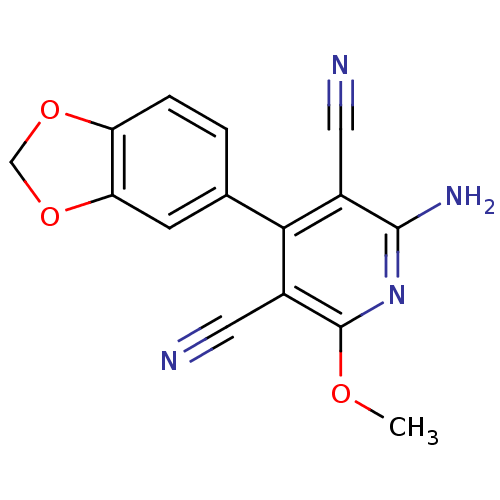
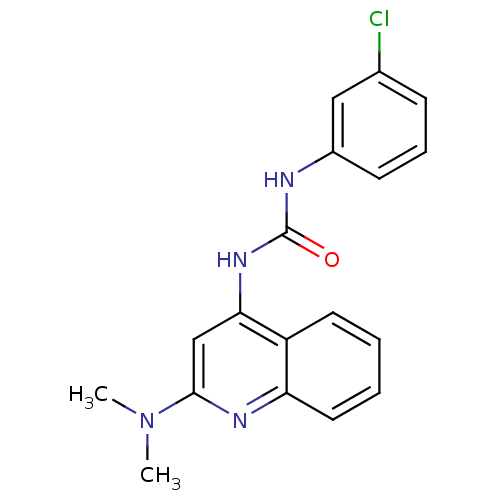
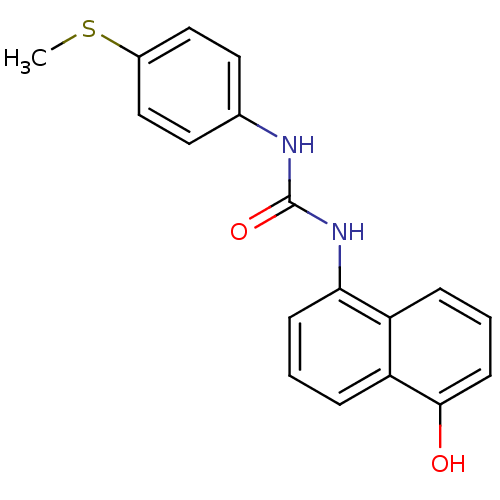
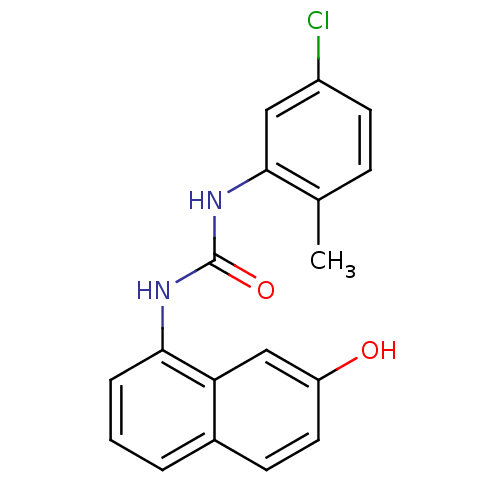
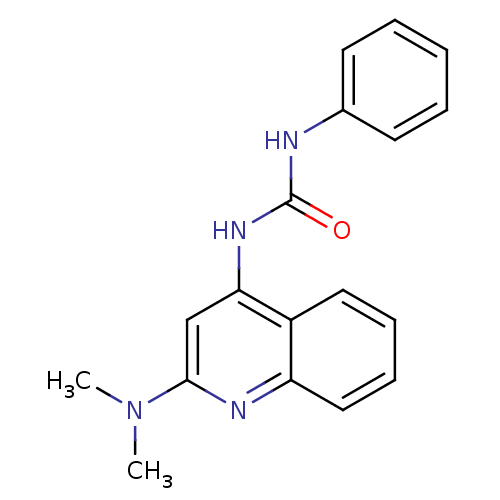
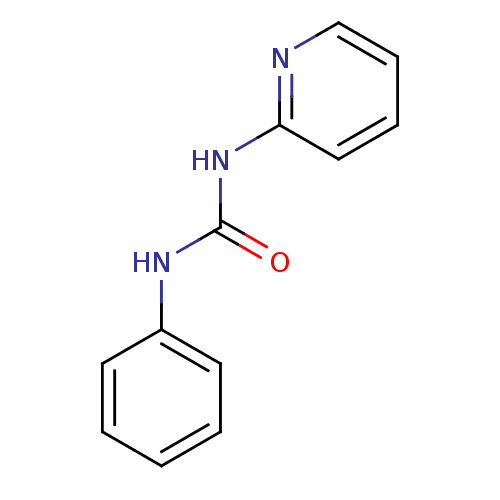
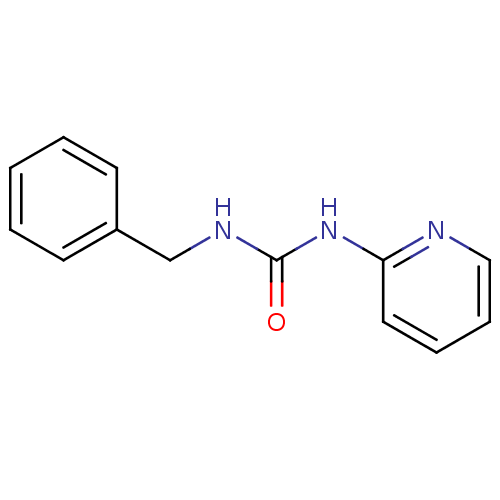
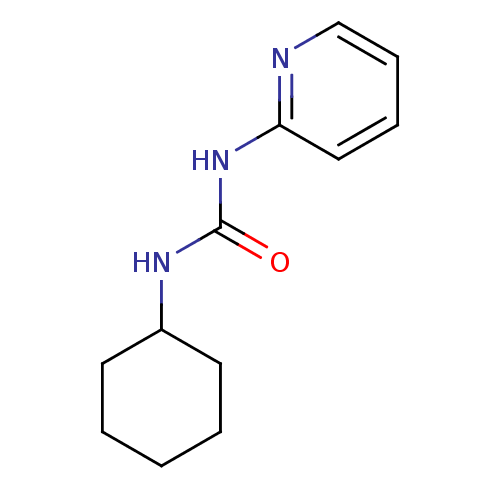
Report error Found 28 Enz. Inhib. hit(s) with all data for entry = 848
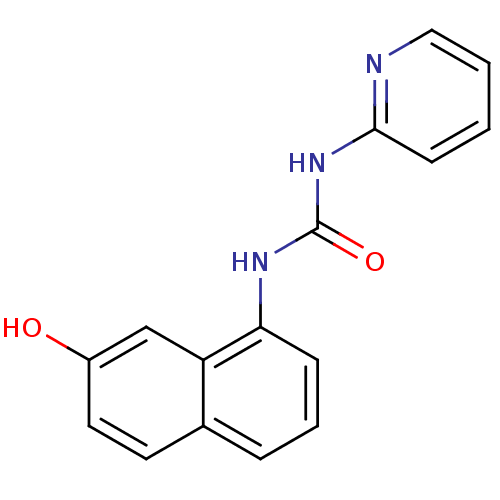
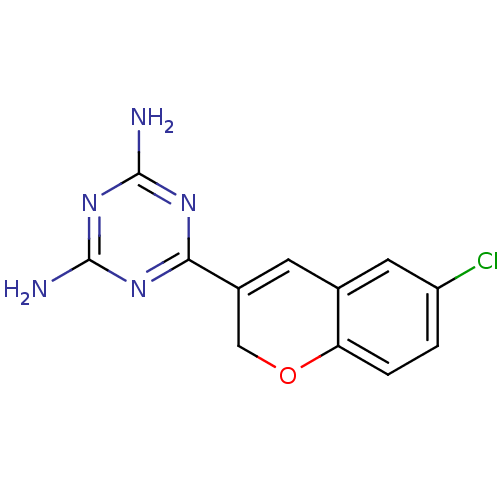
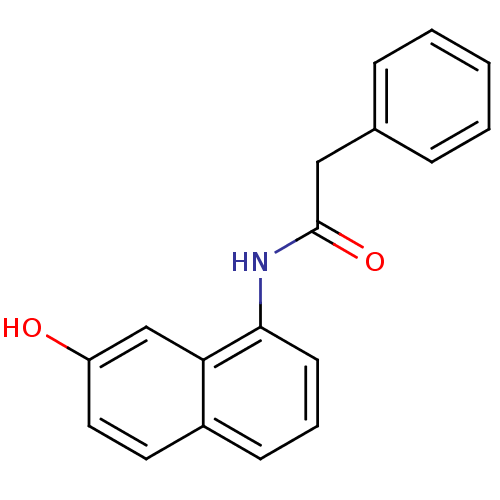
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
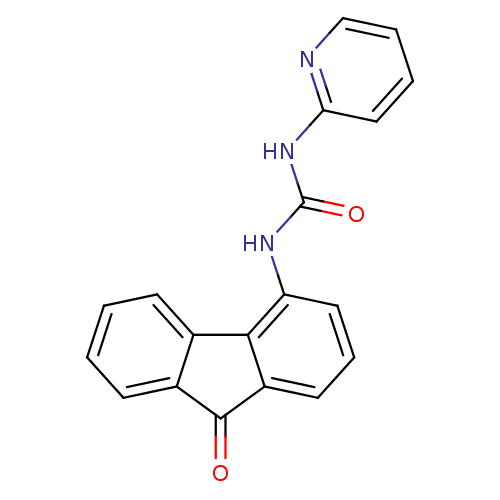
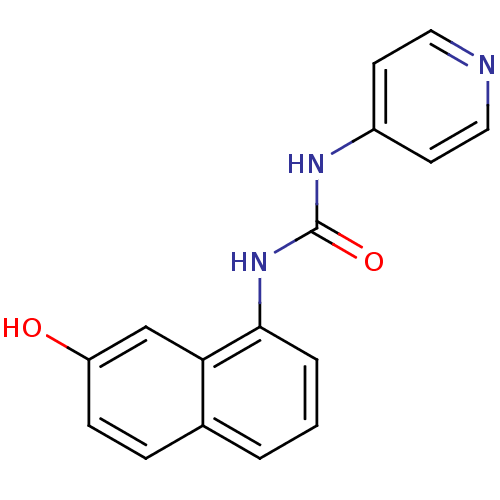
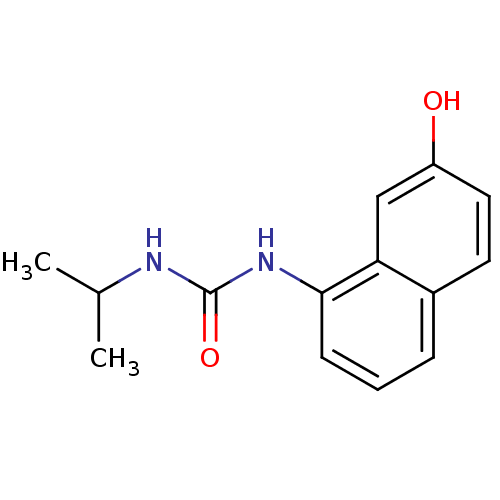
Affinity DataIC50: 100nMT: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
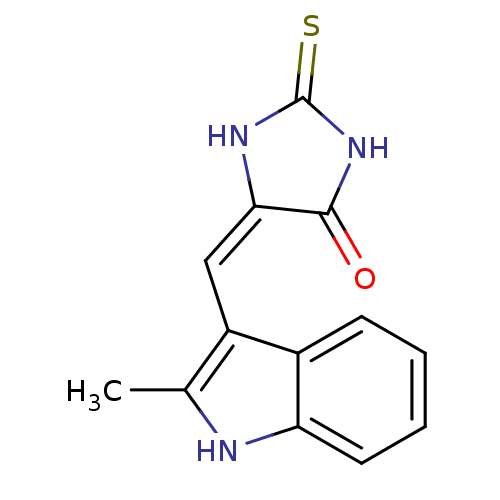
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
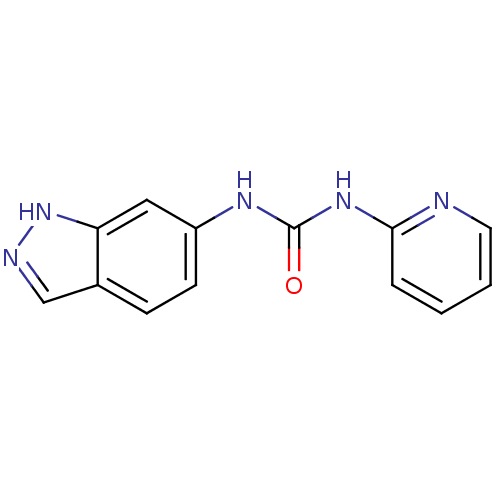
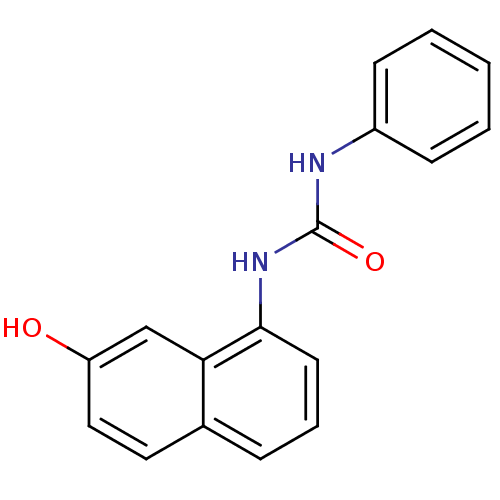
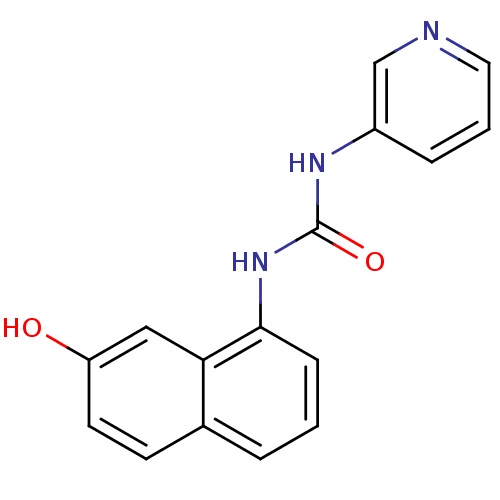
Affinity DataIC50: 670nMT: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
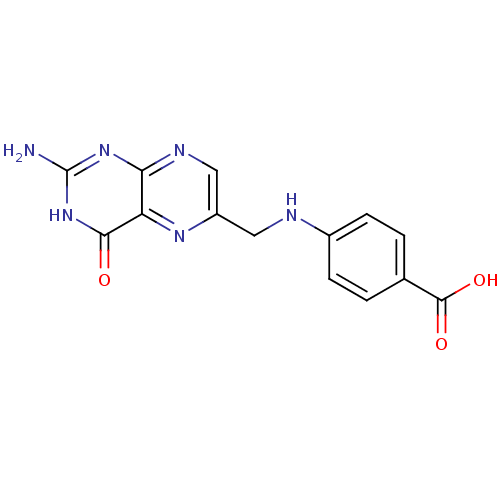
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
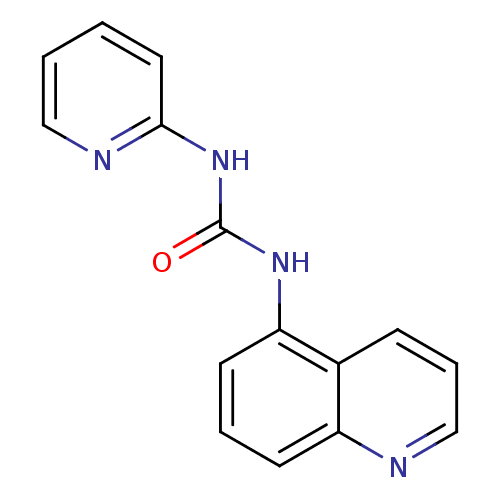
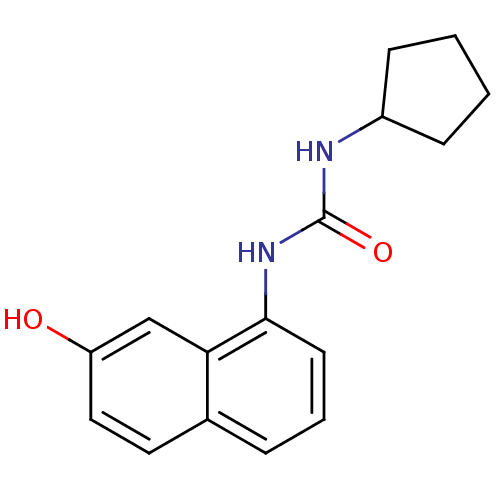
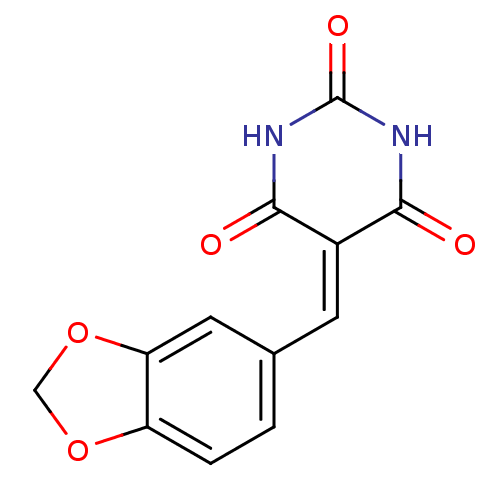
Affinity DataIC50: 2.40E+3nMT: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
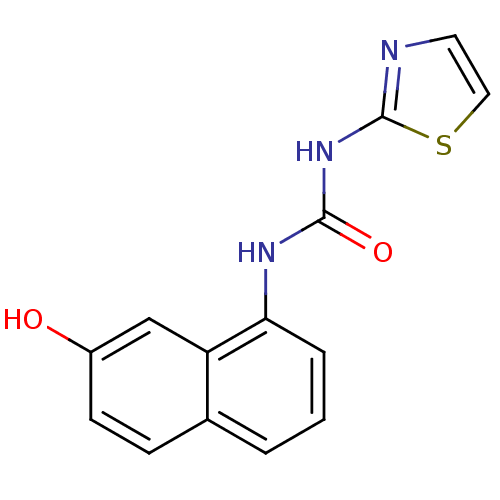
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
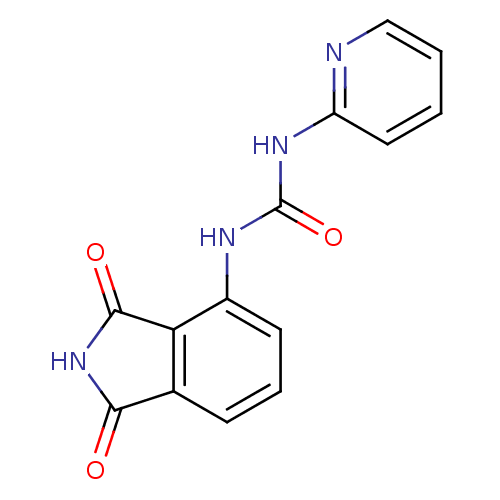
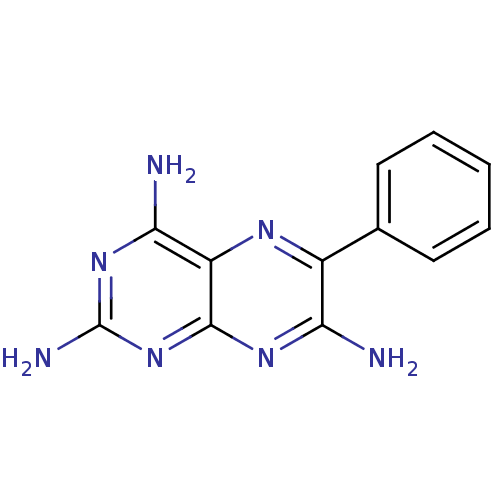
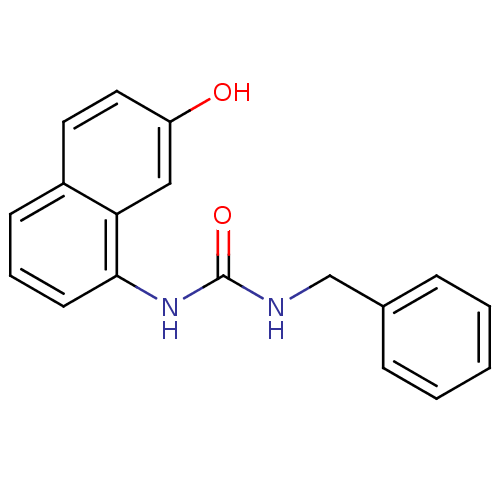
Affinity DataIC50: 3.60E+3nMT: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 7.60E+3nMT: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 1.40E+4nMpH: 7.4 T: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 1.60E+4nMpH: 7.4 T: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 2.30E+4nMT: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 3.60E+4nMpH: 7.4 T: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 3.60E+4nMpH: 7.4 T: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 4.00E+4nMpH: 7.4 T: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 4.40E+4nMpH: 7.4 T: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 7.70E+4nMpH: 7.4 T: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 9.00E+4nMpH: 7.4 T: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 1.00E+5nMpH: 7.4 T: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 1.00E+5nMT: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 1.00E+5nMT: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 1.00E+5nMT: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 1.10E+5nMT: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 1.20E+5nMT: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 1.50E+5nMT: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 2.10E+5nMpH: 7.4 T: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 2.20E+5nMT: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 3.40E+5nMT: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 3.50E+5nMpH: 7.4 T: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 4.10E+5nMT: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 4.50E+5nMpH: 7.4 T: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair
TargetCyclin-dependent kinase 4/G1/S-specific cyclin-D1 [L188C](Human)
Banyu Tsukuba Research Institute
Banyu Tsukuba Research Institute
Affinity DataIC50: 1.00E+6nMT: 2°CAssay Description:In vitro kinase assays using synthetic peptides and purified enzymes were incubated at 30°C for 45 min in buffer that contained 50 uM ATP, and d...More data for this Ligand-Target Pair