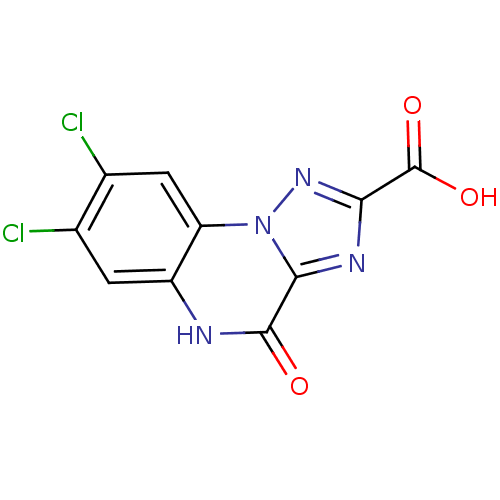
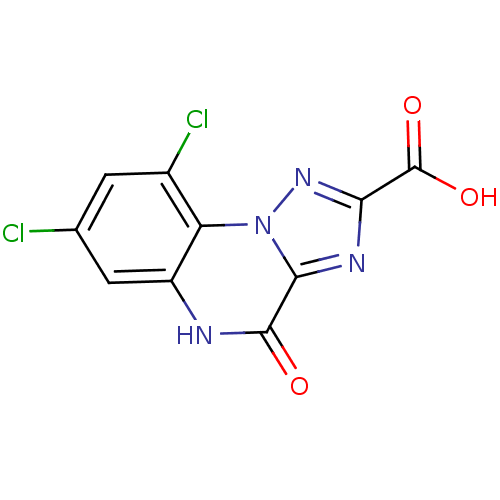
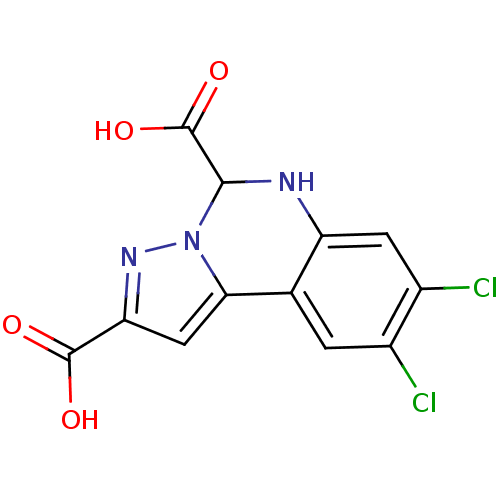
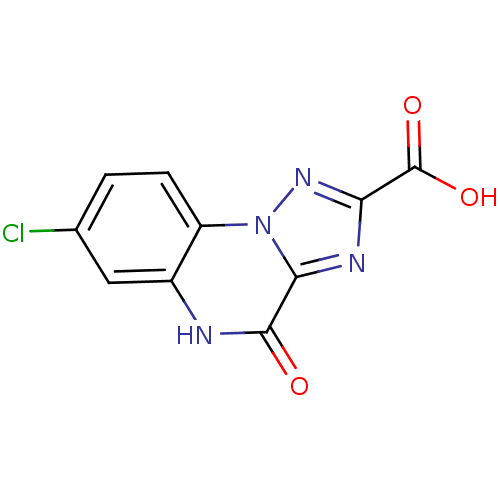
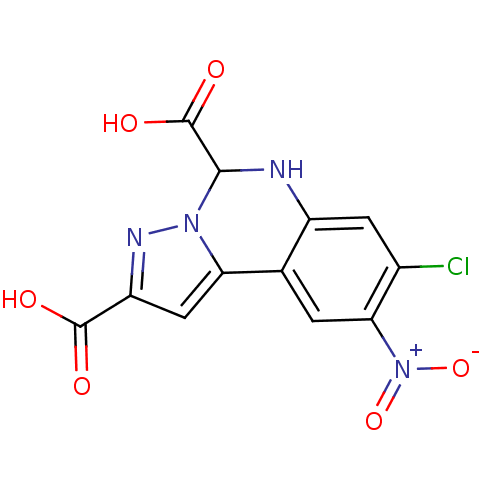
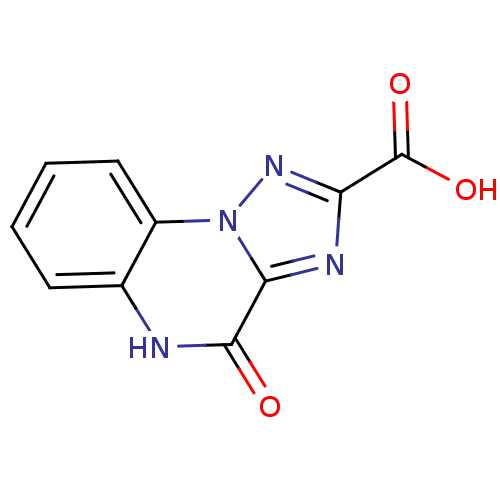
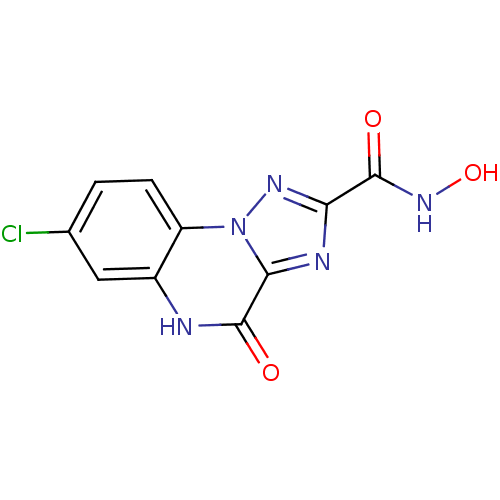
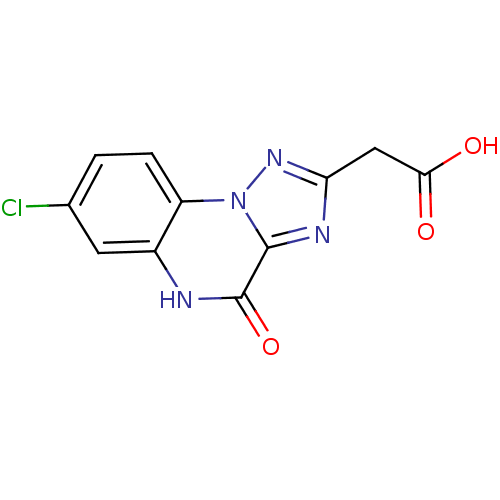
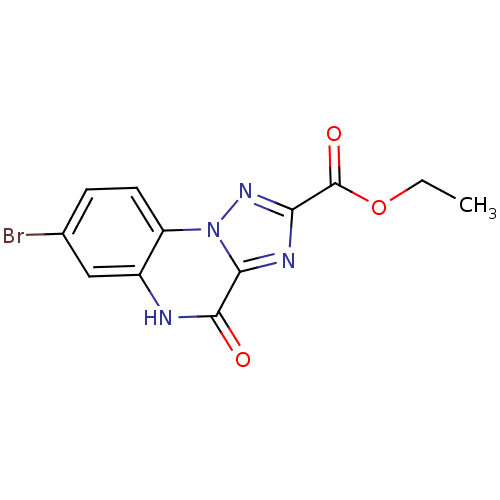
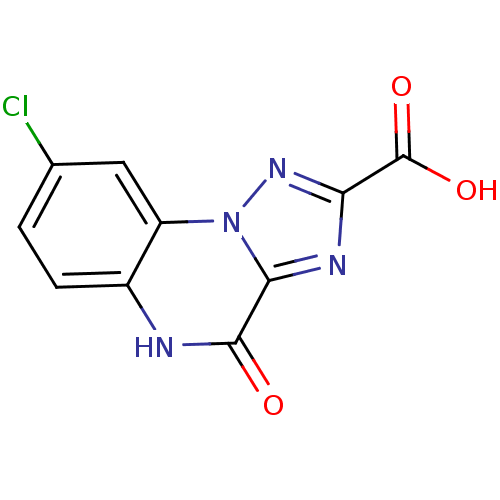
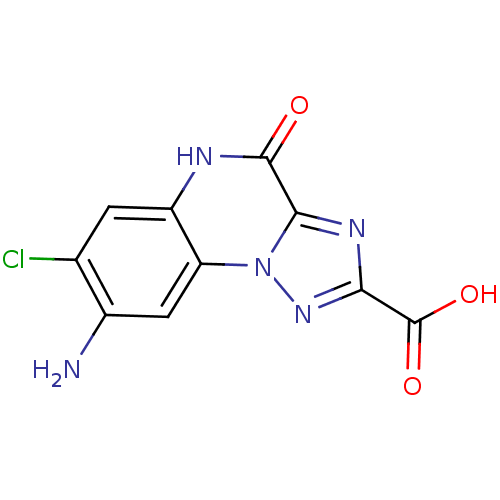
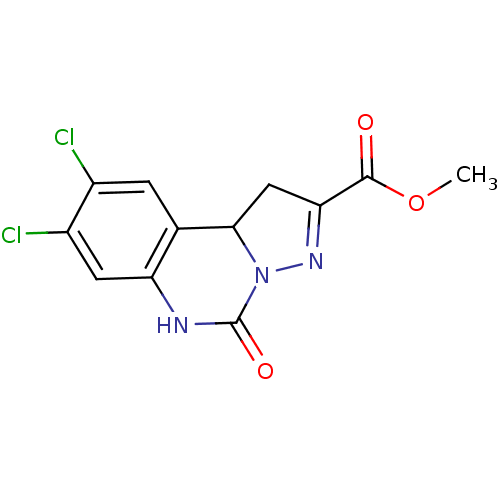
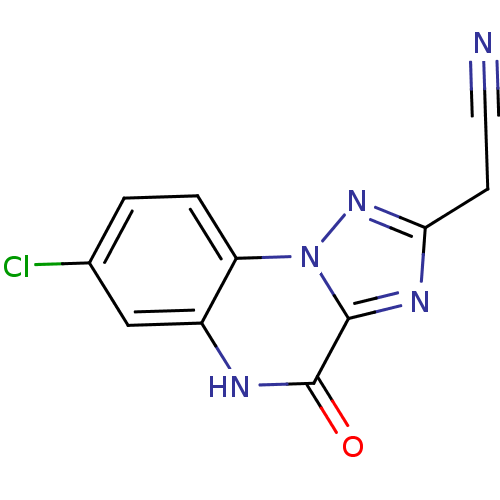
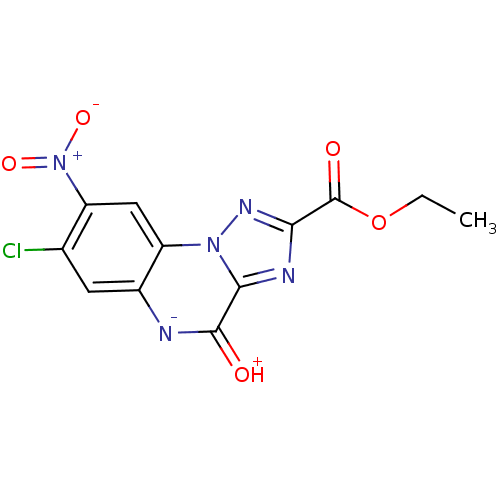
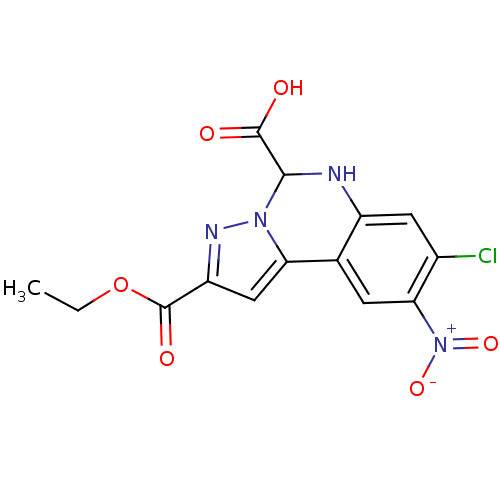
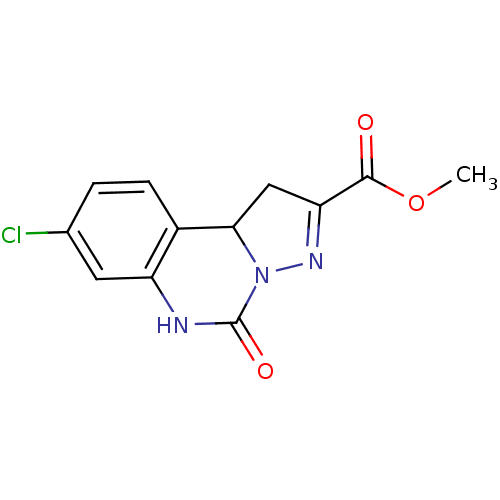
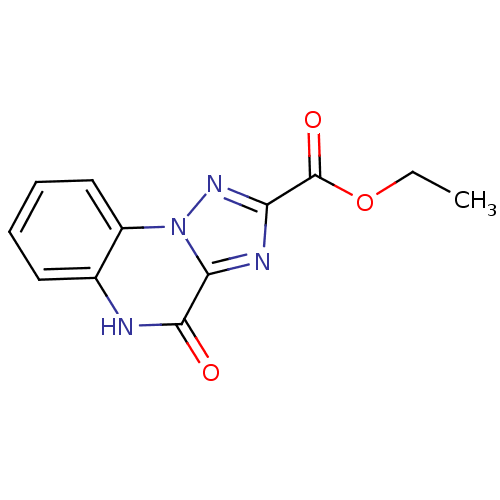
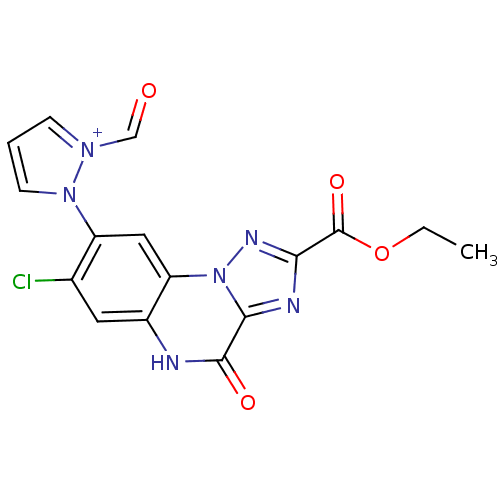
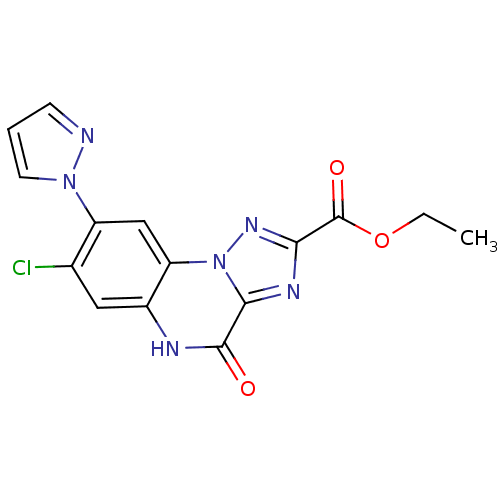
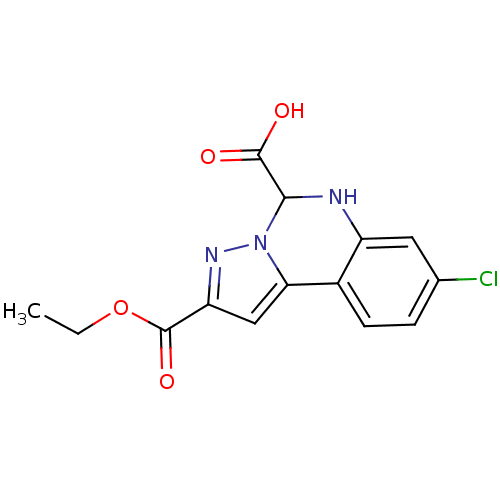
Report error Found 55 Enz. Inhib. hit(s) with all data for entry = 50013659
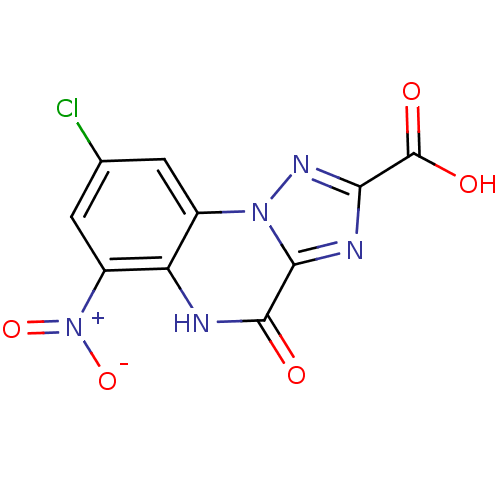
Affinity DataKi: 74nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
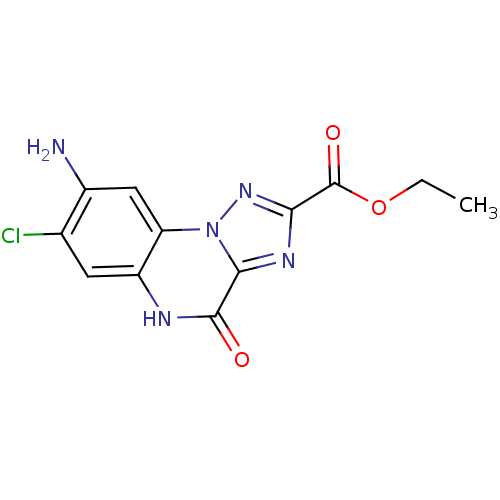
Affinity DataKi: 140nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
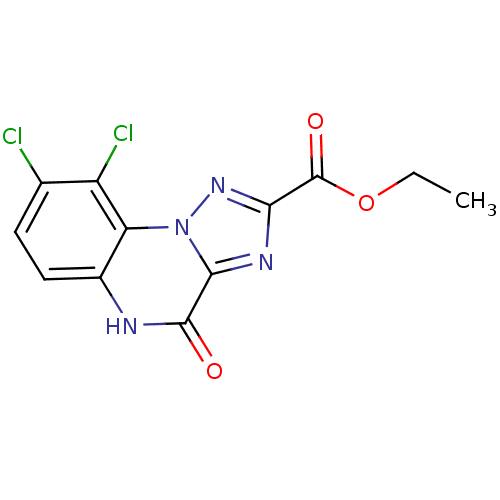
Affinity DataKi: 160nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
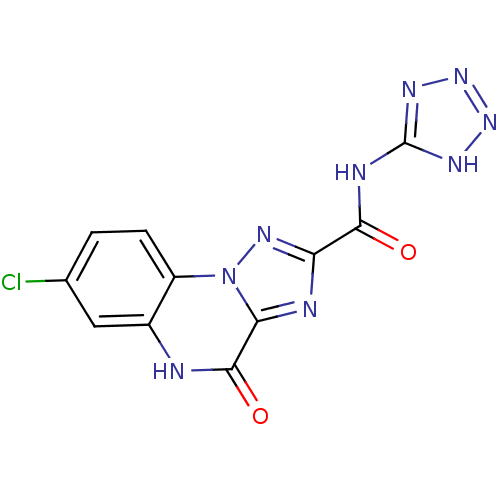
Affinity DataKi: 170nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 190nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 240nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 390nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 480nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 570nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 710nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 710nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 860nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 860nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 1.10E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 1.10E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 1.10E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 1.10E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 1.41E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 1.80E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 2.00E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 2.20E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 2.60E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 3.00E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 3.50E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 3.60E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 4.00E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 4.50E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 4.80E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 4.80E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 5.10E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 5.40E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 6.50E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 7.20E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 7.50E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 8.20E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 8.30E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 8.80E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 9.60E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 9.70E+3nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 1.02E+4nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 1.06E+4nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 1.06E+4nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 1.67E+4nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 1.90E+4nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 2.30E+4nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 2.65E+4nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 2.77E+4nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 2.80E+4nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 3.33E+4nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair
Affinity DataKi: 3.35E+4nMAssay Description:Binding selectivity towards N-methyl-D-aspartate glutamate receptor 1 was determined by using [3H]- glycine/NMDAMore data for this Ligand-Target Pair