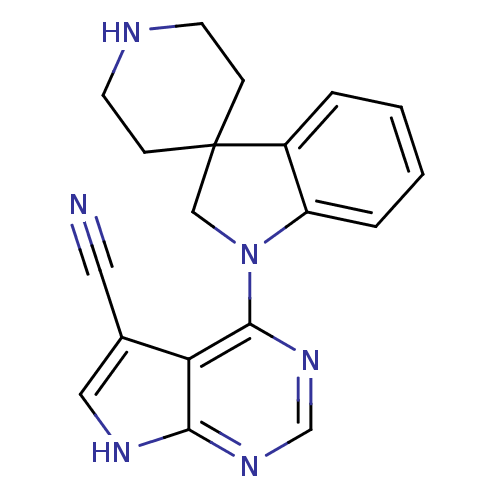
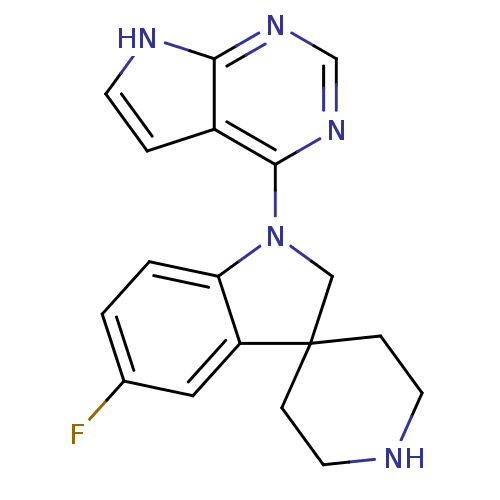
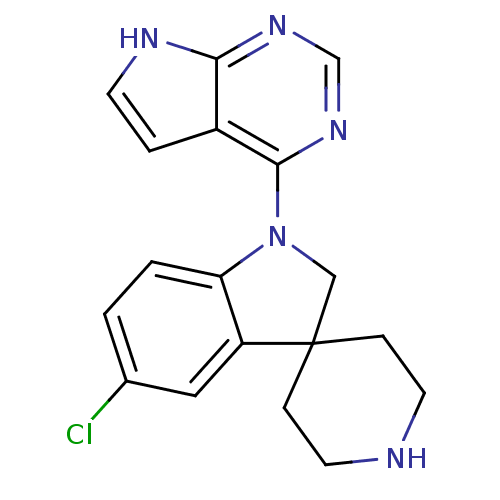
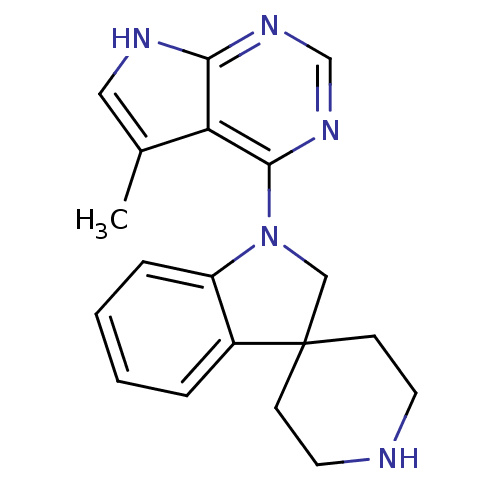
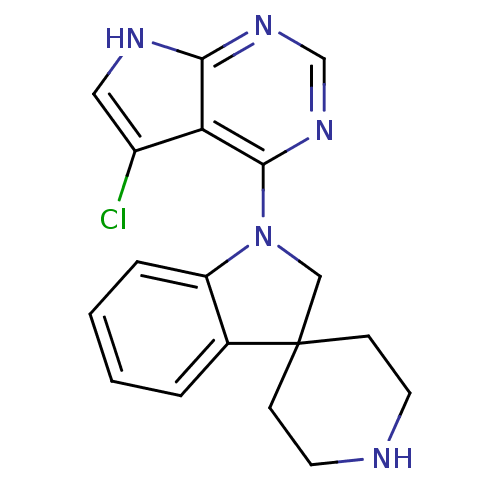
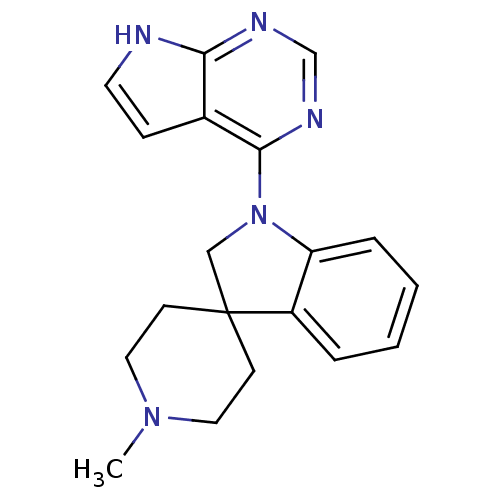
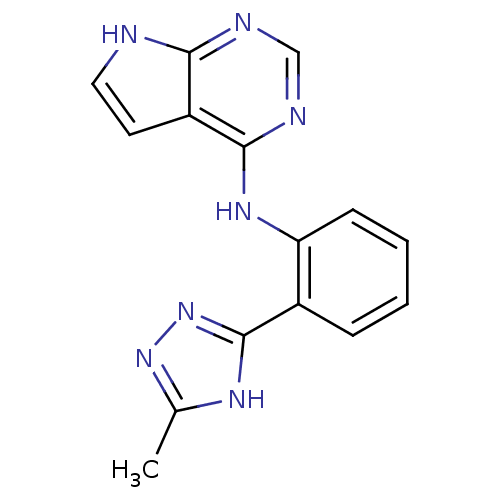
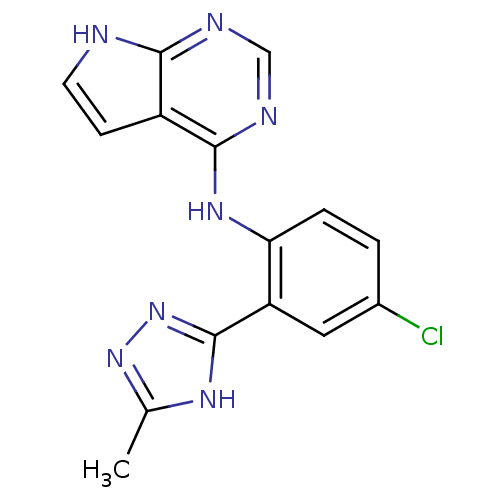
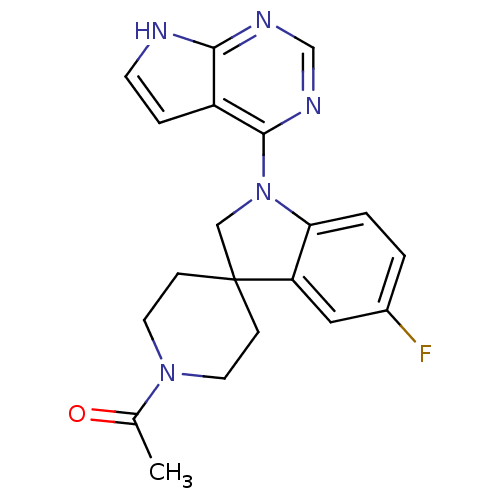
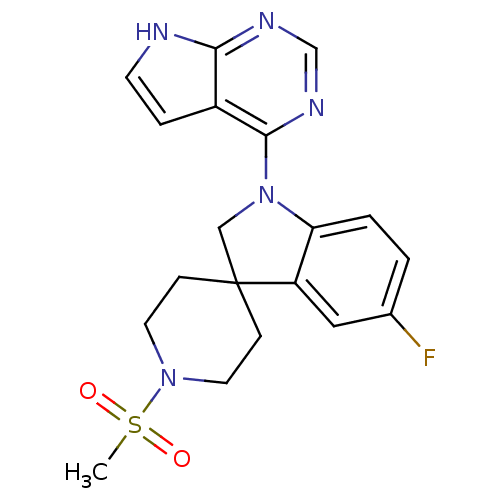
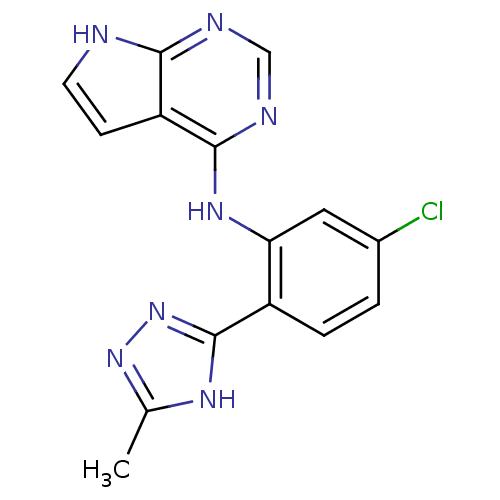
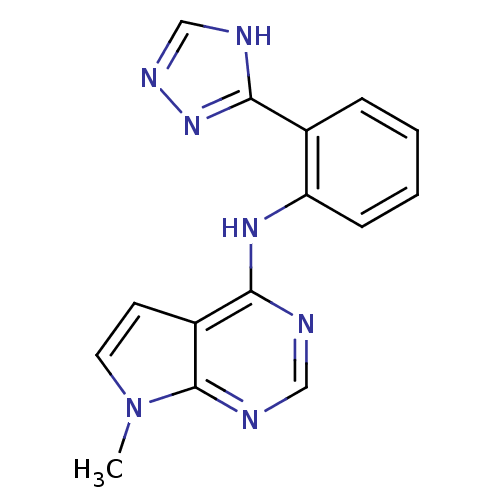
Report error Found 19 Enz. Inhib. hit(s) with all data for entry = 2804
Affinity DataIC50: 2.40nMpH: 7.5 T: 2°CAssay Description:A fluorescence polarization IMAP type assay is used. After a 90 min incubation of reaction mixtures containing enzyme, substrate, and inhibitor, IMAP...More data for this Ligand-Target Pair
Affinity DataIC50: 3.10nMpH: 7.5 T: 2°CAssay Description:A fluorescence polarization IMAP type assay is used. After a 90 min incubation of reaction mixtures containing enzyme, substrate, and inhibitor, IMAP...More data for this Ligand-Target Pair
Affinity DataIC50: 4.80nMpH: 7.5 T: 2°CAssay Description:A fluorescence polarization IMAP type assay is used. After a 90 min incubation of reaction mixtures containing enzyme, substrate, and inhibitor, IMAP...More data for this Ligand-Target Pair
Affinity DataIC50: 4.80nMpH: 7.5 T: 2°CAssay Description:A fluorescence polarization IMAP type assay is used. After a 90 min incubation of reaction mixtures containing enzyme, substrate, and inhibitor, IMAP...More data for this Ligand-Target Pair
Affinity DataIC50: 5.30nMpH: 7.5 T: 2°CAssay Description:A fluorescence polarization IMAP type assay is used. After a 90 min incubation of reaction mixtures containing enzyme, substrate, and inhibitor, IMAP...More data for this Ligand-Target Pair
Affinity DataIC50: 5.80nMpH: 7.5 T: 2°CAssay Description:A fluorescence polarization IMAP type assay is used. After a 90 min incubation of reaction mixtures containing enzyme, substrate, and inhibitor, IMAP...More data for this Ligand-Target Pair
Affinity DataIC50: 12.7nMpH: 7.5 T: 2°CAssay Description:A fluorescence polarization IMAP type assay is used. After a 90 min incubation of reaction mixtures containing enzyme, substrate, and inhibitor, IMAP...More data for this Ligand-Target Pair
Affinity DataIC50: 42nMpH: 7.5 T: 2°CAssay Description:A fluorescence polarization IMAP type assay is used. After a 90 min incubation of reaction mixtures containing enzyme, substrate, and inhibitor, IMAP...More data for this Ligand-Target Pair
Affinity DataIC50: 151nMpH: 7.5 T: 2°CAssay Description:A fluorescence polarization IMAP type assay is used. After a 90 min incubation of reaction mixtures containing enzyme, substrate, and inhibitor, IMAP...More data for this Ligand-Target Pair
Affinity DataIC50: 171nMpH: 7.5 T: 2°CAssay Description:A fluorescence polarization IMAP type assay is used. After a 90 min incubation of reaction mixtures containing enzyme, substrate, and inhibitor, IMAP...More data for this Ligand-Target Pair
Affinity DataIC50: 210nMpH: 7.5 T: 2°CAssay Description:A fluorescence polarization IMAP type assay is used. After a 90 min incubation of reaction mixtures containing enzyme, substrate, and inhibitor, IMAP...More data for this Ligand-Target Pair
Affinity DataIC50: 294nMpH: 7.5 T: 2°CAssay Description:A fluorescence polarization IMAP type assay is used. After a 90 min incubation of reaction mixtures containing enzyme, substrate, and inhibitor, IMAP...More data for this Ligand-Target Pair
Affinity DataIC50: 318nMpH: 7.5 T: 2°CAssay Description:A fluorescence polarization IMAP type assay is used. After a 90 min incubation of reaction mixtures containing enzyme, substrate, and inhibitor, IMAP...More data for this Ligand-Target Pair
Affinity DataIC50: 425nMpH: 7.5 T: 2°CAssay Description:A fluorescence polarization IMAP type assay is used. After a 90 min incubation of reaction mixtures containing enzyme, substrate, and inhibitor, IMAP...More data for this Ligand-Target Pair
Affinity DataIC50: 1.33E+3nMpH: 7.5 T: 2°CAssay Description:A fluorescence polarization IMAP type assay is used. After a 90 min incubation of reaction mixtures containing enzyme, substrate, and inhibitor, IMAP...More data for this Ligand-Target Pair
Affinity DataIC50: 2.79E+3nMpH: 7.5 T: 2°CAssay Description:A fluorescence polarization IMAP type assay is used. After a 90 min incubation of reaction mixtures containing enzyme, substrate, and inhibitor, IMAP...More data for this Ligand-Target Pair
Affinity DataIC50: 3.19E+3nMpH: 7.5 T: 2°CAssay Description:A fluorescence polarization IMAP type assay is used. After a 90 min incubation of reaction mixtures containing enzyme, substrate, and inhibitor, IMAP...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 7.5 T: 2°CAssay Description:A fluorescence polarization IMAP type assay is used. After a 90 min incubation of reaction mixtures containing enzyme, substrate, and inhibitor, IMAP...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 7.5 T: 2°CAssay Description:A fluorescence polarization IMAP type assay is used. After a 90 min incubation of reaction mixtures containing enzyme, substrate, and inhibitor, IMAP...More data for this Ligand-Target Pair