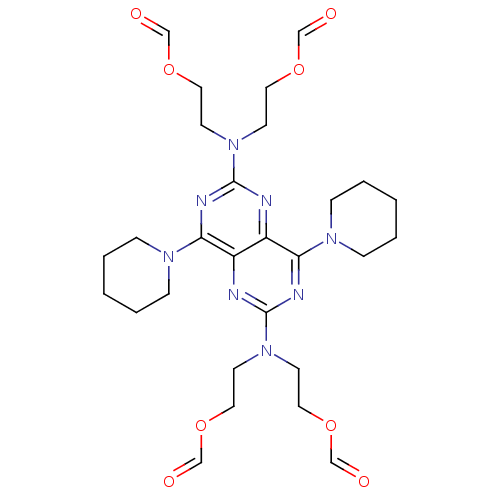
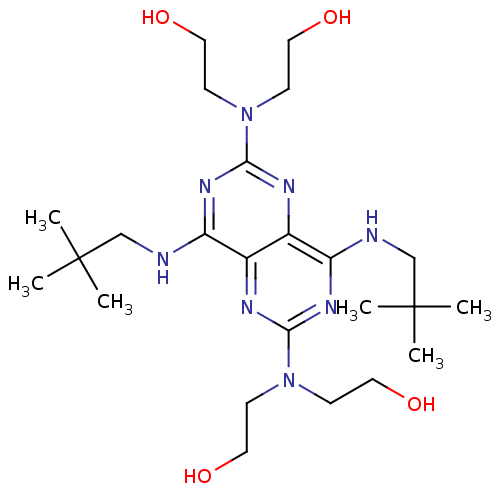
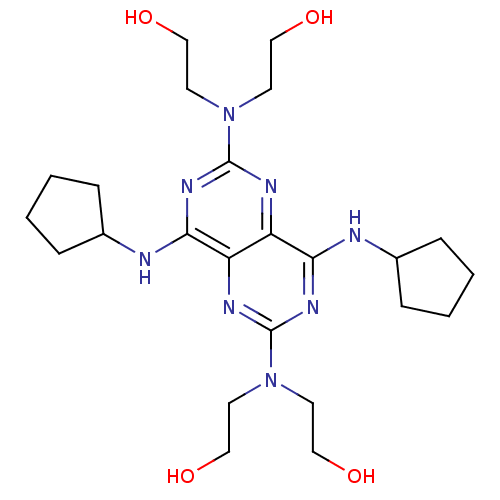
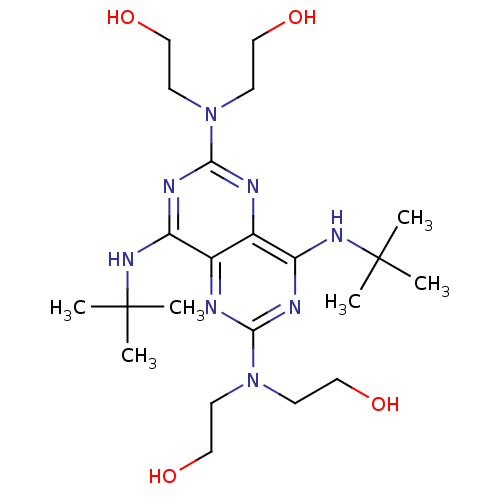
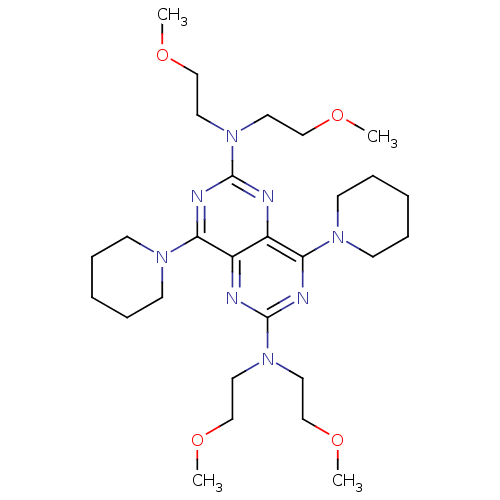
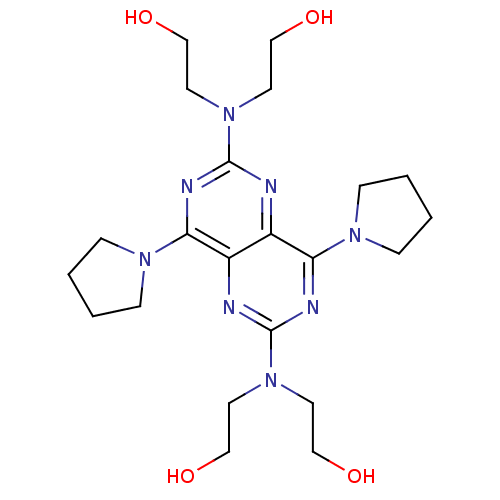
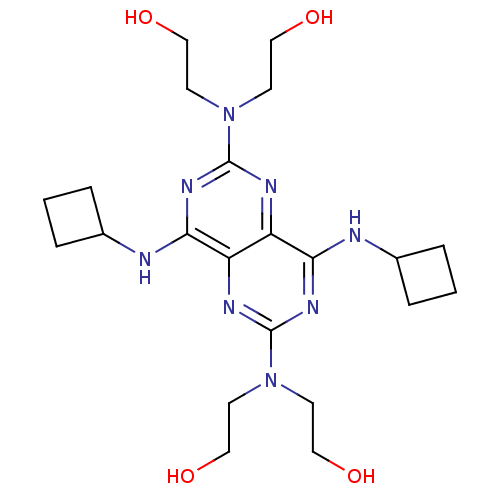
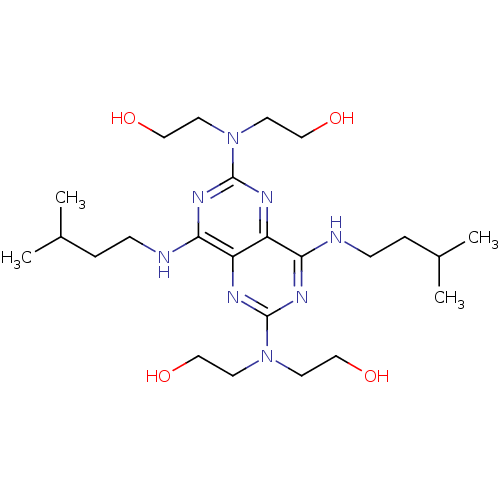
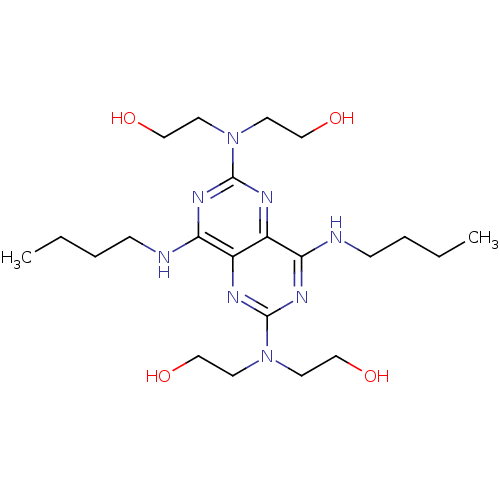
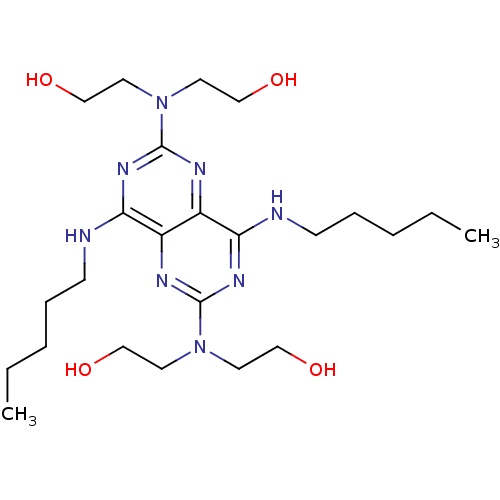
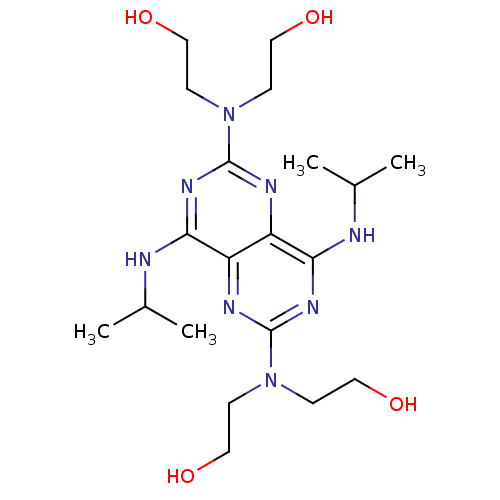
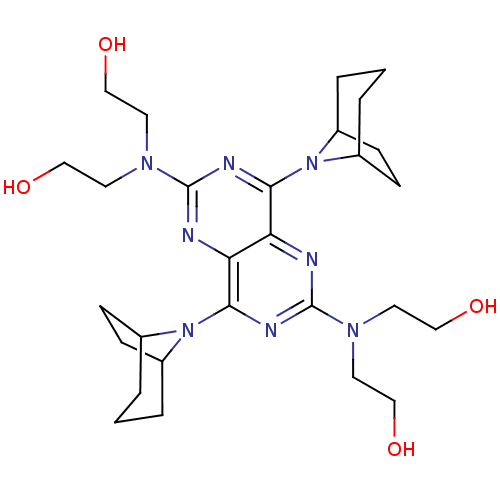
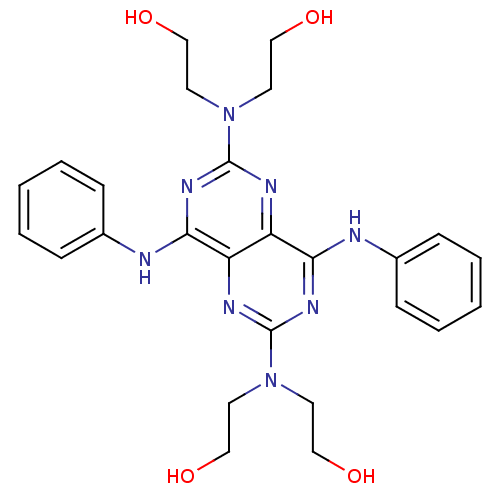
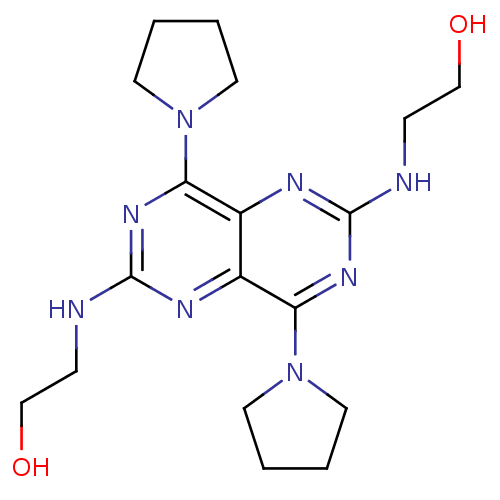
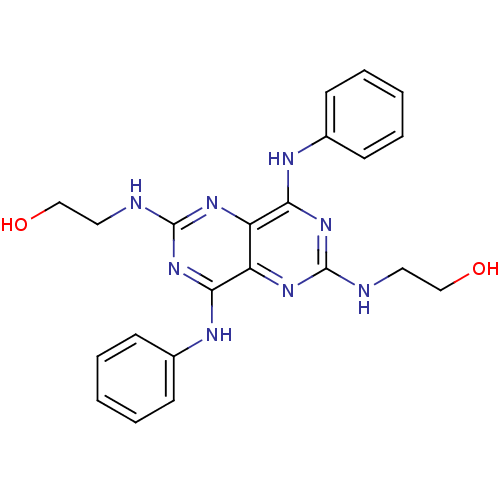
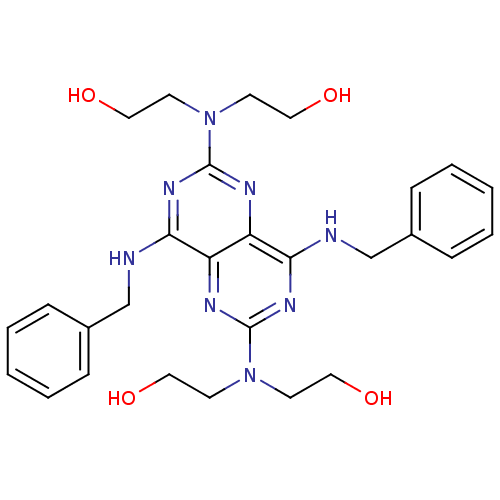
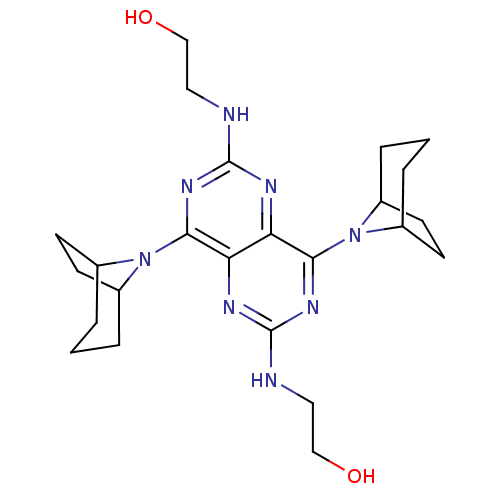
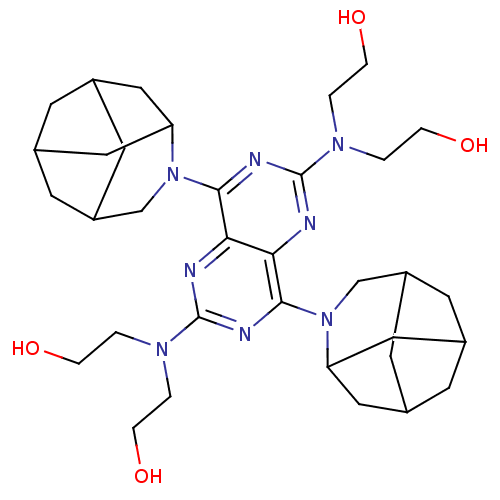
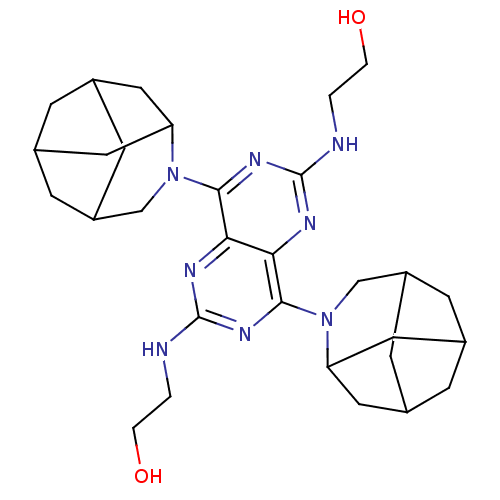
Report error Found 80 Enz. Inhib. hit(s) with all data for entry = 2707
Affinity DataKi: 0.430nM ΔG°: -52.9kJ/mole IC50: 7.60nMpH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 0.490nM ΔG°: -52.6kJ/mole IC50: 8.67nMpH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 0.770nM ΔG°: -51.5kJ/mole IC50: 13.6nMpH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 0.860nM ΔG°: -51.2kJ/mole IC50: 15.2nMpH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 4.30nM IC50: 76nMAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 8.18nM ΔG°: -45.7kJ/mole IC50: 145nMpH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 8.20nM IC50: 145nMAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 14.7nM IC50: 260nMAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 15.8nM IC50: 280nMAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 16.8nM IC50: 297nMAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 17.1nM IC50: 302nMAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 21.2nM ΔG°: -43.4kJ/mole IC50: 375nMpH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 38nM IC50: 673nMAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 38nM ΔG°: -41.9kJ/mole IC50: 672nMpH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 53.1nM IC50: 940nMAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 91.6nM IC50: 1.62E+3nMAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 99.7nM ΔG°: -39.6kJ/mole IC50: 1.76E+3nMpH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 104nM IC50: 1.84E+3nMAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 121nM IC50: 2.14E+3nMAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 136nM IC50: 2.41E+3nMAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 140nM IC50: 2.48E+3nMAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 191nM IC50: 3.38E+3nMAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 193nM ΔG°: -37.9kJ/mole IC50: 3.42E+3nMpH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 216nM ΔG°: -37.7kJ/mole IC50: 3.83E+3nMpH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 217nM ΔG°: -37.7kJ/mole IC50: 3.83E+3nMpH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 280nM ΔG°: -37.0kJ/mole IC50: 4.95E+3nMpH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 300nM IC50: 5.31E+3nMAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 325nM IC50: 5.75E+3nMAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 393nM ΔG°: -36.2kJ/mole IC50: 6.96E+3nMpH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 428nM IC50: 7.55E+3nMAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 449nM ΔG°: -35.9kJ/mole IC50: 7.95E+3nMpH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 489nM IC50: 8.66E+3nMAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataKi: 691nM ΔG°: -34.8kJ/mole IC50: 1.22E+4nMpH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DatapH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DatapH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DatapH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DatapH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DatapH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DataAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DatapH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DatapH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
Affinity DatapH: 7.4 T: 2°CAssay Description:The compounds were tested to determine their ENT1 nucleoside transporter binding ability by a flow cytometric assay using human leukemia K562 cells i...More data for this Ligand-Target Pair
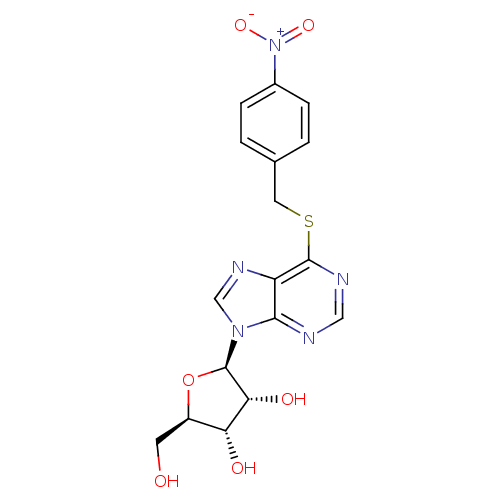
 3D Structure (crystal)
3D Structure (crystal)