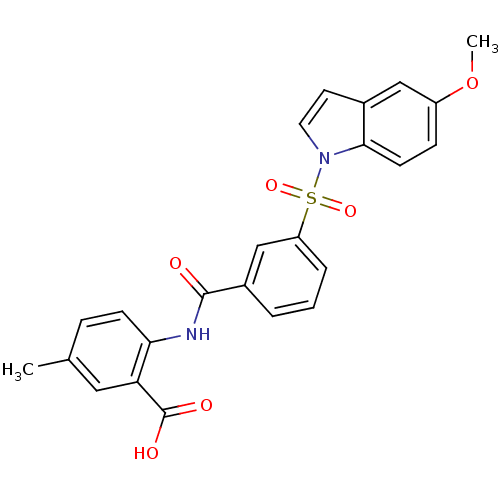
Report error Found 105 Enz. Inhib. hit(s) with all data for entry = 3143
Affinity DataIC50: 4nMpH: 7.0 T: 2°CAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMpH: 7.0 T: 2°CAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMpH: 7.0 T: 2°CAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMpH: 7.0 T: 2°CAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 8nMpH: 7.0 T: 2°CAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 8nMpH: 7.0 T: 2°CAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMpH: 7.0 T: 2°CAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMpH: 7.0 T: 2°CAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMpH: 7.0 T: 2°CAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMpH: 7.0 T: 2°CAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMpH: 7.0 T: 2°CAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMpH: 7.0 T: 2°CAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMpH: 7.0 T: 2°CAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 25nMpH: 7.0 T: 2°CAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 25nMpH: 7.0 T: 2°CAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 32nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 50nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 63nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 126nMpH: 7.0 T: 2°CAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 158nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 794nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 794nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 1.26E+3nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 1.26E+3nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 1.26E+3nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 1.59E+3nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 1.59E+3nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 1.59E+3nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 1.59E+3nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 1.59E+3nMpH: 7.0 T: 2°CAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMpH: 7.0 T: 2°CAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Competition-binding curves for test compounds were determined with expressed human PPAR LBD. Plots of inhibitor concentration versus cpm of radioliga...More data for this Ligand-Target Pair
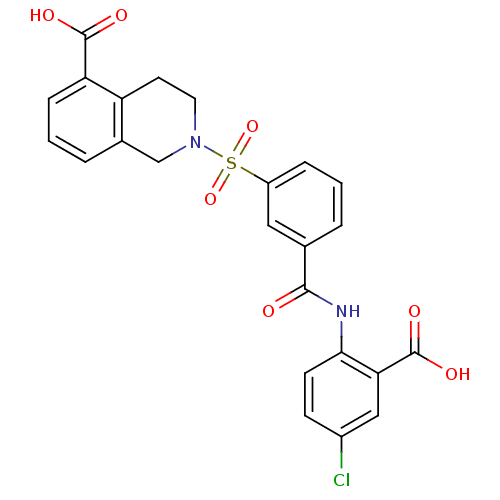
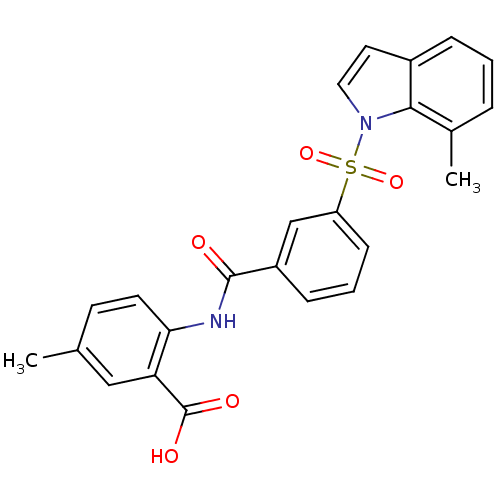
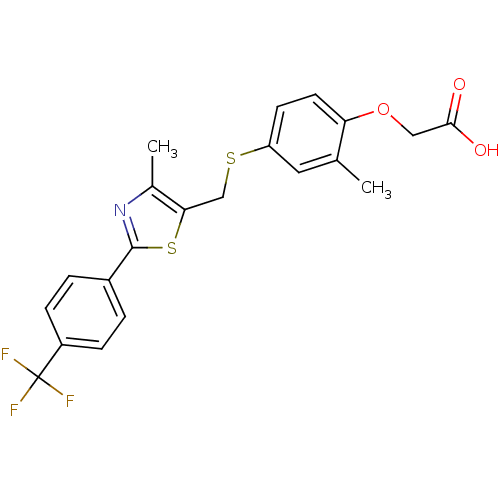
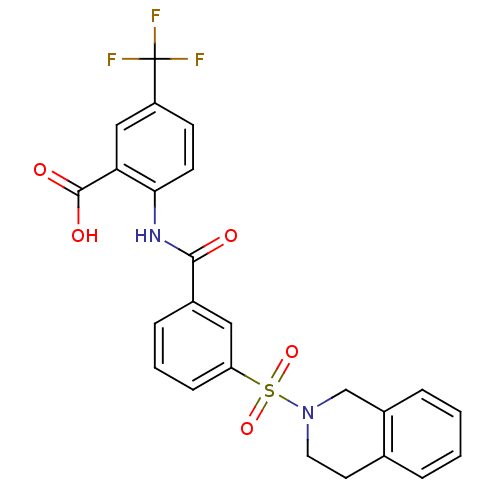
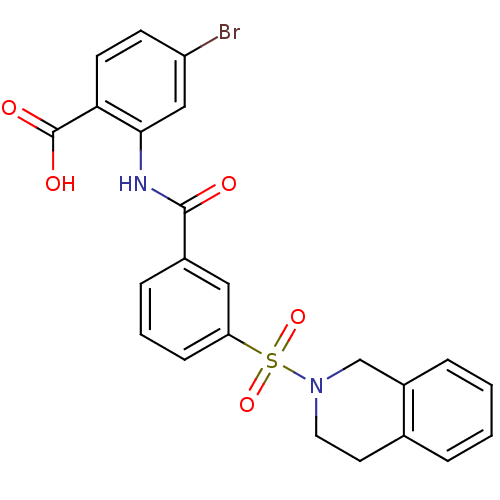
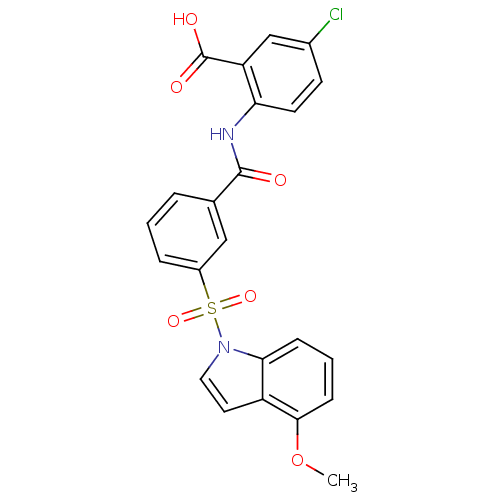
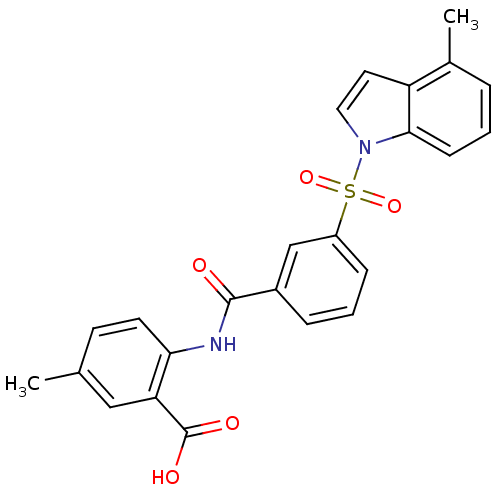
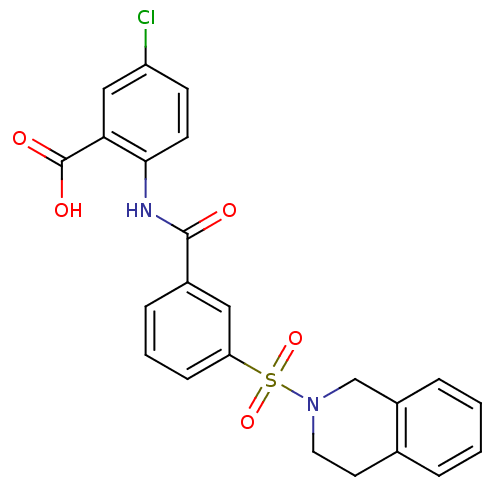
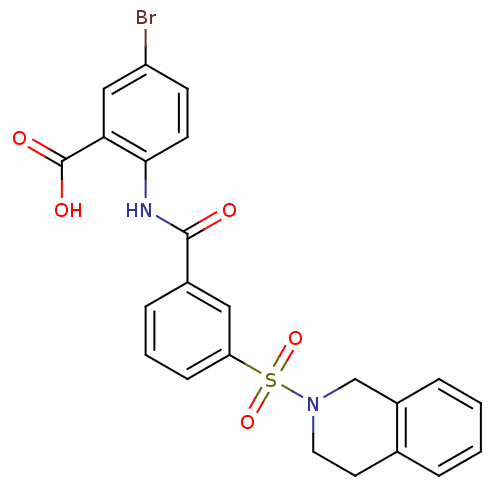
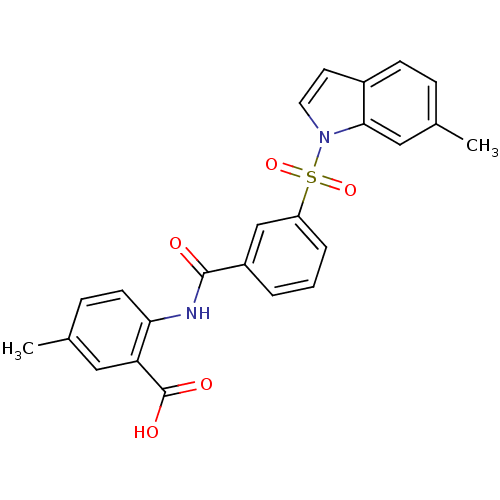
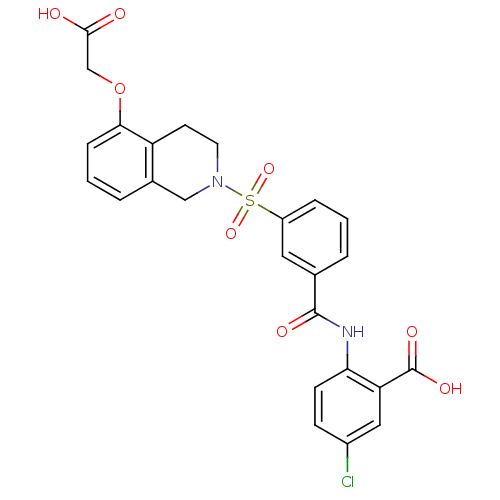
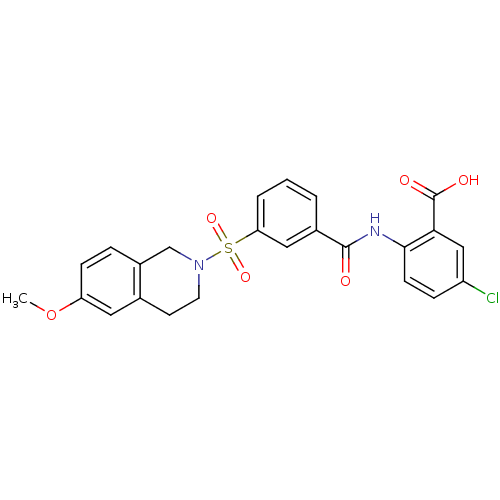
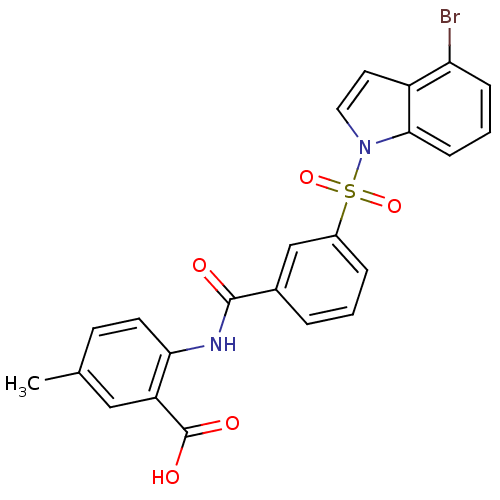
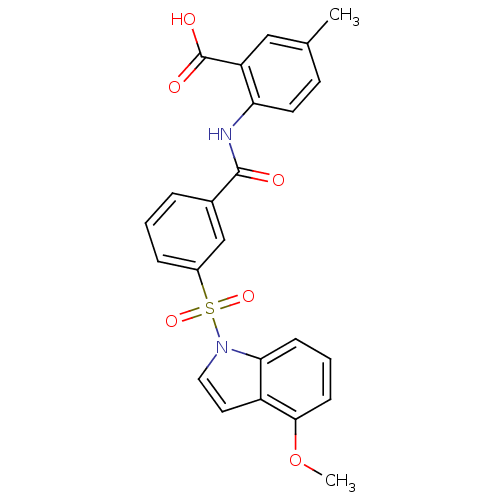
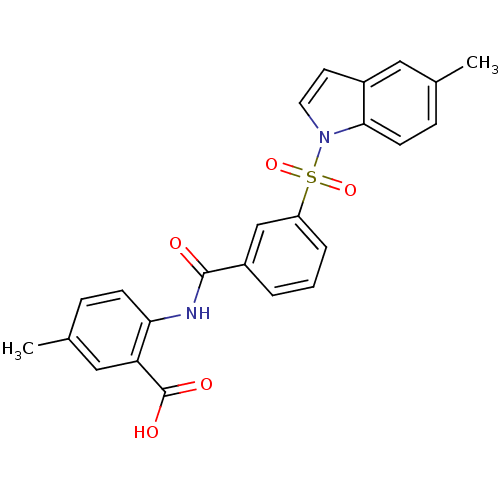
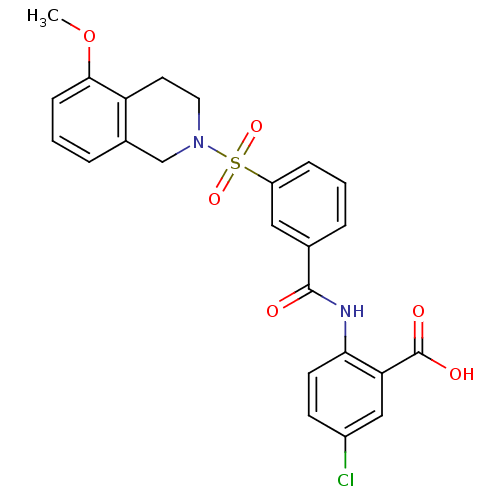
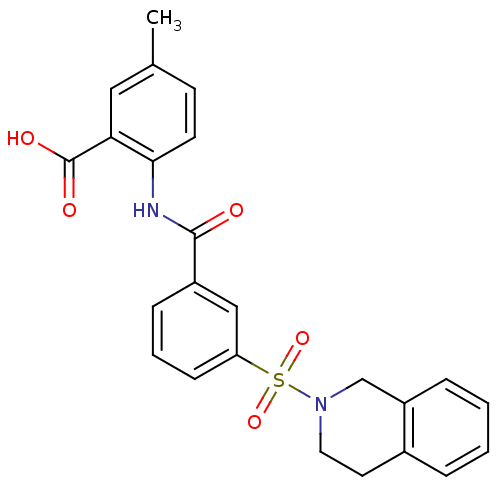
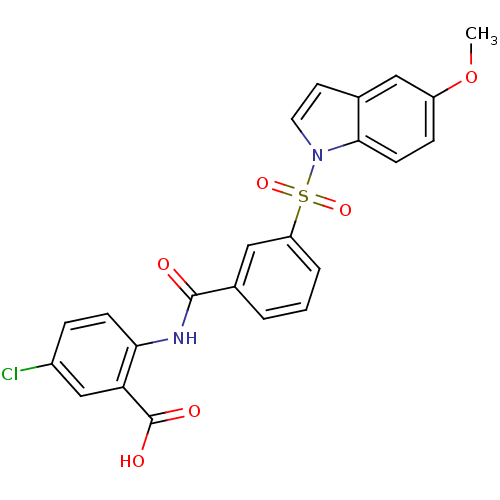
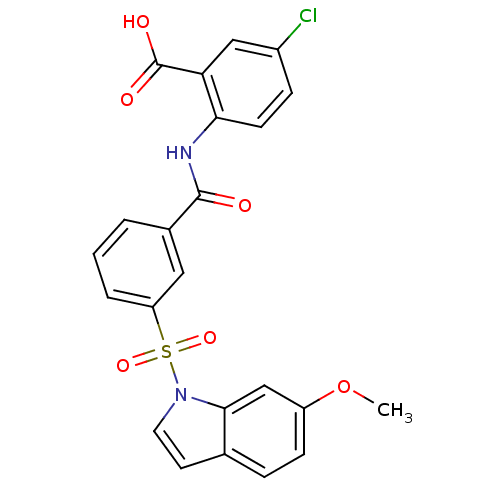
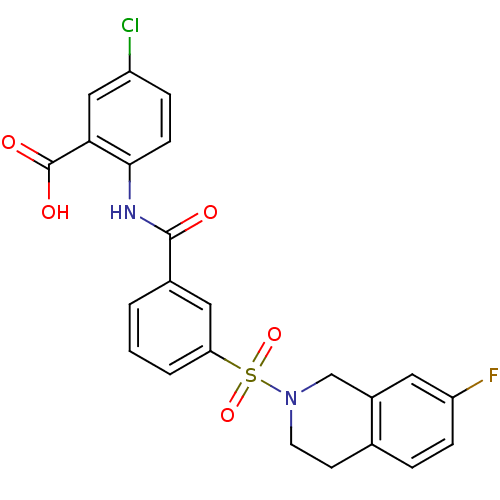
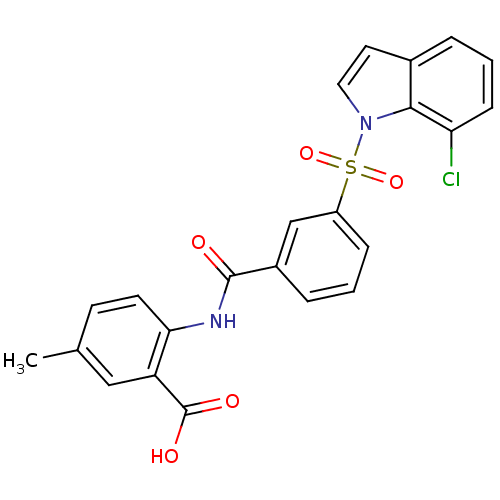
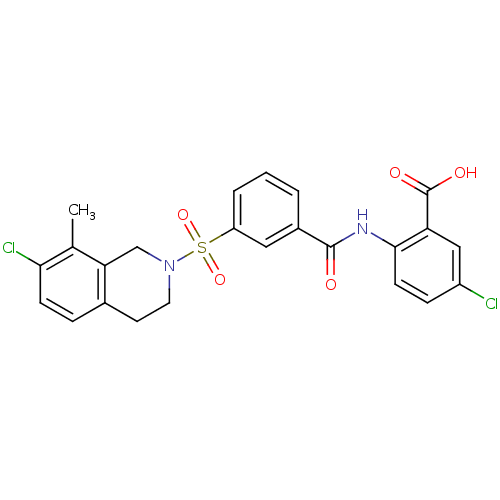
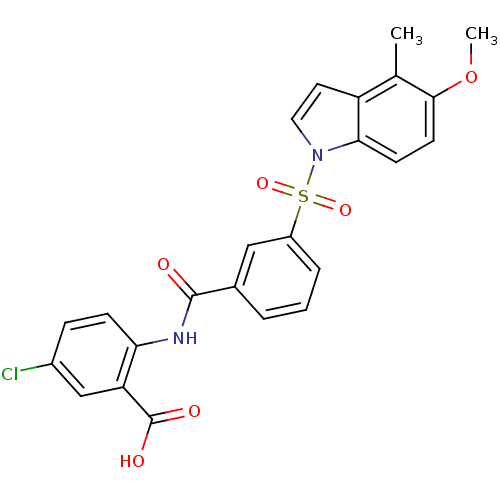
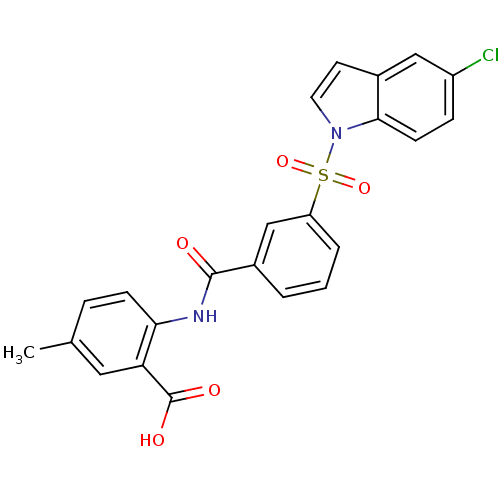
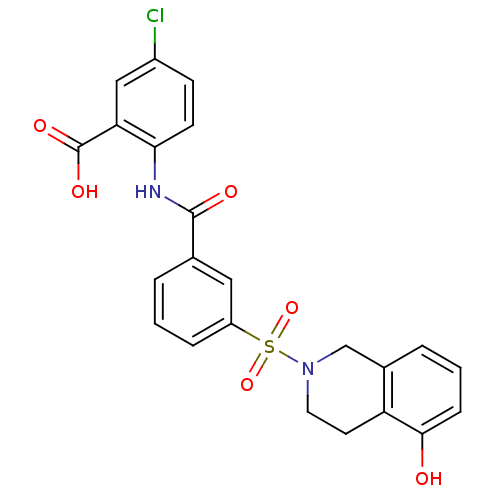
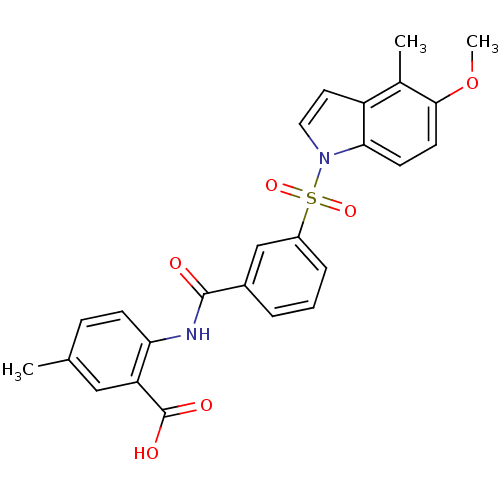
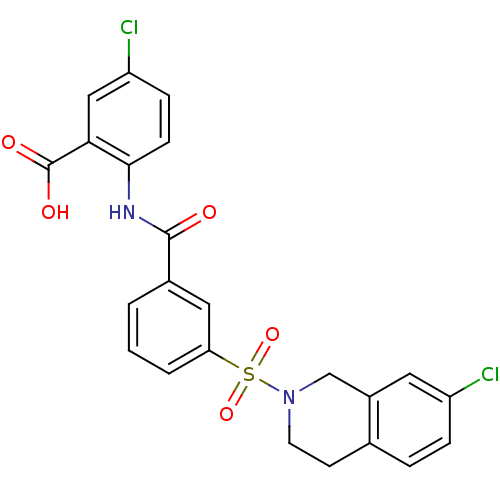
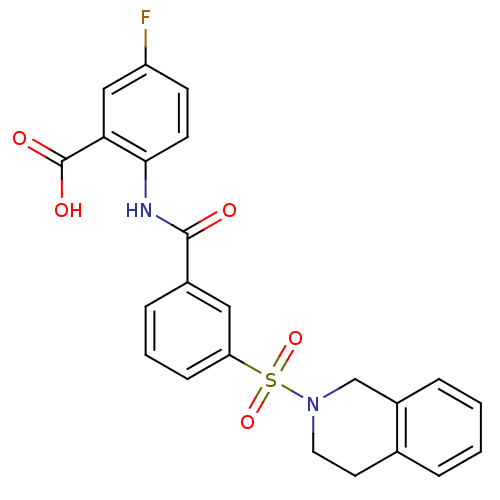
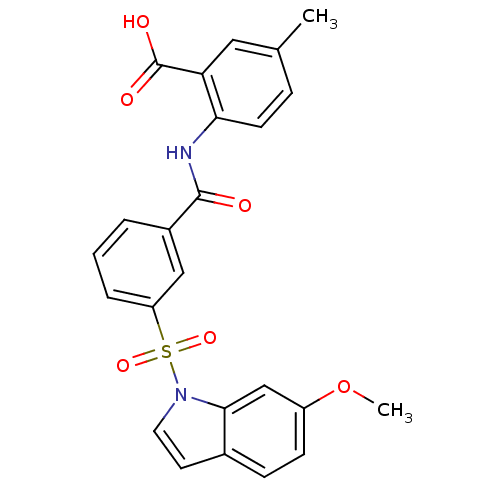
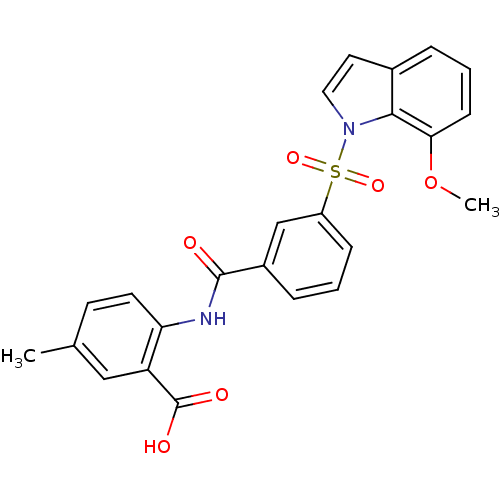
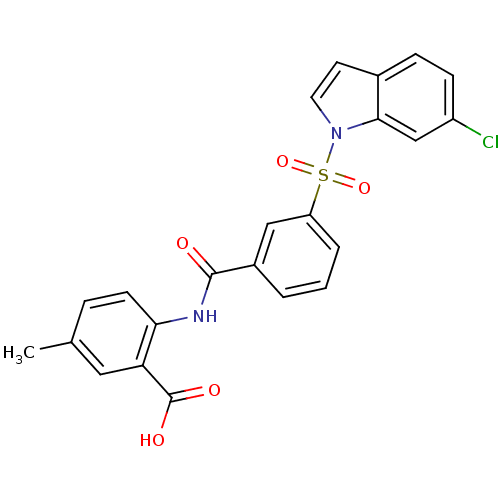
 3D Structure (crystal)
3D Structure (crystal)