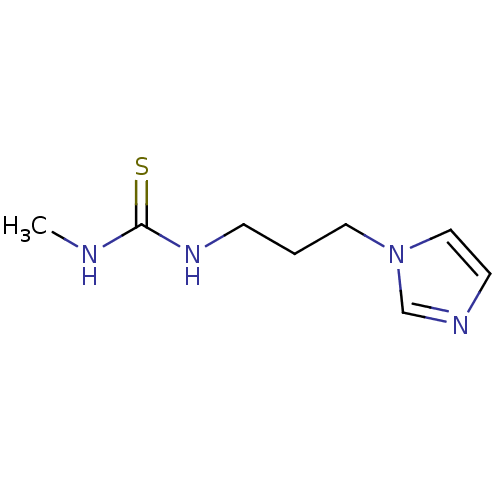
Report error Found 56 Enz. Inhib. hit(s) with all data for entry = 993
Affinity DataKi: 60nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 90nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 340nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 340nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 390nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 400nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 490nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 550nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 560nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 700nM ΔG°: -35.7kJ/molepH: 8.0 T: 2°CAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 700nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 750nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 760nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 890nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 970nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 990nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 1.12E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 1.24E+3nM ΔG°: -34.3kJ/molepH: 8.0 T: 2°CAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 1.55E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 1.57E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 1.66E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 1.79E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 1.86E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 2.03E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 2.14E+3nM ΔG°: -32.9kJ/molepH: 8.0 T: 2°CAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 2.22E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 2.33E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 2.68E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 2.78E+3nM ΔG°: -32.2kJ/molepH: 8.0 T: 2°CAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 2.79E+3nM ΔG°: -32.2kJ/molepH: 8.0 T: 2°CAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 3.35E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 3.51E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 3.66E+3nM ΔG°: -31.6kJ/molepH: 8.0 T: 2°CAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 3.73E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 4.48E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 4.48E+3nM ΔG°: -31.0kJ/molepH: 8.0 T: 2°CAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 4.65E+3nM ΔG°: -30.9kJ/molepH: 8.0 T: 2°CAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 4.70E+3nM ΔG°: -30.9kJ/molepH: 8.0 T: 2°CAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 4.73E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 4.83E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 4.88E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 5.66E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 5.67E+3nM ΔG°: -30.4kJ/molepH: 8.0 T: 2°CAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 7.00E+3nM ΔG°: -29.9kJ/molepH: 8.0 T: 2°CAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 7.33E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 7.34E+3nMAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 7.70E+3nM ΔG°: -29.7kJ/molepH: 8.0 T: 2°CAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 1.13E+4nM ΔG°: -28.7kJ/molepH: 8.0 T: 2°CAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
Affinity DataKi: 1.30E+4nM ΔG°: -28.4kJ/molepH: 8.0 T: 2°CAssay Description:QC activity was evaluated fluorometrically using Gln-AMC as substrate, and pyroglutamyl peptidase as the auxiliary enzyme. After conversion of Gln-AM...More data for this Ligand-Target Pair
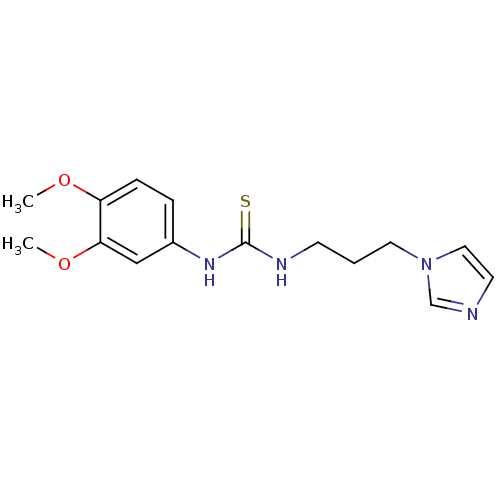
 3D Structure (crystal)
3D Structure (crystal)