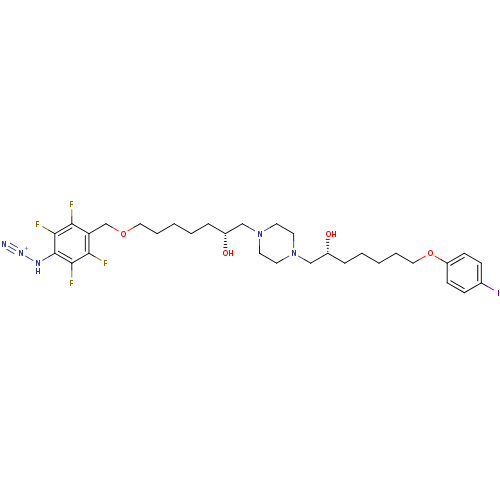
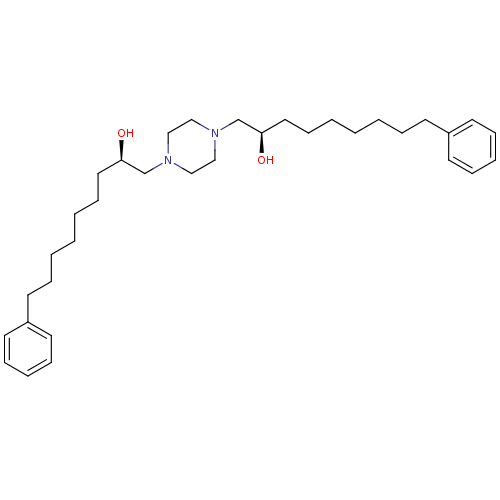
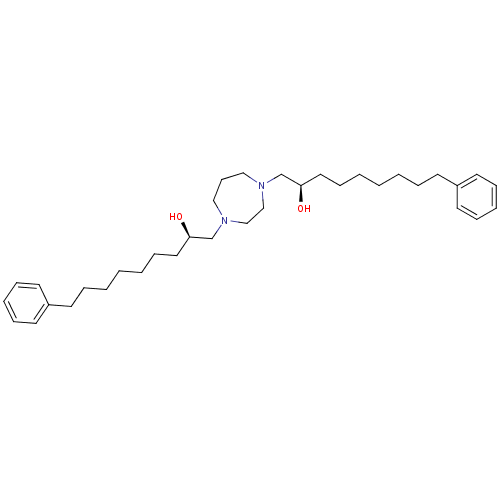
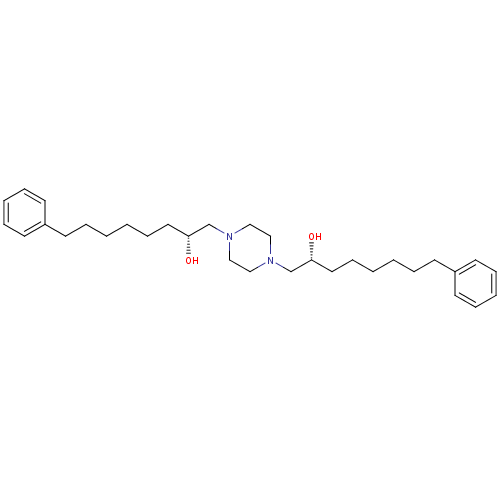
Report error Found 21 Enz. Inhib. hit(s) with all data for entry = 5735
Affinity DataIC50: 1nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 1.20nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 1.70nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 1.70nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 2.20nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 2.30nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 2.60nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 2.80nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 3.60nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 5.90nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 26nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 110nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 670nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 930nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 4.70E+3nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+4nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMAssay Description:Inhibition assay using NADH-ubiquinone oxidoreductase (complex I).More data for this Ligand-Target Pair