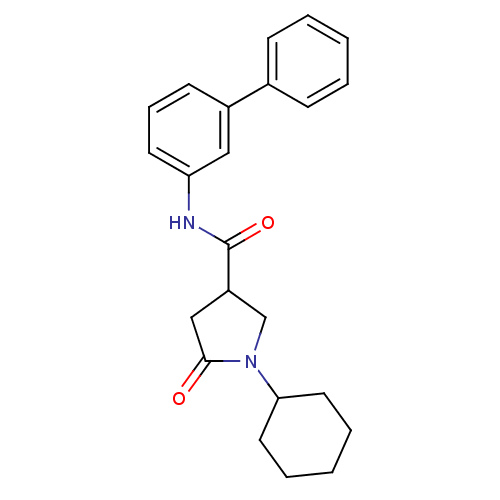
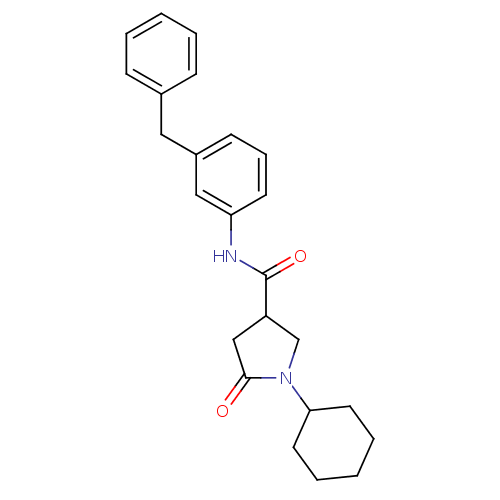
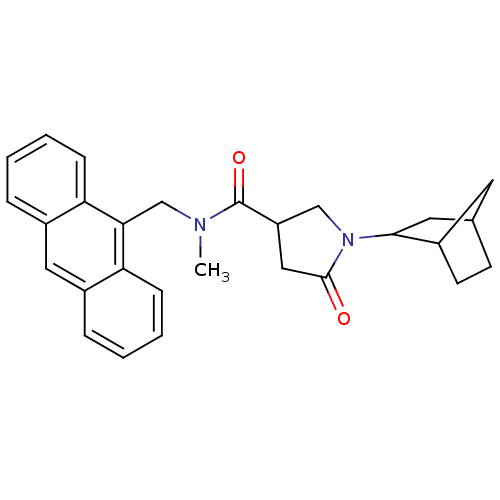
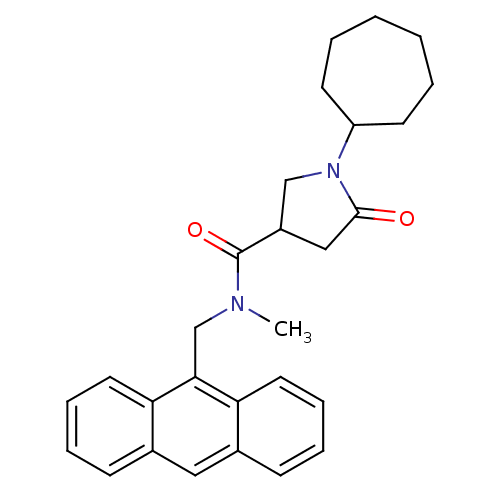
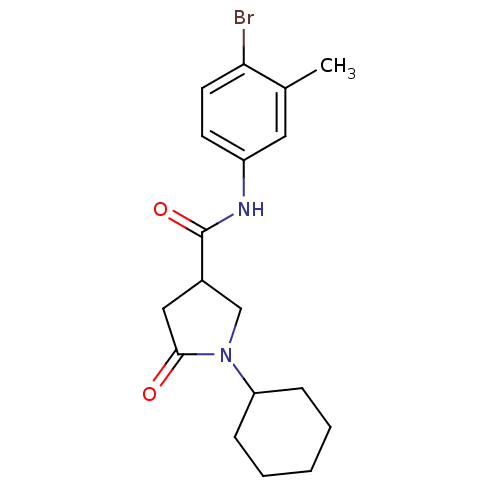
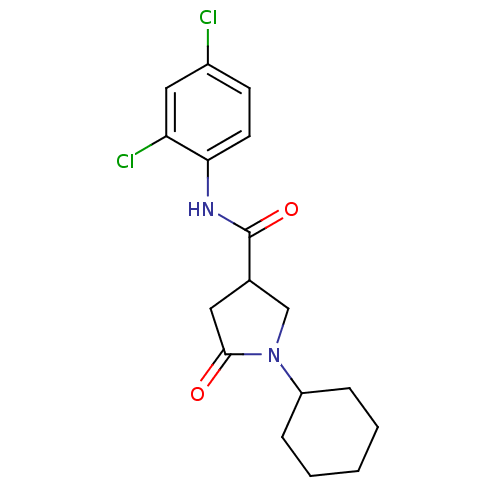
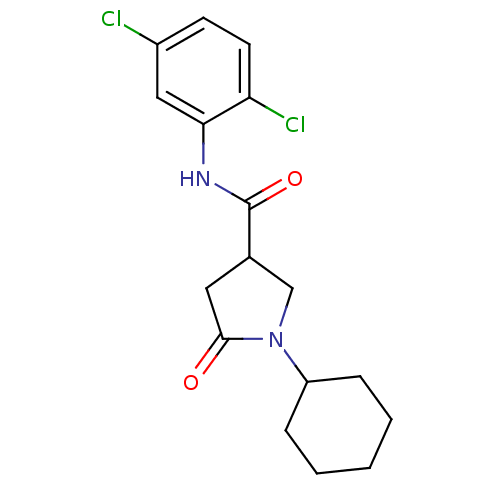
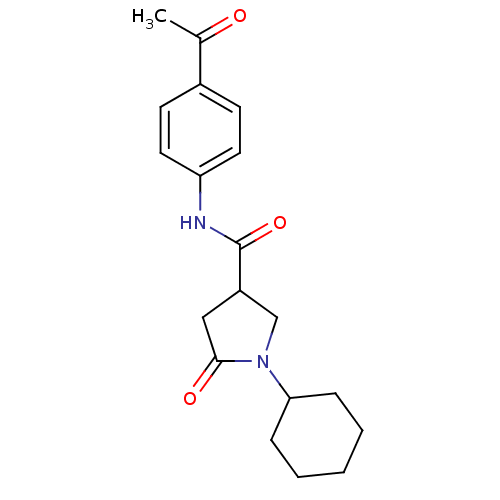
Report error Found 66 Enz. Inhib. hit(s) with all data for entry = 1973
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
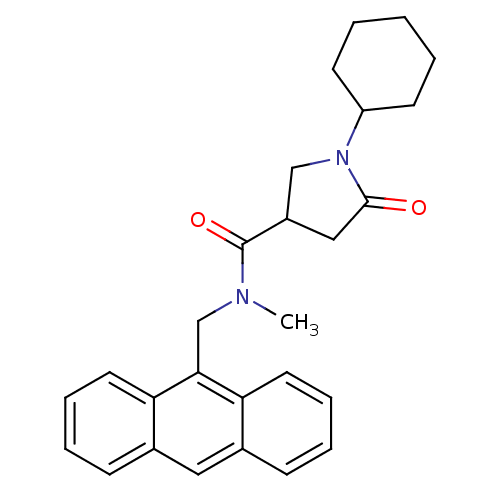
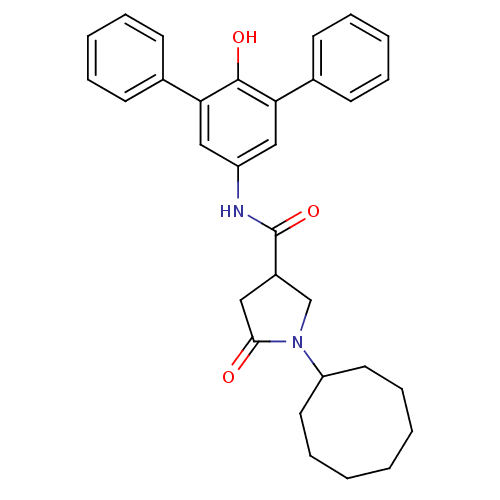
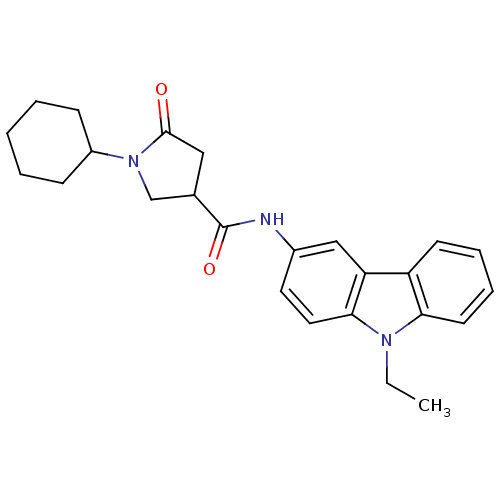
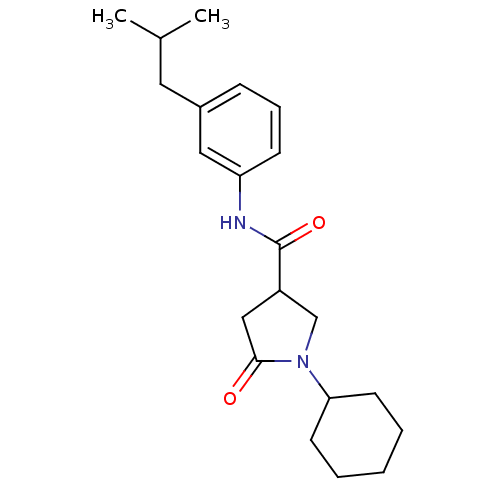
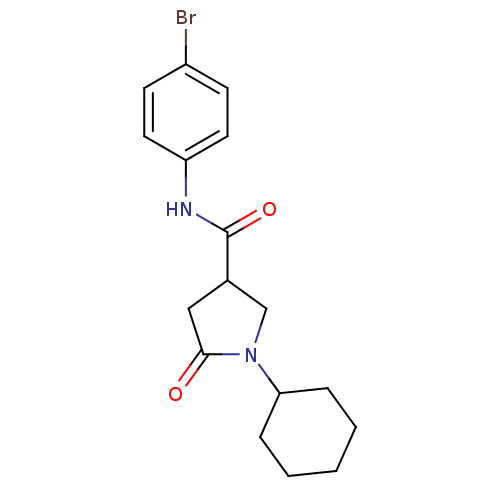
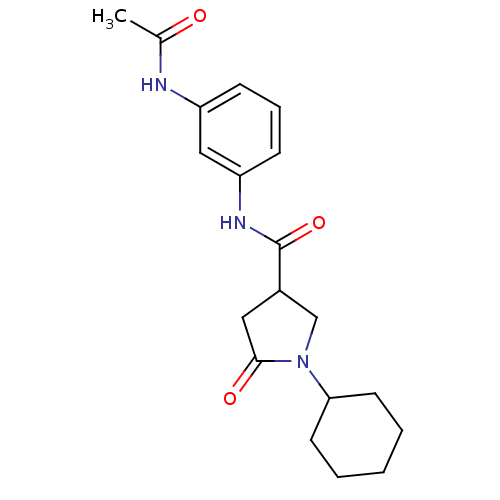
Affinity DataIC50: 140nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
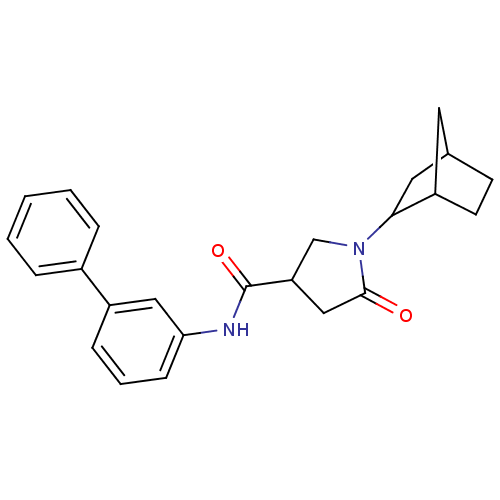
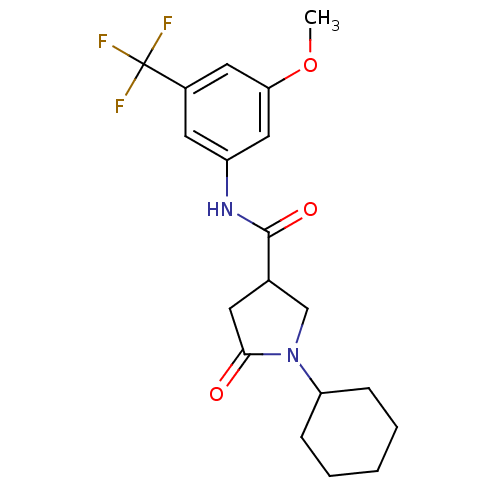
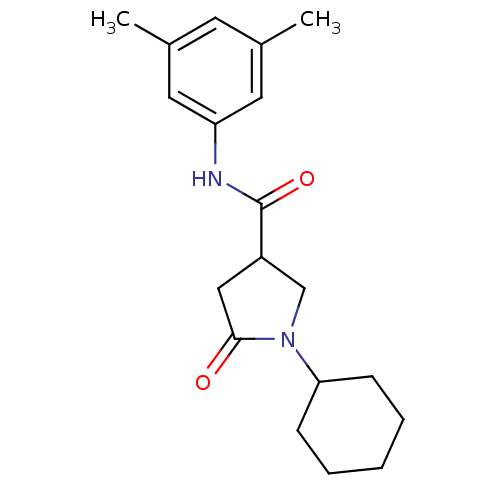
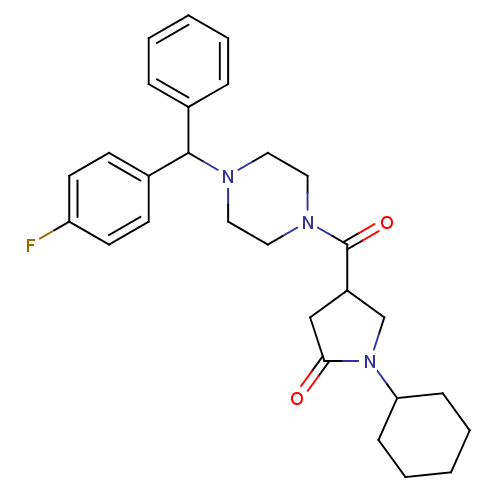
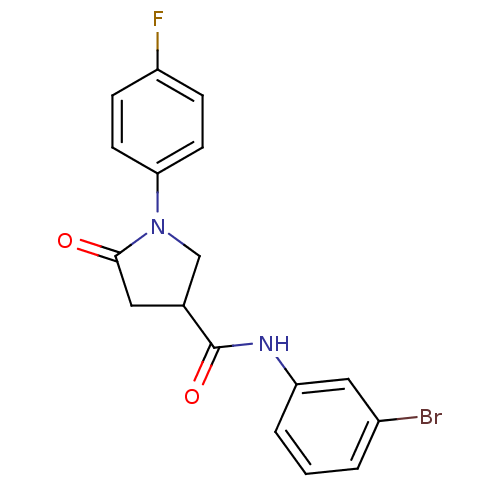
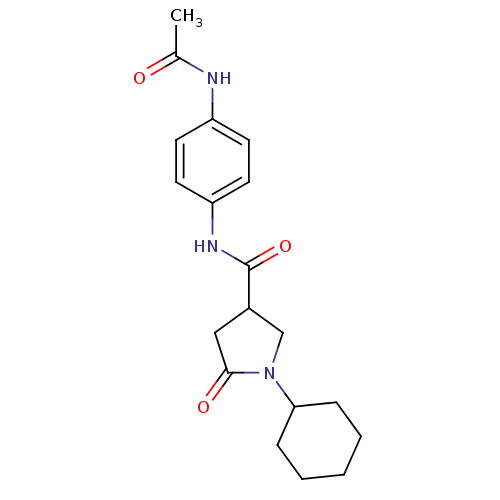
Affinity DataIC50: 320nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
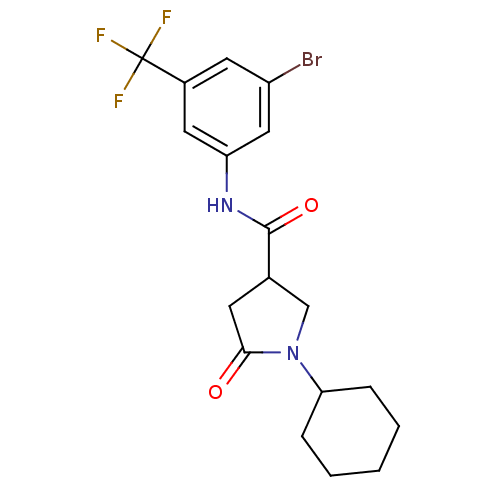
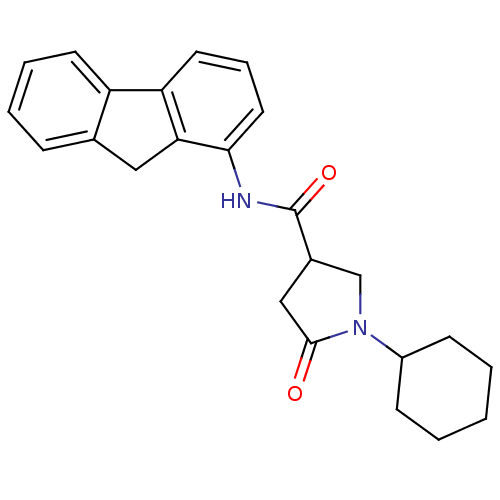
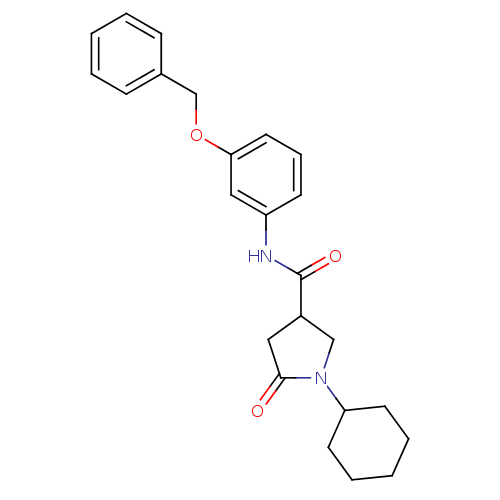
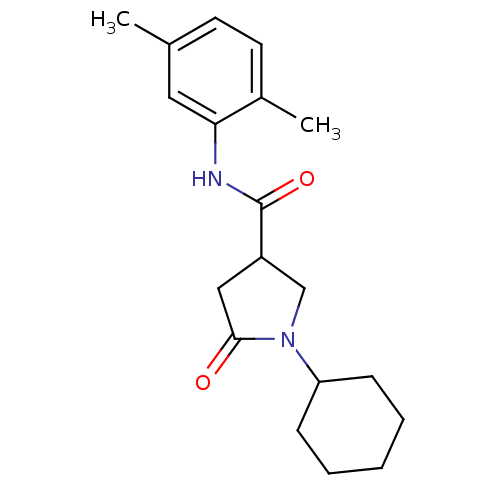
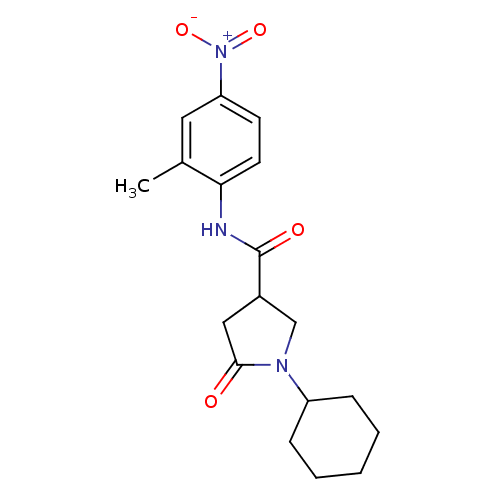
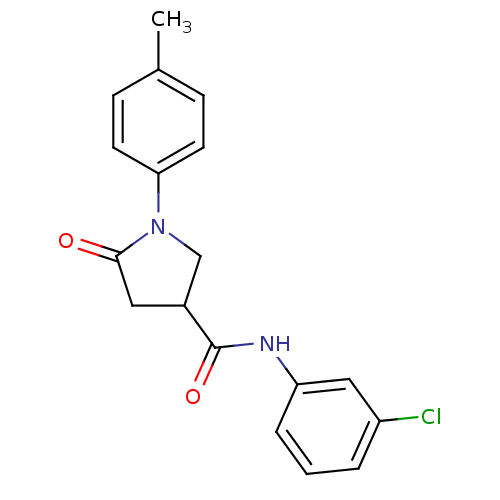
Affinity DataIC50: 360nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
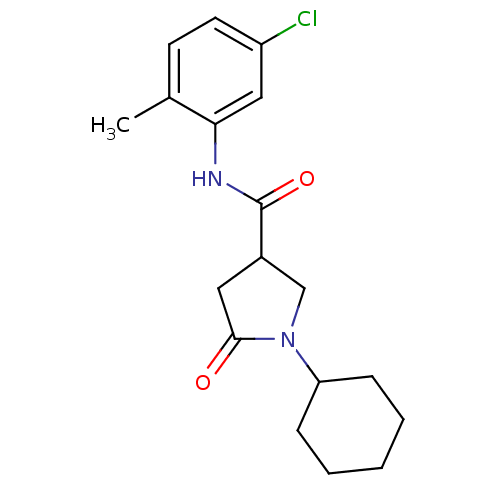
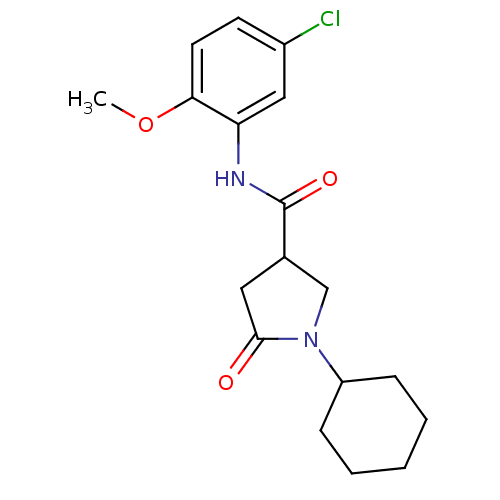
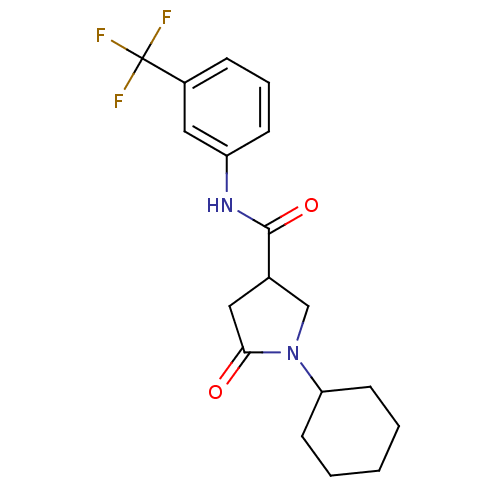
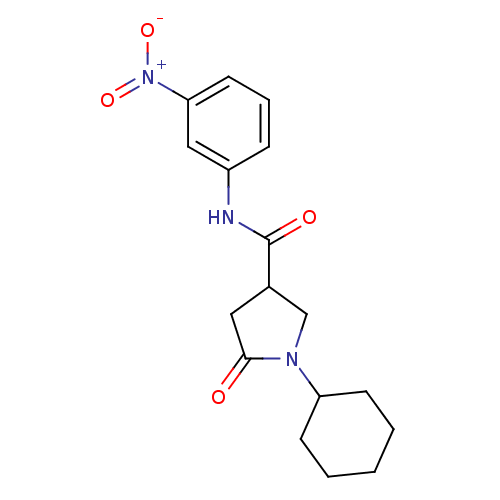
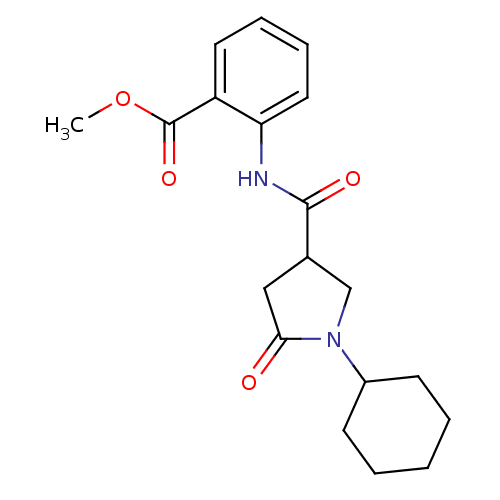
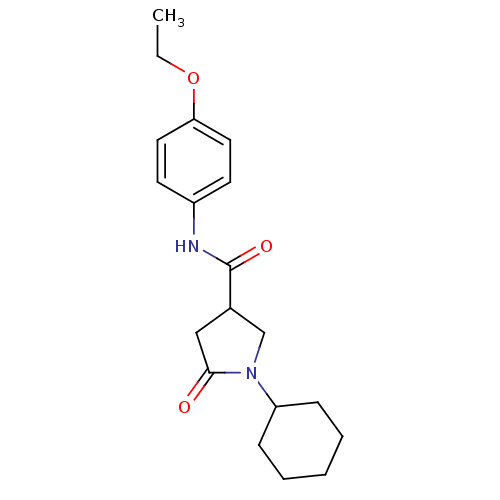
Affinity DataIC50: 390nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 390nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 410nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 460nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 620nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 750nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 845nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 850nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 890nMpH: 6.8 T: 2°CAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 970nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 1.29E+3nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 1.30E+3nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 1.35E+3nMpH: 6.8 T: 2°CAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 1.39E+3nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 1.49E+3nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 1.60E+3nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 2.57E+3nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 3.14E+3nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 3.39E+3nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 3.51E+3nMpH: 6.8 T: 2°CAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 3.67E+3nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 3.94E+3nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 4.47E+3nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 5.18E+3nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 5.51E+3nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 5.55E+3nMpH: 6.8 T: 2°CAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 6.41E+3nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 1.01E+4nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 1.06E+4nMpH: 6.8 T: 2°CAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 1.07E+4nMpH: 6.8 T: 2°CAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 1.36E+4nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 1.45E+4nMpH: 6.8 T: 2°CAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 1.48E+4nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 1.68E+4nMpH: 6.8 T: 2°CAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 2.31E+4nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 2.80E+4nMpH: 6.8 T: 2°CAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 2.92E+4nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 3.14E+4nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 3.49E+4nMpH: 6.8 T: 2°CAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 3.74E+4nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 5.60E+4nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 5.65E+4nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 7.36E+4nMpH: 6.8 T: 2°CAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 7.50E+4nMpH: 6.8 T: 2°CAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 1.00E+5nMpH: 6.8 T: 2°CAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 1.00E+5nMAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
TargetEnoyl-[acyl-carrier-protein] reductase [NADH](Mycobacterium tuberculosis (strain ATCC 25618 / H3...)
University of California San Francisco
University of California San Francisco
Affinity DataIC50: 1.00E+5nMpH: 6.8 T: 2°CAssay Description:The assay measured the NADH-dependent catalysis of an octenoyl-CoA substrate as a decrease in 340 nm absorbance resulting from conversion of NADH to ...More data for this Ligand-Target Pair
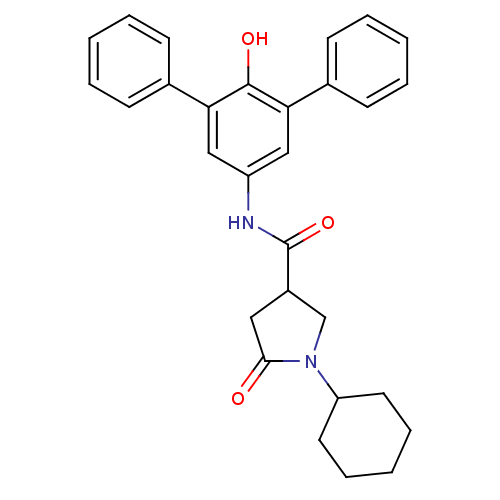
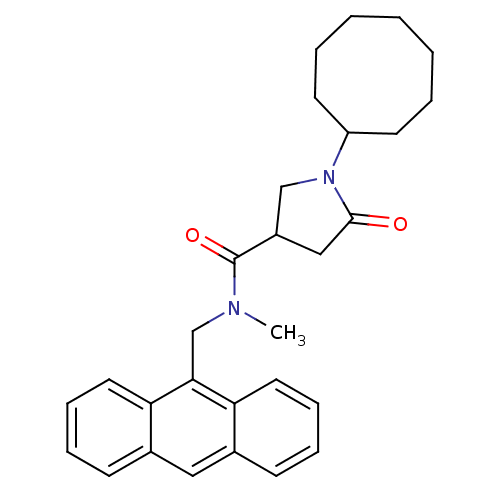
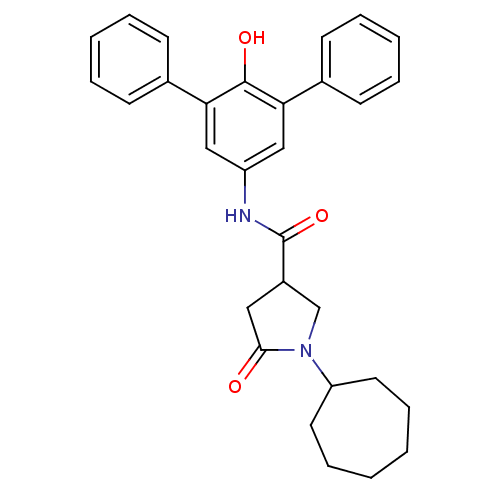

 3D Structure (crystal)
3D Structure (crystal)