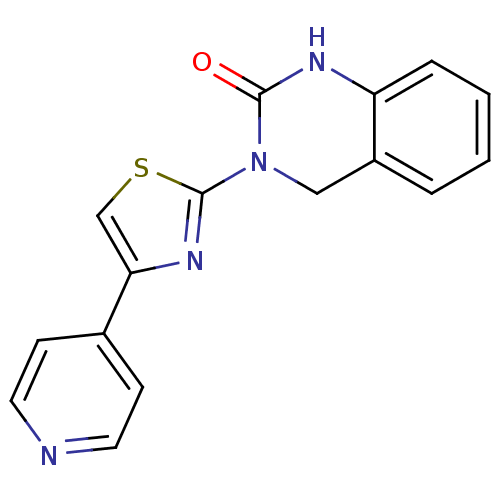
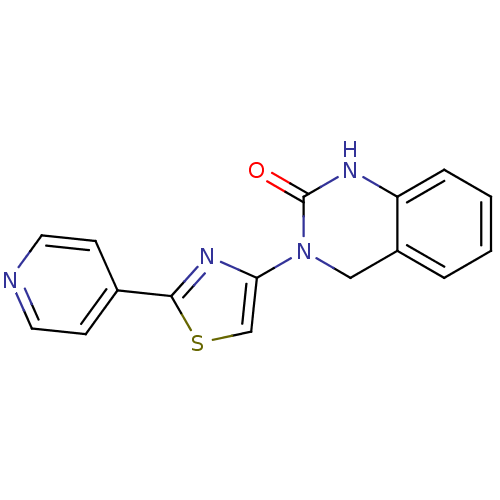
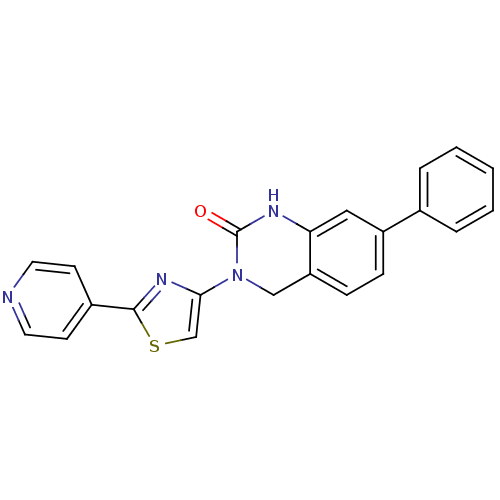
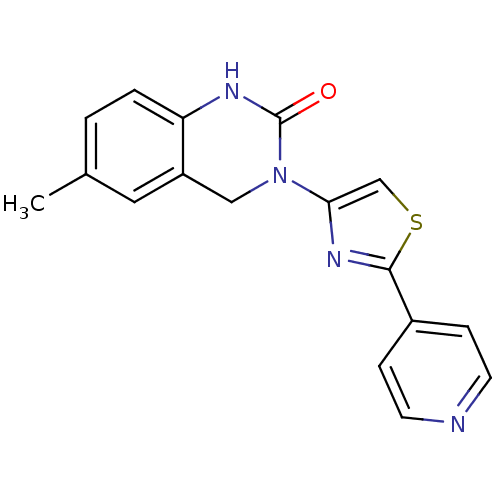
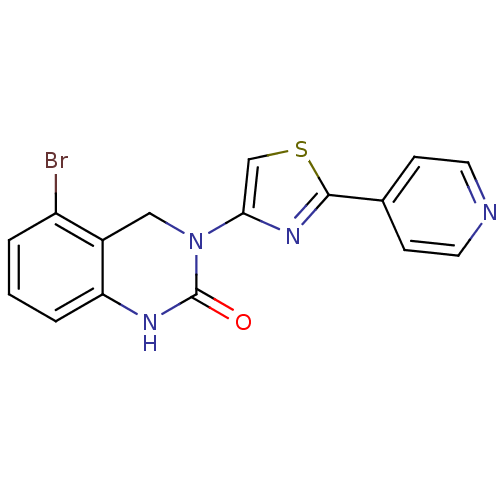
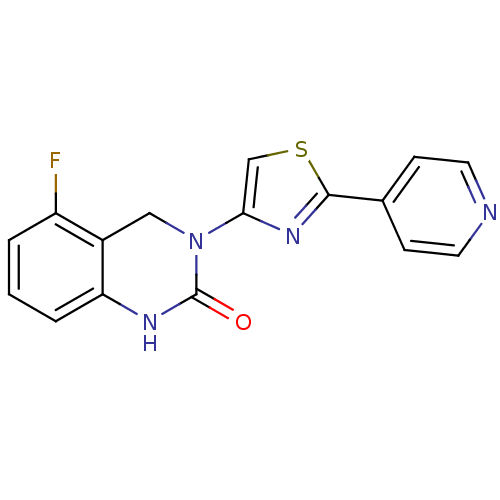
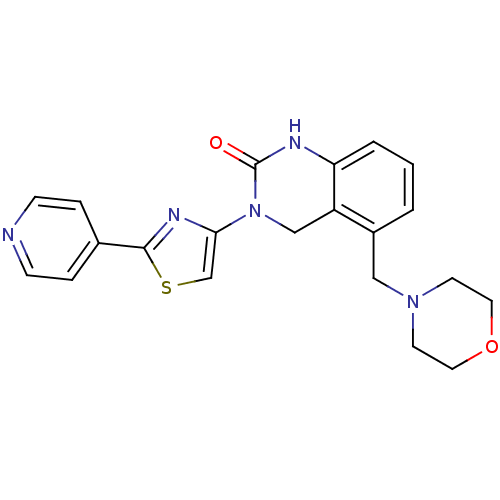
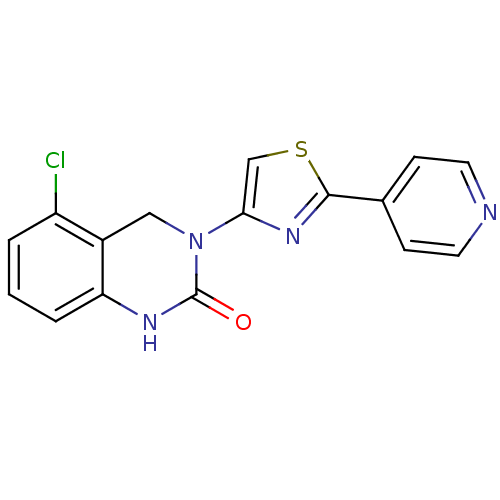
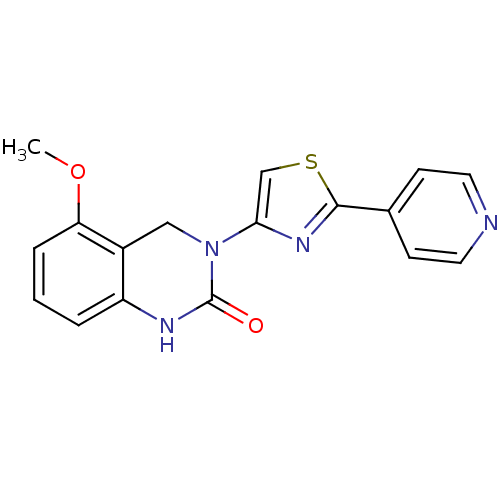
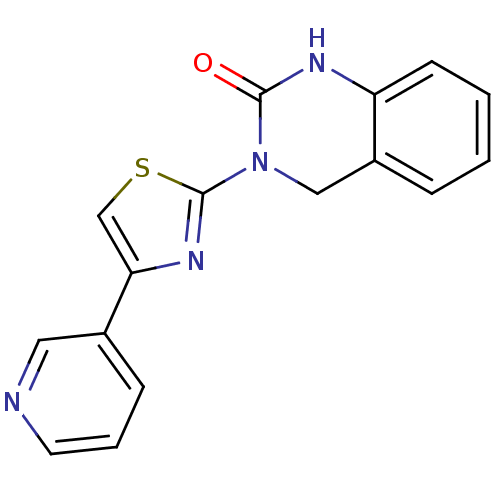
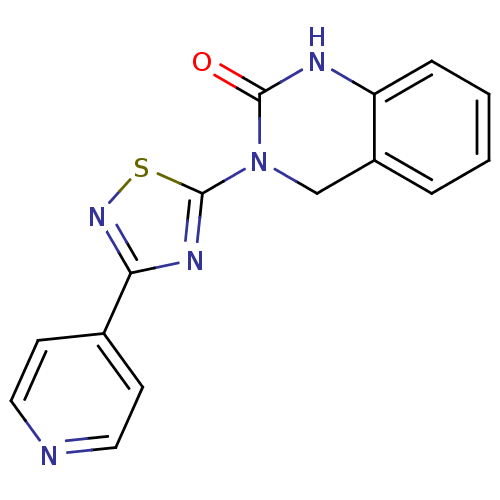
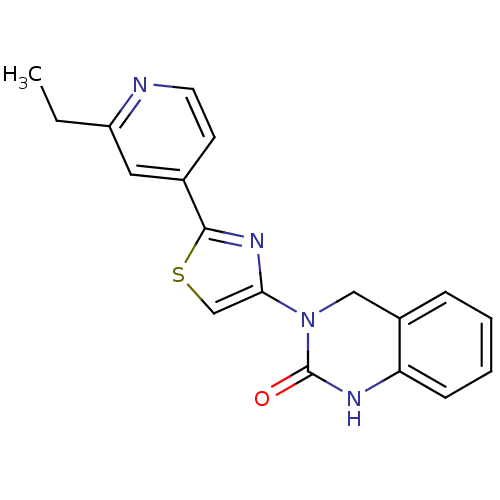
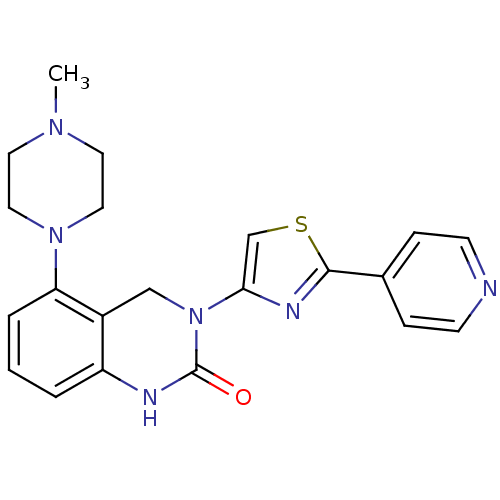
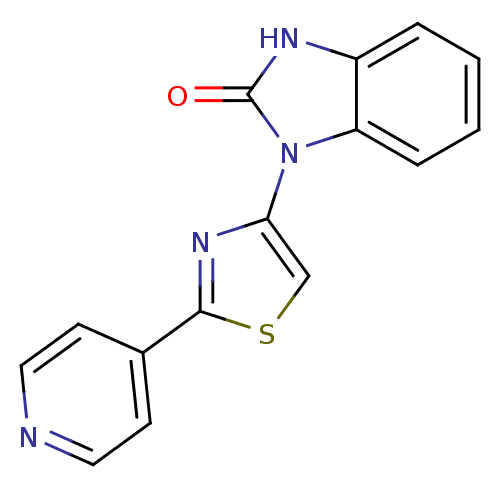
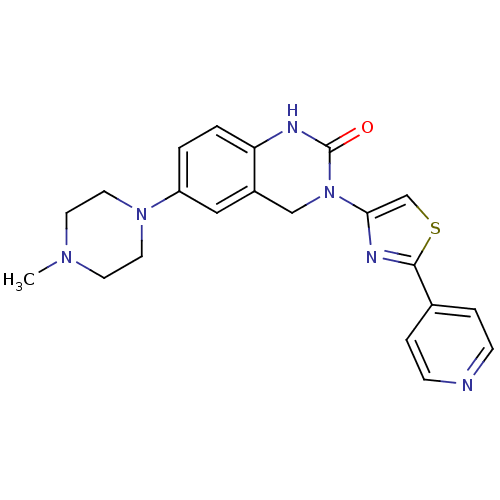
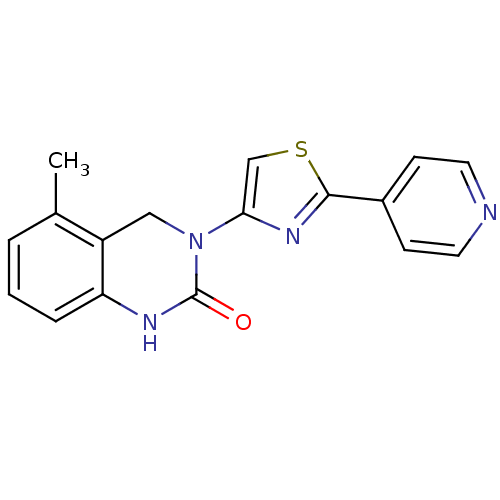
Report error Found 29 Enz. Inhib. hit(s) with all data for entry = 2460
Affinity DataIC50: 16nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 38nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 52nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 62nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 72nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 77nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 79nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 113nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 142nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 165nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 228nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 240nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 284nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 332nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 362nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 399nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 529nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 616nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 746nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 829nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 1.16E+3nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 1.17E+3nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 1.26E+3nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 2.99E+3nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 7.36E+3nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 7.5 T: 2°CAssay Description:In vitro CDK assay using purified enzyme, was incubated at 25 °C with substrate, and test compounds in the presence of ATP/ [gamma-33P] ATP. 33P...More data for this Ligand-Target Pair