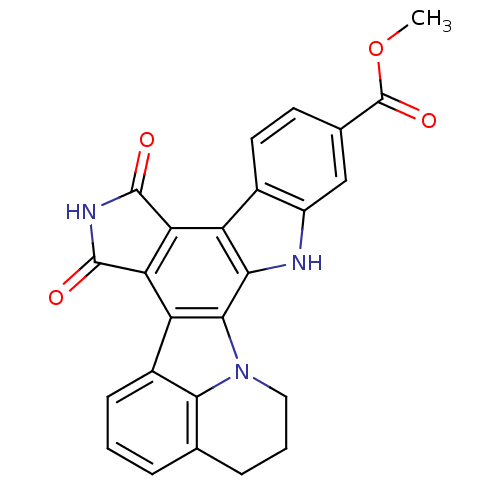
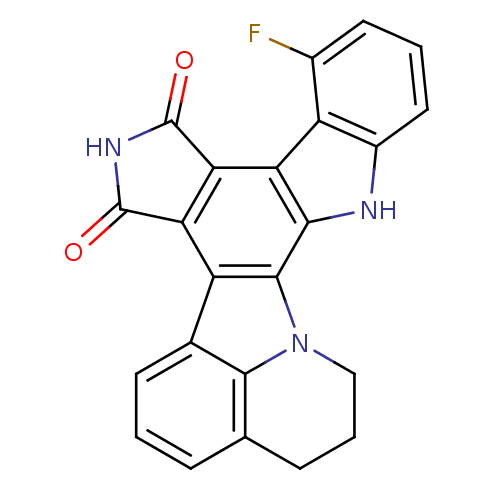
Report error Found 32 Enz. Inhib. hit(s) with all data for entry = 859
Affinity DataIC50: 50nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 65nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 75nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 75nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 79nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 87nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 90nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 90nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 94nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 110nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 110nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 120nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 160nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 170nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 240nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 290nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 420nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 590nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMT: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMT: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMT: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMT: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMT: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMT: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMT: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMT: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMpH: 7.0 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at room temperature with substrate, and test compounds in the presence of ATP/[gamma-33P]ATP....More data for this Ligand-Target Pair