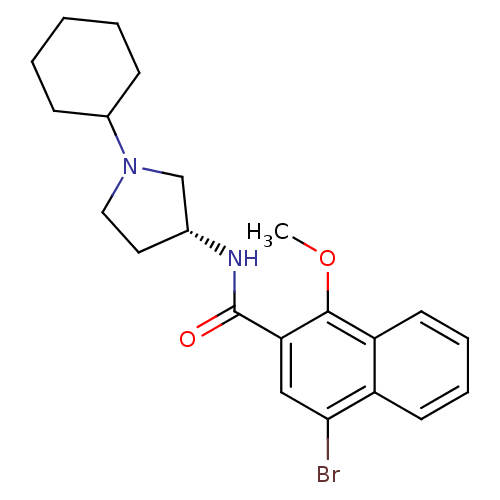
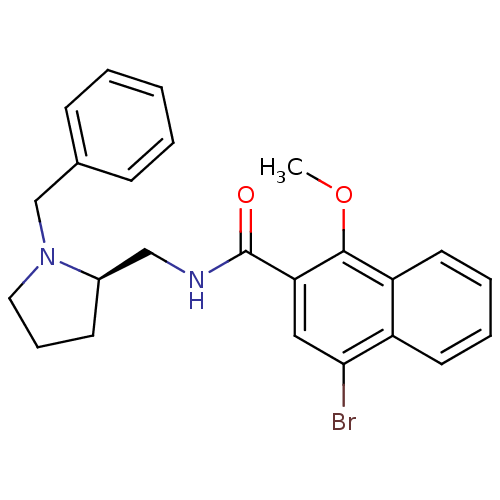
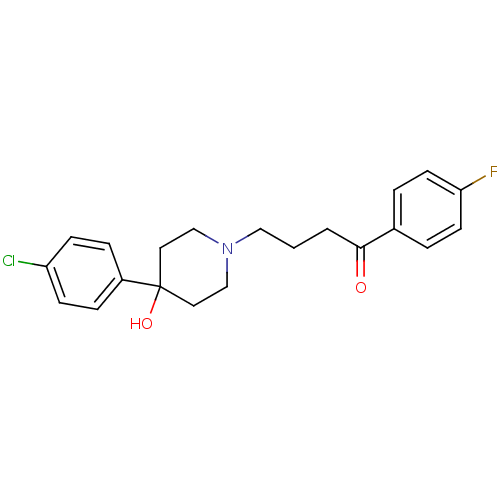
 3D Structure (crystal)
3D Structure (crystal)