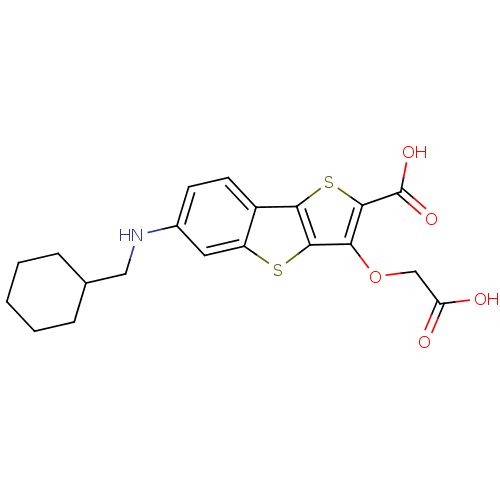
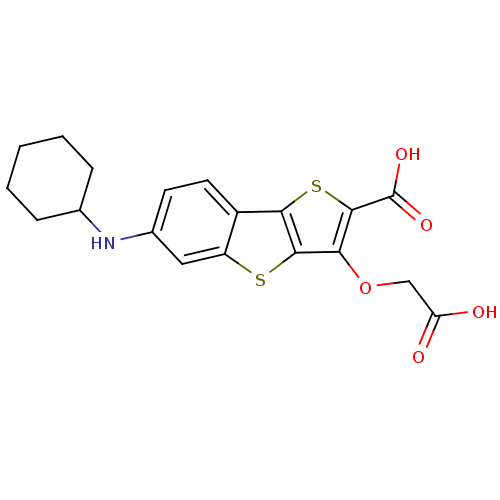
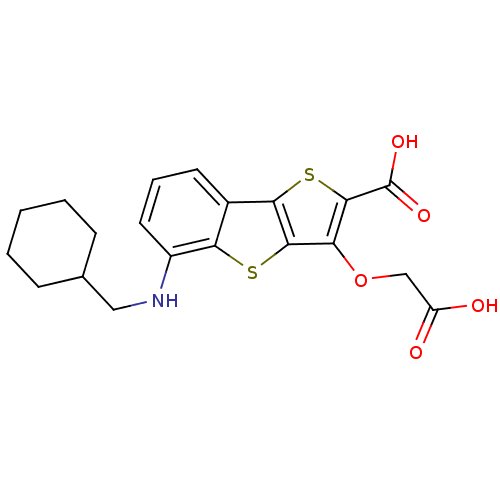
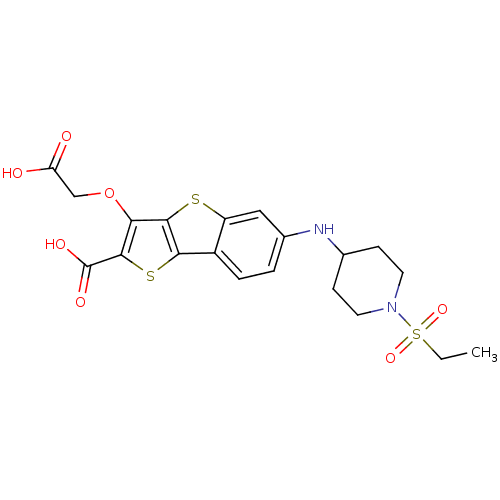
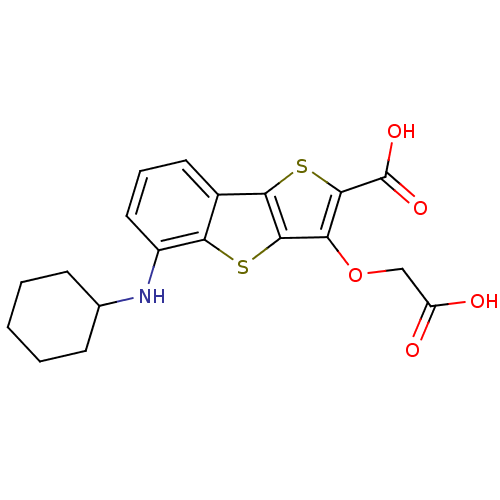
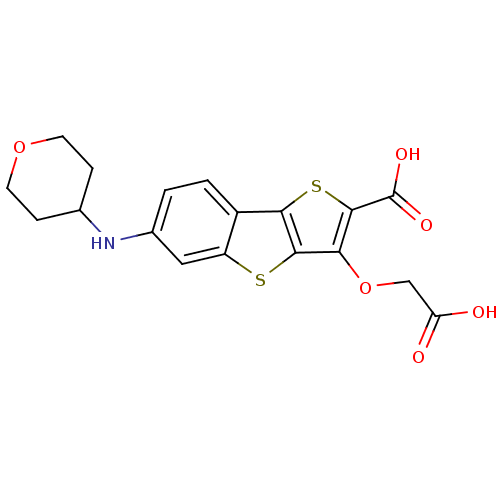
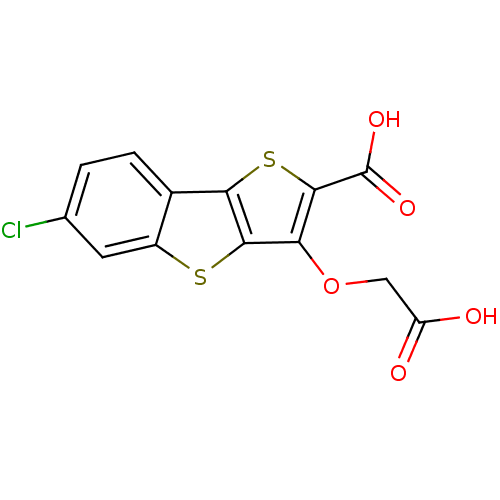
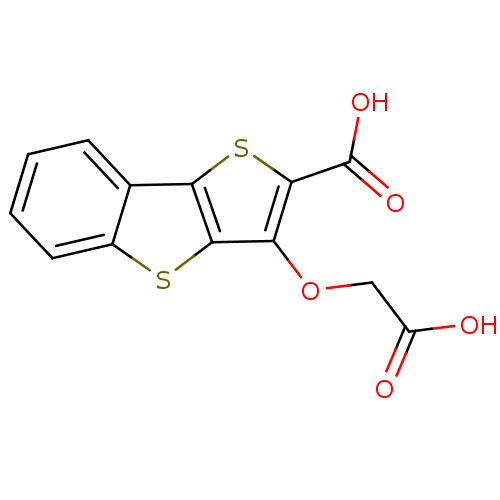
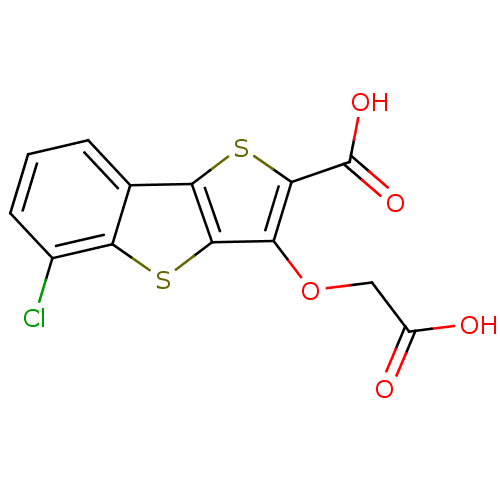
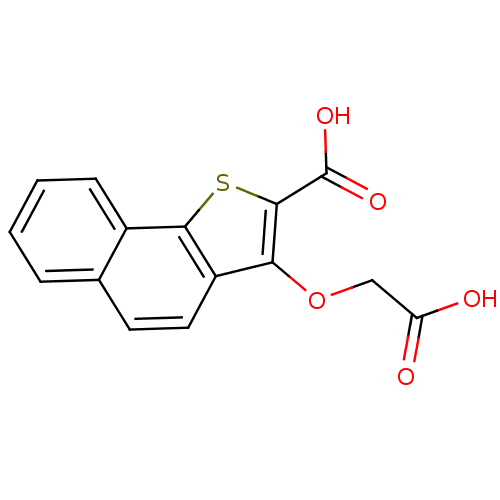
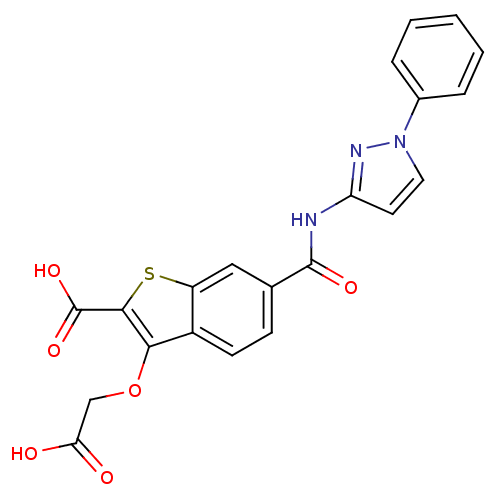
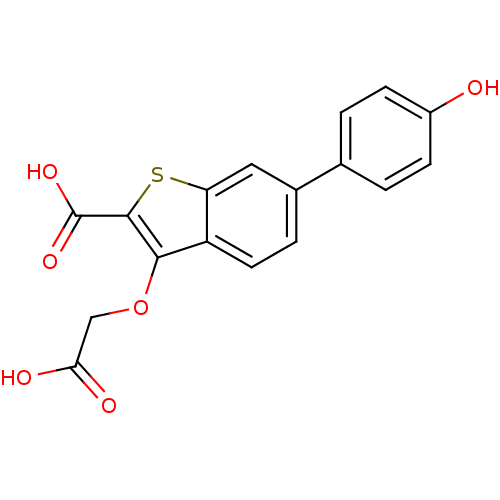
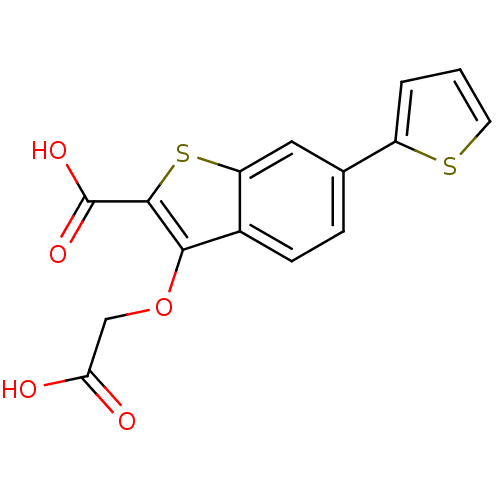
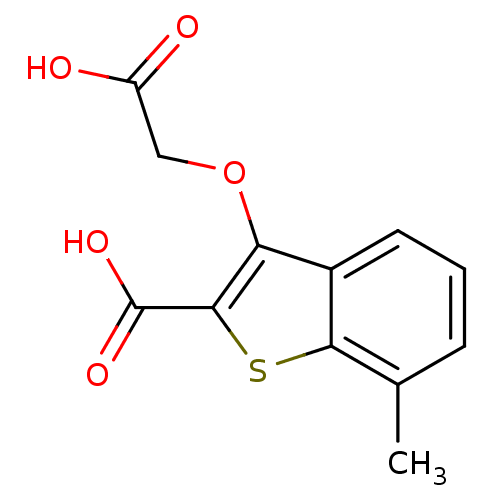
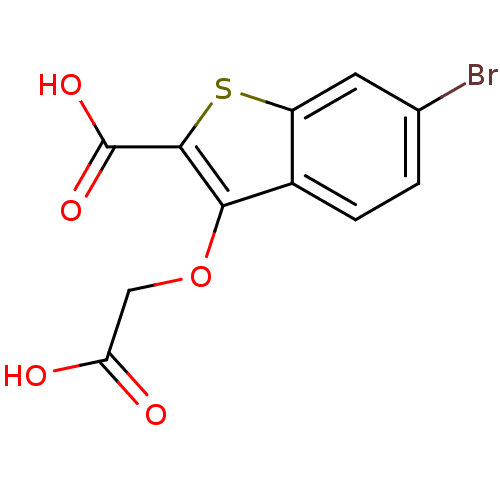
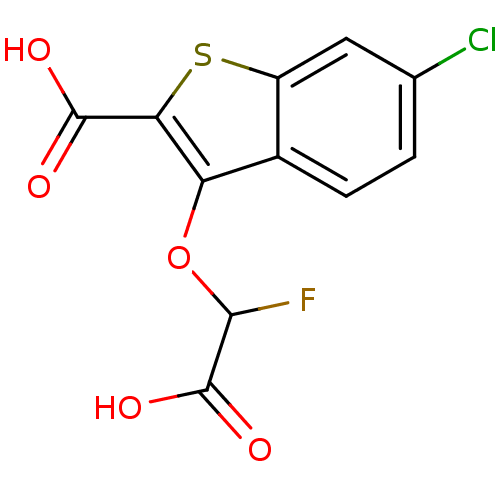
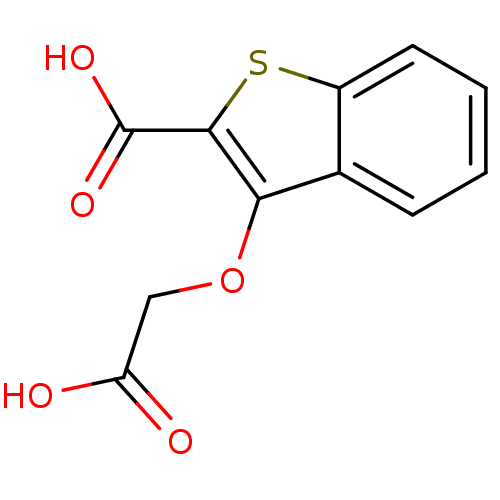
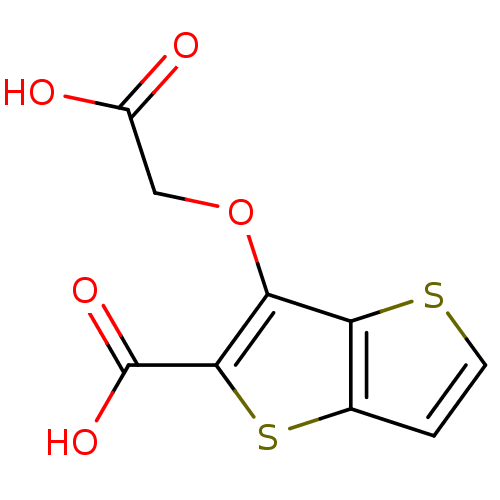
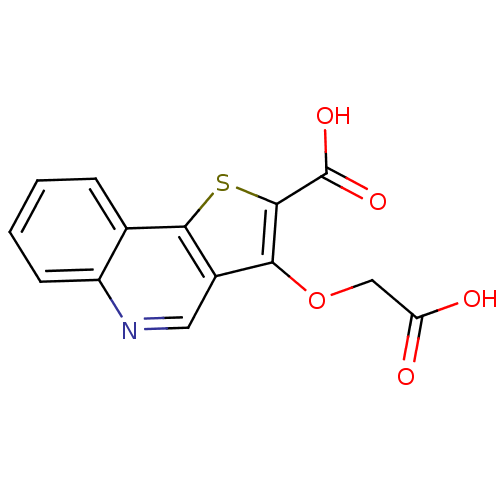
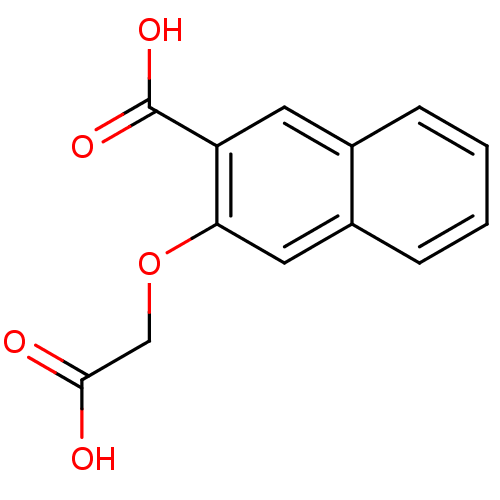
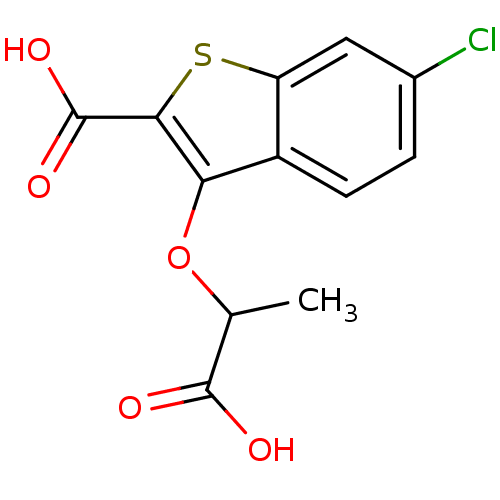
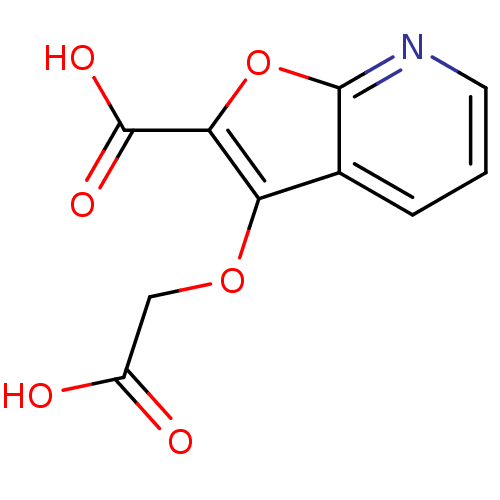
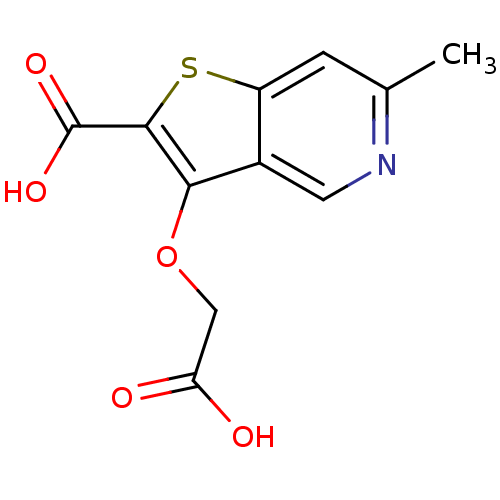
Report error Found 31 Enz. Inhib. hit(s) with all data for entry = 1806
Affinity DataKi: 370nM ΔG°: -36.7kJ/moleT: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 380nM ΔG°: -36.6kJ/moleT: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 680nM ΔG°: -35.2kJ/moleT: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 740nM ΔG°: -35.0kJ/moleT: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 920nM ΔG°: -34.5kJ/moleT: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 1.60E+3nM ΔG°: -33.1kJ/moleT: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 1.70E+3nM ΔG°: -32.9kJ/moleT: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 2.40E+3nM ΔG°: -32.1kJ/moleT: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 3.50E+3nM ΔG°: -31.1kJ/moleT: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 4.10E+3nM ΔG°: -30.7kJ/moleT: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 9.20E+3nM ΔG°: -28.7kJ/moleT: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 1.00E+4nM ΔG°: -28.5kJ/moleT: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 1.10E+4nM ΔG°: -28.3kJ/moleT: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+4nM ΔG°: -26.6kJ/molepH: 7.0 T: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 2.60E+4nM ΔG°: -25.9kJ/molepH: 7.0 T: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 3.00E+4nM ΔG°: -25.6kJ/molepH: 7.0 T: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 3.70E+4nM ΔG°: -25.3kJ/moleT: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 4.20E+4nM ΔG°: -24.7kJ/molepH: 7.0 T: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 5.20E+4nM ΔG°: -24.2kJ/molepH: 7.0 T: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 6.10E+4nM ΔG°: -23.8kJ/molepH: 7.0 T: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 7.70E+4nM ΔG°: -23.2kJ/molepH: 7.0 T: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 1.19E+5nM ΔG°: -22.4kJ/moleT: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 1.28E+5nM ΔG°: -22.0kJ/molepH: 7.0 T: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 1.60E+5nM ΔG°: -21.4kJ/molepH: 7.0 T: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 2.30E+5nM ΔG°: -20.6kJ/molepH: 7.4 T: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 2.80E+5nM ΔG°: -20.1kJ/molepH: 7.0 T: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: >5.00E+5nM ΔG°: >-18.8kJ/moleT: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: 8.60E+5nM ΔG°: -17.3kJ/molepH: 7.0 T: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: >2.50E+6nM ΔG°: >-14.7kJ/molepH: 7.0 T: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: >2.50E+6nM ΔG°: >-14.7kJ/molepH: 7.0 T: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
Affinity DataKi: >2.50E+6nM ΔG°: >-14.7kJ/molepH: 7.0 T: 2°CAssay Description:The enzymatic assay was carried out at room temperature in 96-well plates. The initial rate of PTPase-catalyzed hydrolysis of p-nitrophenol phosphate...More data for this Ligand-Target Pair
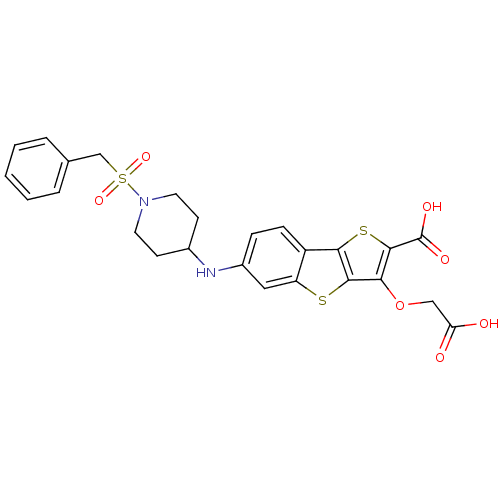
 3D Structure (crystal)
3D Structure (crystal)