Report error Found 27 Enz. Inhib. hit(s) with all data for entry = 50020638
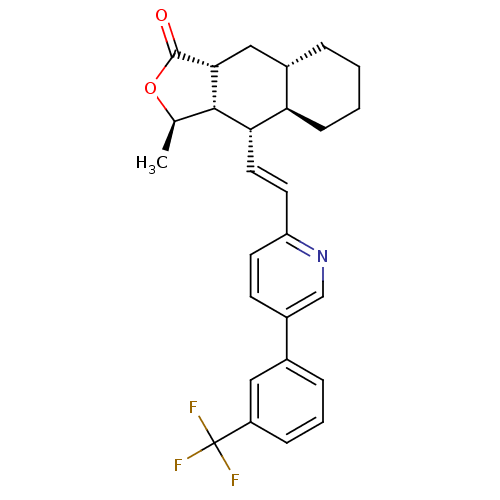
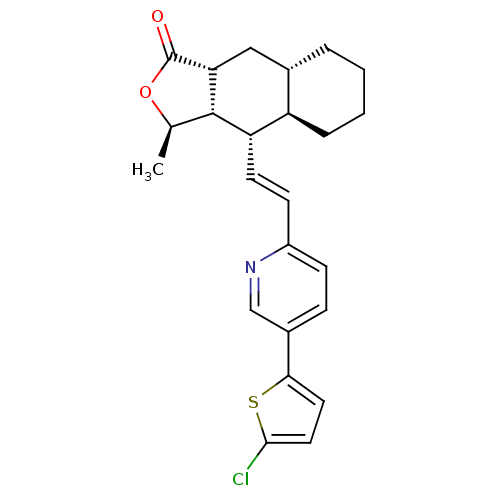
Affinity DataIC50: 11nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
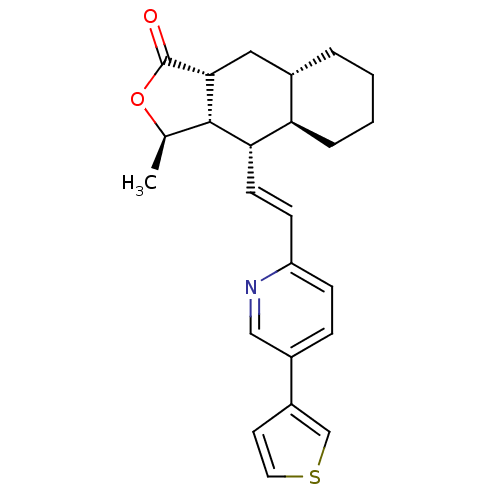
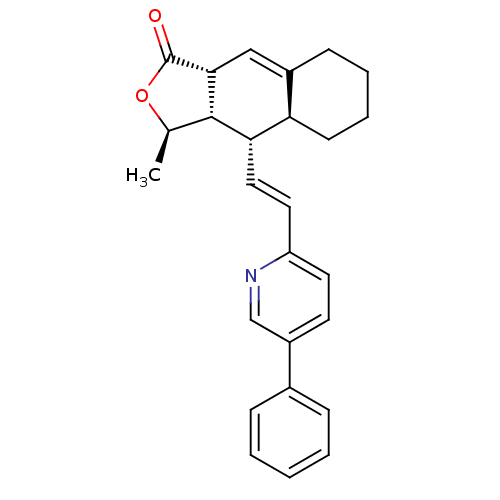
Affinity DataIC50: 11nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
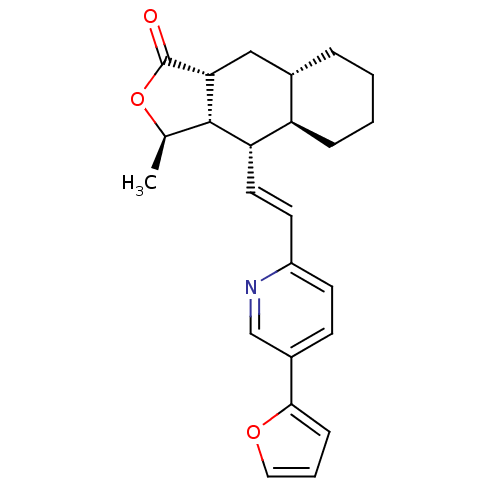
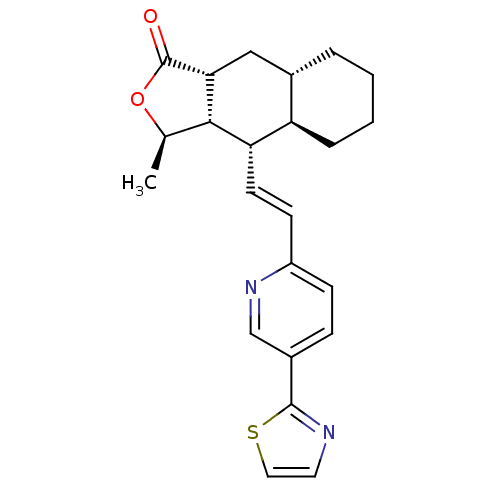
Affinity DataIC50: 12nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
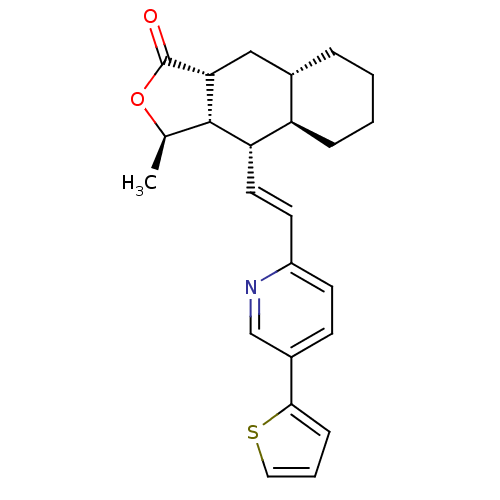
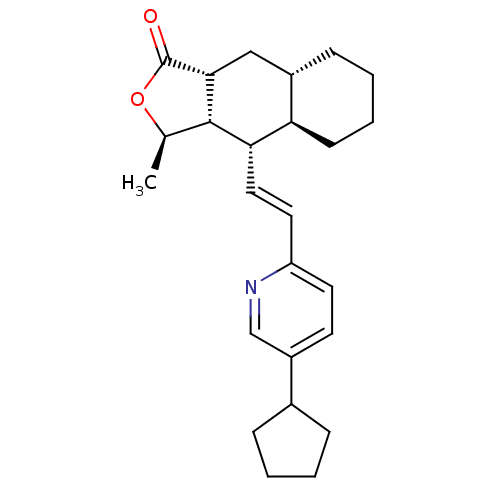
Affinity DataIC50: 16nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 27nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 29nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 36nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 43nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 47nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 88nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 115nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 130nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 145nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 207nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 263nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 285nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 350nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 440nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataKi: 487nMAssay Description:Binding affinity to cannabinoid CB2 receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 1.32E+3nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 2.14E+3nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+3nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 3.60E+3nMAssay Description:Displacement of [3H]haTRAP from human PAR1 receptor in platelet membraneMore data for this Ligand-Target Pair
Affinity DataKi: 1.30E+4nMAssay Description:Binding affinity to cannabinoid CB2 receptorMore data for this Ligand-Target Pair