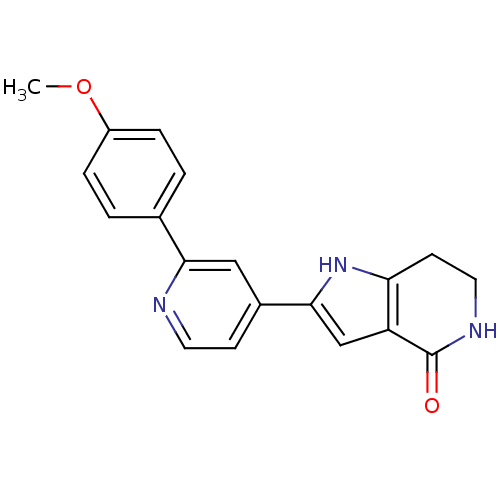
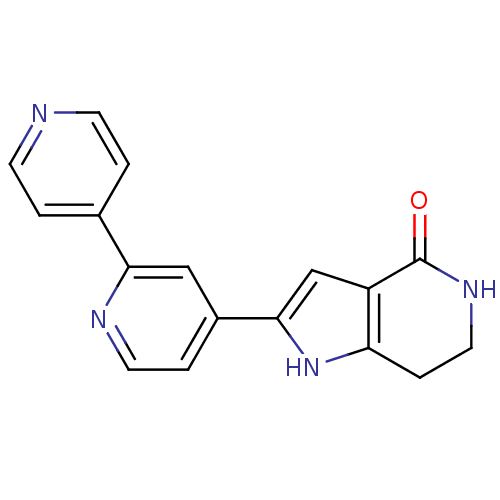
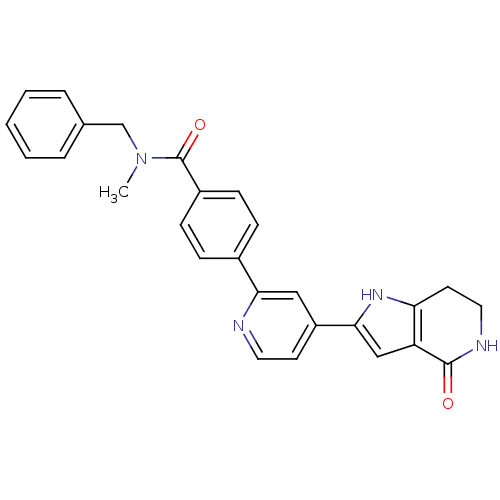
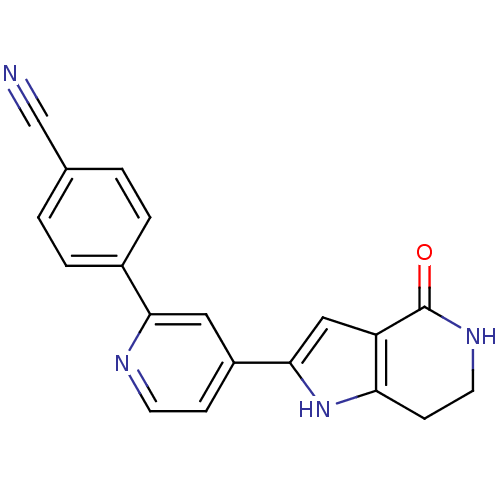
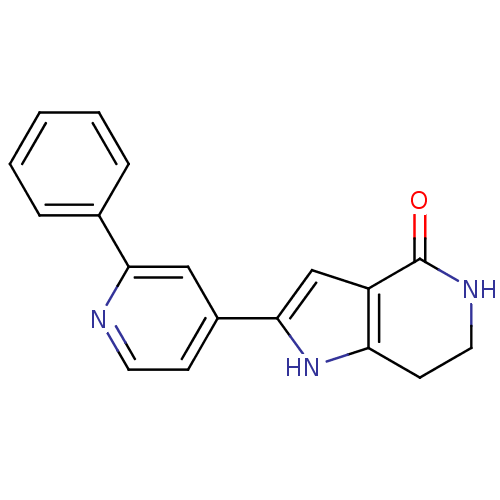
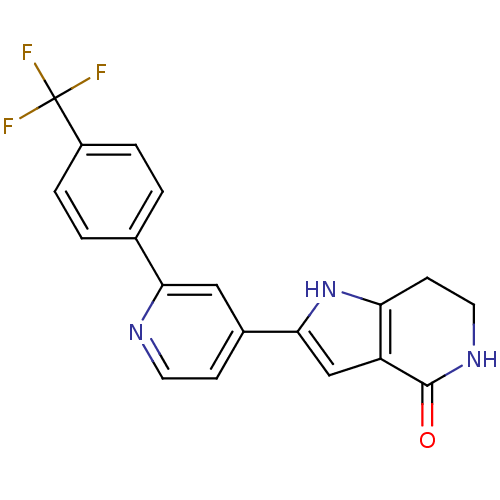
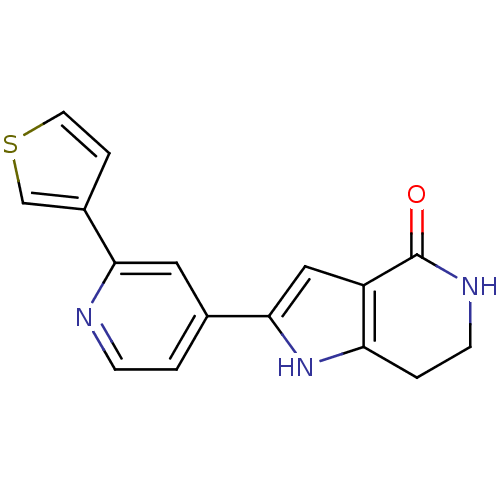
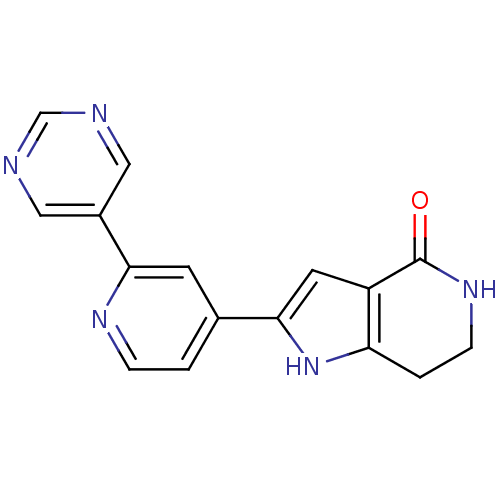
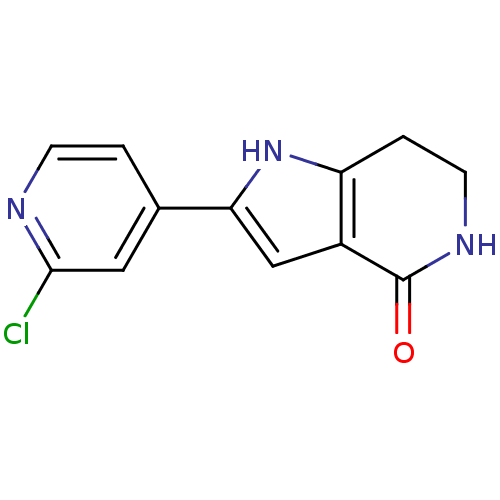
Report error Found 53 Enz. Inhib. hit(s) with all data for entry = 3270
Affinity DataIC50: 8nMAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 8.5nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 17nMAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 21nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 25nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 32nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 37nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 41nMAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 46nMAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 48nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 50nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 51nMAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 52nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 55nMAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 56nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 56nMAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 62nMAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 66nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 71nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 76nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 81nMpH: 7.5 T: 2°CAssay Description:Compounds were evaluated as inhibitors of kinase by measuring their effect on kinase induced phosphorylation of substrate. IC50 values were reported ...More data for this Ligand-Target Pair
Affinity DataIC50: 83nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 126nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 140nMpH: 7.5 T: 2°CAssay Description:Compounds were evaluated as inhibitors of kinase by measuring their effect on kinase induced phosphorylation of substrate. IC50 values were reported ...More data for this Ligand-Target Pair
Affinity DataIC50: 171nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 210nMpH: 7.5 T: 2°CAssay Description:Compounds were evaluated as inhibitors of kinase by measuring their effect on kinase induced phosphorylation of substrate. IC50 values were reported ...More data for this Ligand-Target Pair
Affinity DataIC50: 210nMpH: 7.5 T: 2°CAssay Description:Compounds were evaluated as inhibitors of kinase by measuring their effect on kinase induced phosphorylation of substrate. IC50 values were reported ...More data for this Ligand-Target Pair
Affinity DataIC50: 410nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 500nMpH: 7.5 T: 2°CAssay Description:Compounds were evaluated as inhibitors of kinase by measuring their effect on kinase induced phosphorylation of substrate. IC50 values were reported ...More data for this Ligand-Target Pair
Affinity DataIC50: 606nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 608nMpH: 7.5 T: 2°CAssay Description:The phosphorylation of HSP27 peptide by MAPKAPK2 was measured using an anion exchange resin capture assay method. The reaction was carried out in rea...More data for this Ligand-Target Pair
Affinity DataIC50: 660nMpH: 7.5 T: 2°CAssay Description:Compounds were evaluated as inhibitors of kinase by measuring their effect on kinase induced phosphorylation of substrate. IC50 values were reported ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMpH: 7.5 T: 2°CAssay Description:Compounds were evaluated as inhibitors of kinase by measuring their effect on kinase induced phosphorylation of substrate. IC50 values were reported ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.34E+3nMAssay Description:Compounds were evaluated as inhibitors of kinase by measuring their effect on kinase induced phosphorylation of substrate. IC50 values were reported ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.67E+3nMAssay Description:Compounds were evaluated as inhibitors of kinase by measuring their effect on kinase induced phosphorylation of substrate. IC50 values were reported ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.01E+3nMAssay Description:Compounds were evaluated as inhibitors of kinase by measuring their effect on kinase induced phosphorylation of substrate. IC50 values were reported ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.44E+3nMpH: 7.5 T: 2°CAssay Description:Compounds were evaluated as inhibitors of kinase by measuring their effect on kinase induced phosphorylation of substrate. IC50 values were reported ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.60E+3nMpH: 7.5 T: 2°CAssay Description:Compounds were evaluated as inhibitors of kinase by measuring their effect on kinase induced phosphorylation of substrate. IC50 values were reported ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.70E+3nMAssay Description:Compounds were evaluated as inhibitors of kinase by measuring their effect on kinase induced phosphorylation of substrate. IC50 values were reported ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Compounds were evaluated as inhibitors of kinase by measuring their effect on kinase induced phosphorylation of substrate. IC50 values were reported ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+4nMAssay Description:Compounds were evaluated as inhibitors of kinase by measuring their effect on kinase induced phosphorylation of substrate. IC50 values were reported ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.77E+4nMpH: 7.5 T: 2°CAssay Description:Compounds were evaluated as inhibitors of kinase by measuring their effect on kinase induced phosphorylation of substrate. IC50 values were reported ...More data for this Ligand-Target Pair
Affinity DataIC50: 5.80E+4nMAssay Description:Compounds were evaluated as inhibitors of kinase by measuring their effect on kinase induced phosphorylation of substrate. IC50 values were reported ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.46E+5nMAssay Description:Compounds were evaluated as inhibitors of kinase by measuring their effect on kinase induced phosphorylation of substrate. IC50 values were reported ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.48E+5nMAssay Description:Compounds were evaluated as inhibitors of kinase by measuring their effect on kinase induced phosphorylation of substrate. IC50 values were reported ...More data for this Ligand-Target Pair