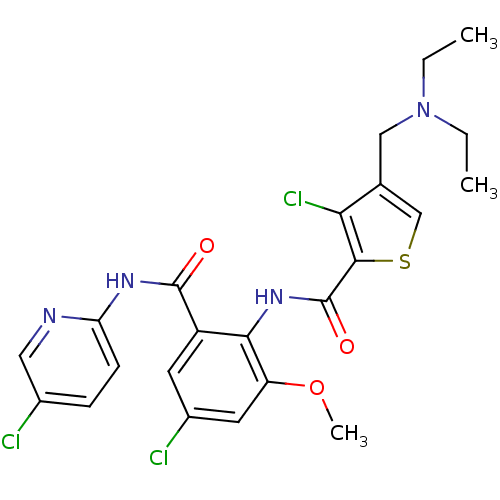
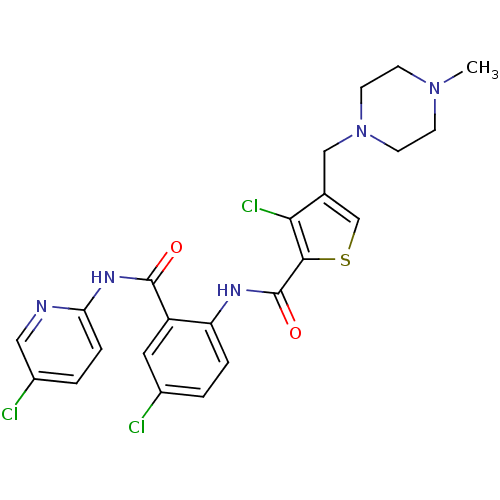
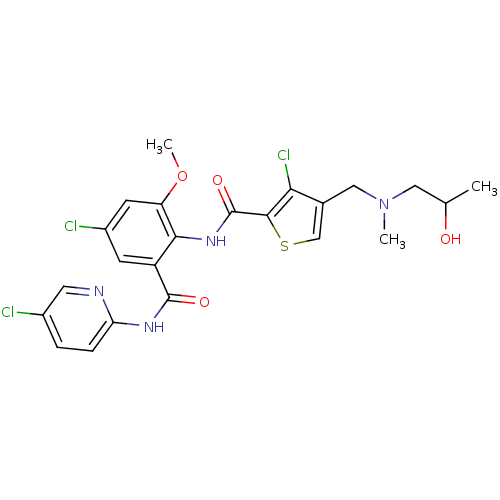
Report error Found 118 Enz. Inhib. hit(s) with all data for entry = 2117
Affinity DataKi: 0.00400nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.00400nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.00400nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.00500nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.00500nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.00600nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.00700nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.00800nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0100nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0100nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0100nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0100nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0100nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0100nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0100nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0120nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0120nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0190nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0200nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0200nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0220nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0220nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0240nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0240nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0260nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0260nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0310nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0340nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0360nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0450nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0500nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0590nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.0600nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.120nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.120nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.150nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.160nM ΔG°: -55.3kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.180nM ΔG°: -55.1kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.210nM ΔG°: -54.7kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.220nM ΔG°: -54.6kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.240nM ΔG°: -54.4kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.260nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.260nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.290nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.310nMAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.320nM ΔG°: -53.6kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.360nM ΔG°: -53.4kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.400nM ΔG°: -53.1kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.400nM ΔG°: -53.1kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
Affinity DataKi: 0.430nM ΔG°: -52.9kJ/molepH: 7.5 T: 2°CAssay Description:The enzyme activities were determined kinetically as the initial rate of cleavage of a peptide p-nitroanilide. Km for enzyme and substrate was determ...More data for this Ligand-Target Pair
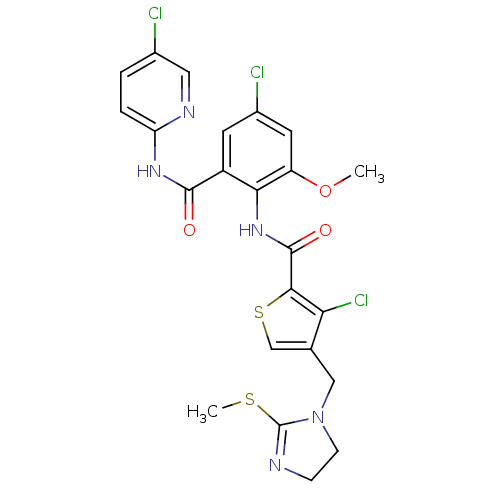
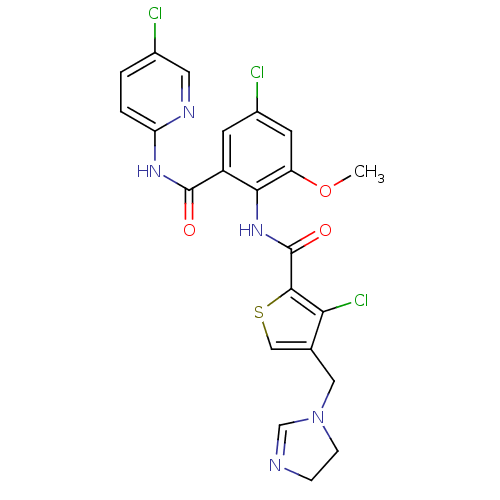
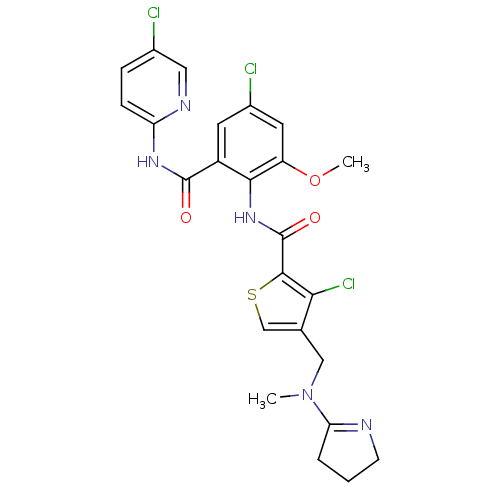
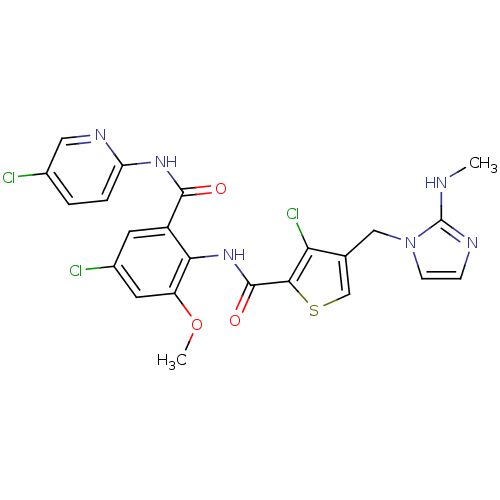
 3D Structure (crystal)
3D Structure (crystal)