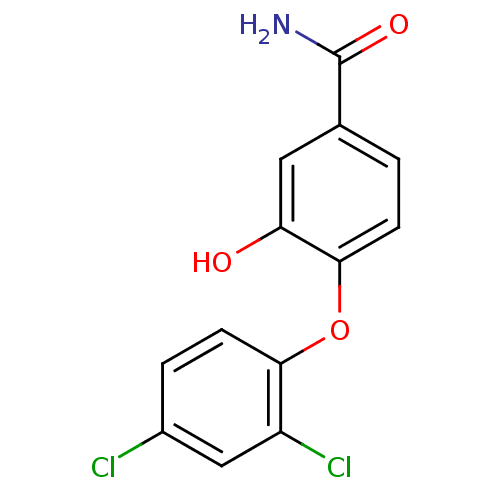
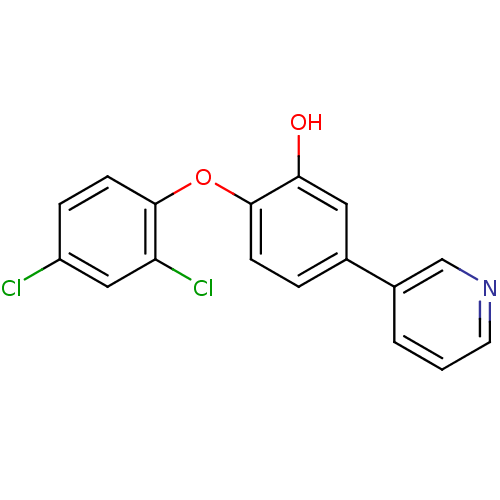
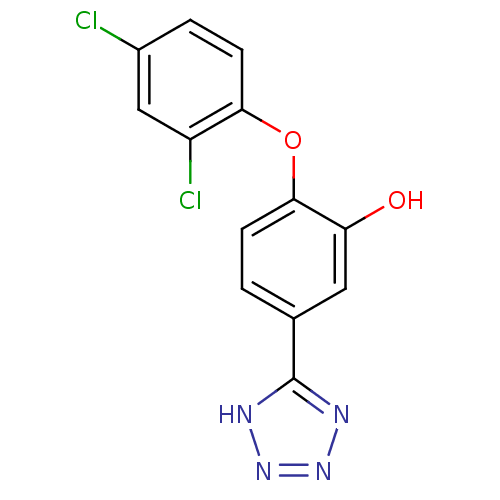
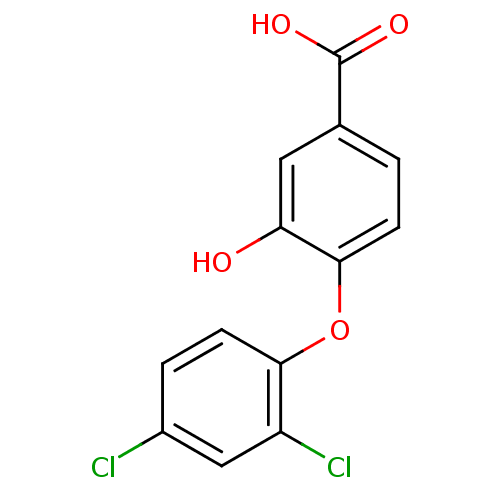
Report error Found 28 Enz. Inhib. hit(s) with all data for entry = 2858
Affinity DataIC50: 38nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 49nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 71nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 73nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 76nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 110nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 120nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 140nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 180nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 190nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 200nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 210nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 230nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 290nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 410nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 440nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 440nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 480nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 530nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 530nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 640nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 700nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 840nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+3nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.10E+4nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+4nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMpH: 7.9 T: 2°CAssay Description:PfENR inhibition assays were carried out on a Cary 100 Bio Spectrophotometer or POLARstar Optima by monitoring oxidation of NADH at 340 nm. The IC50 ...More data for this Ligand-Target Pair
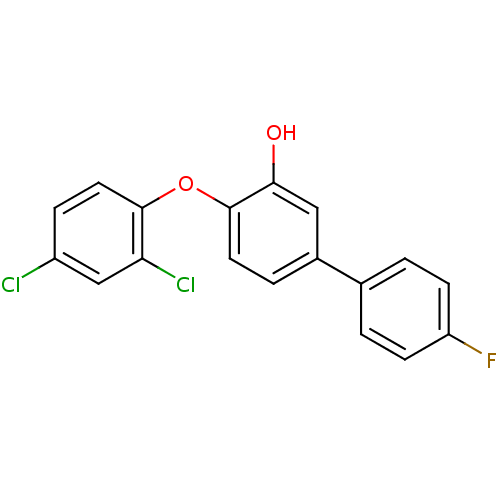
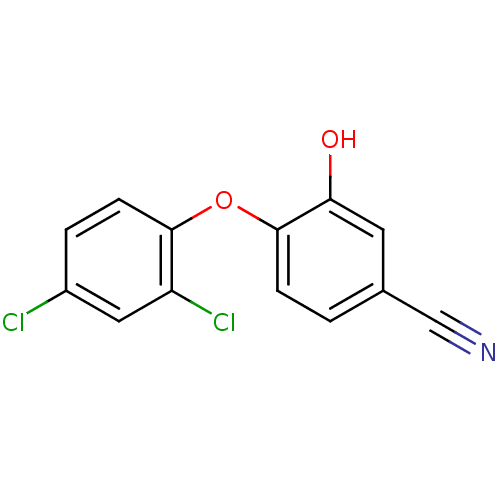
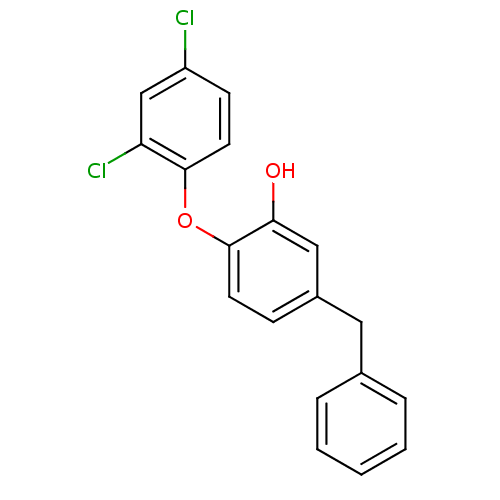
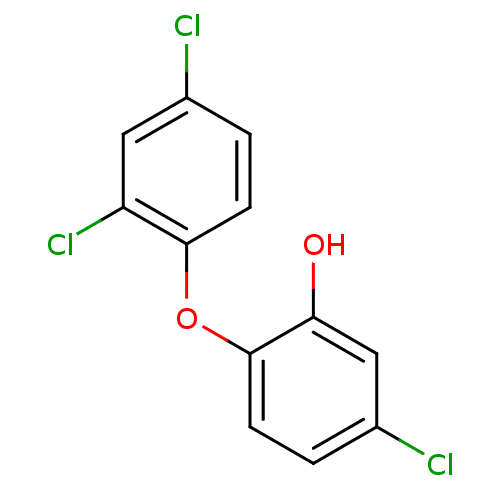
 3D Structure (crystal)
3D Structure (crystal)