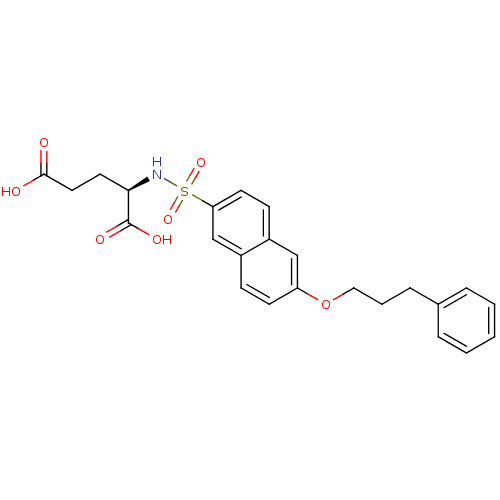
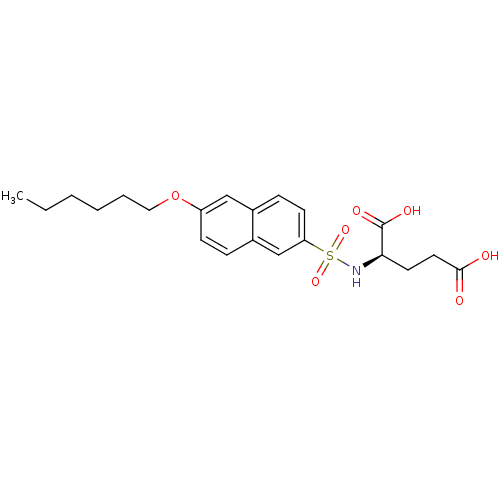
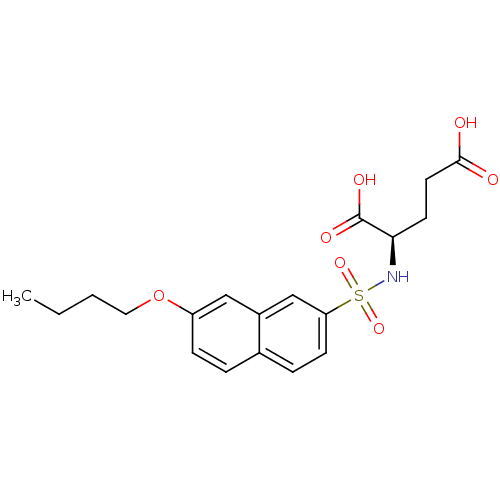
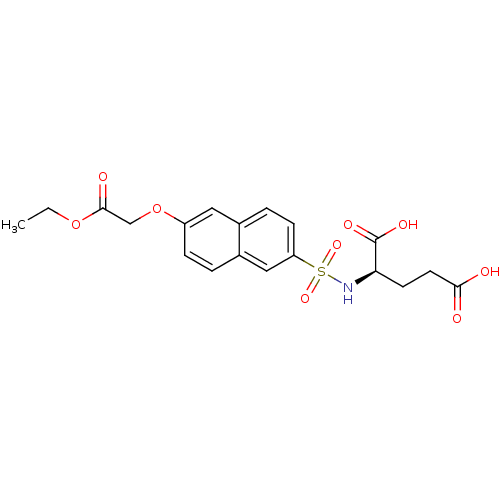
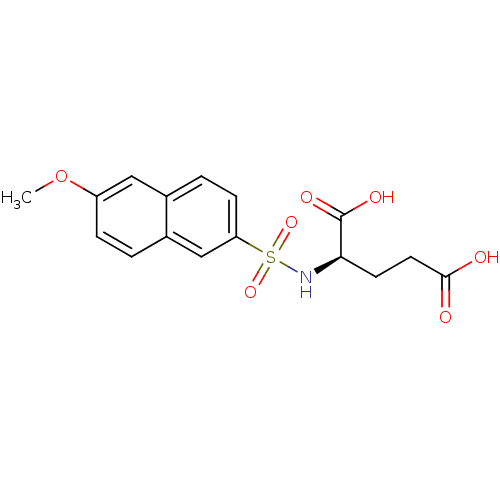
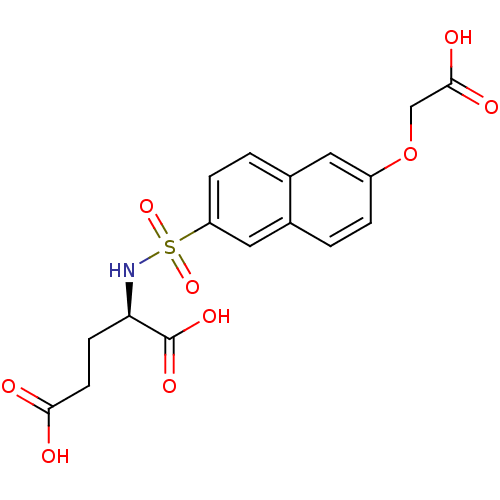
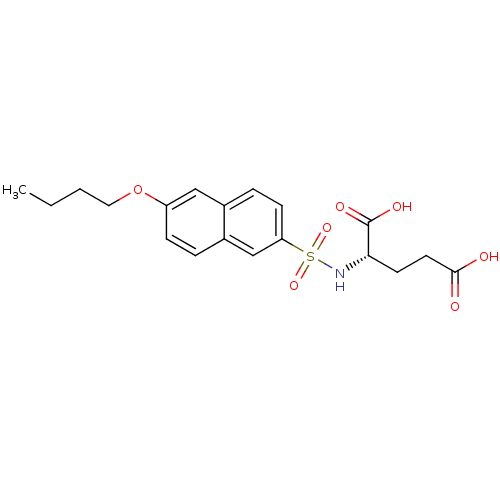
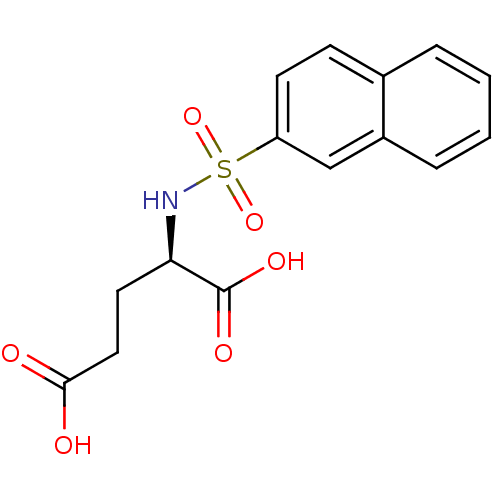
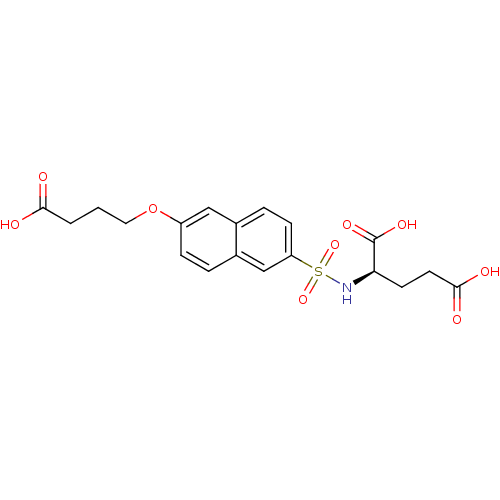
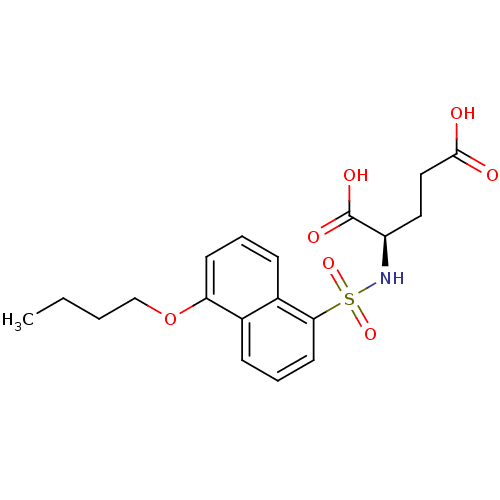
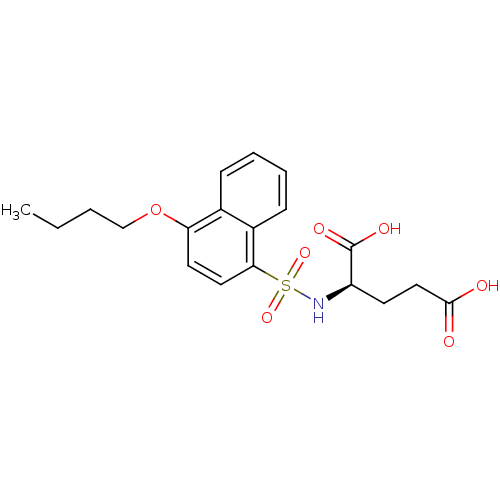
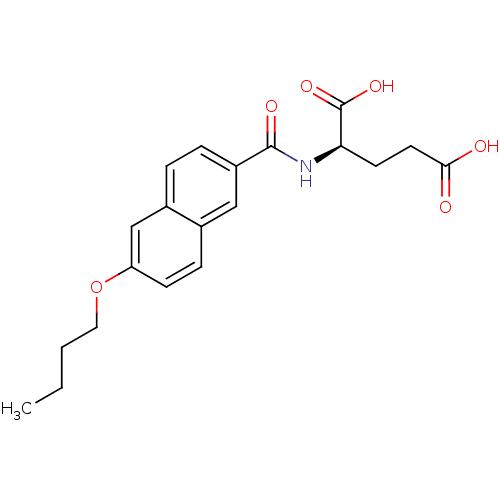
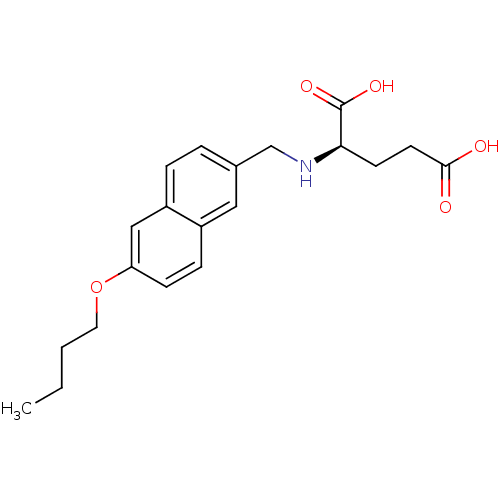
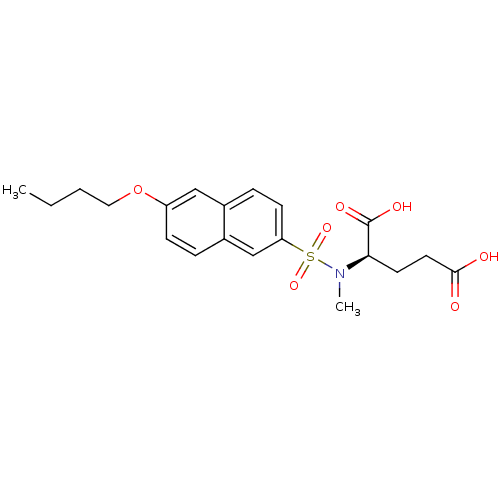
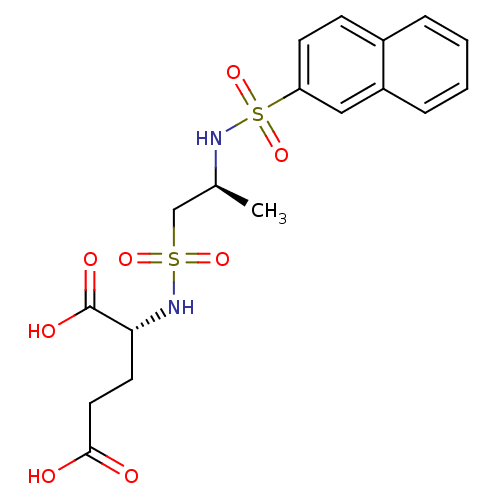
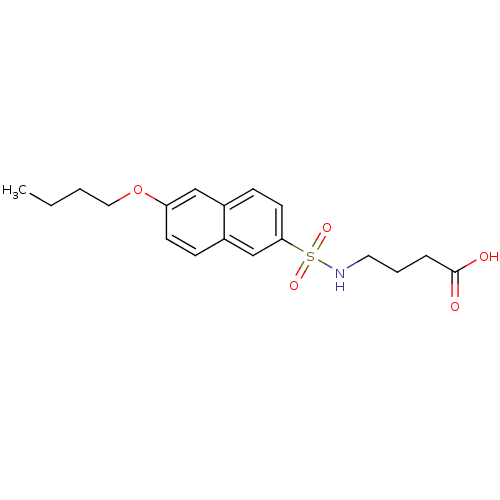
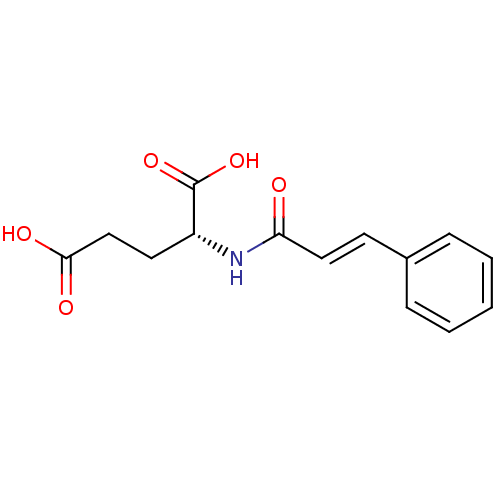
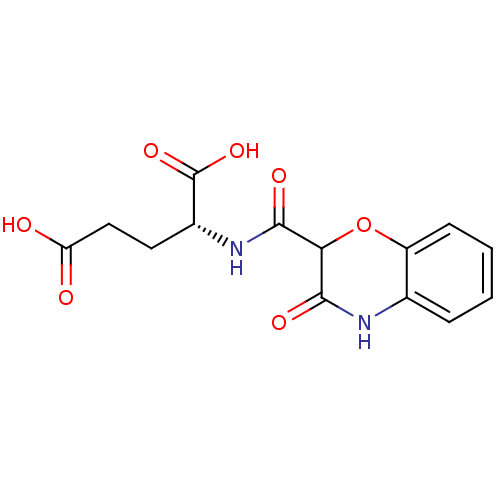
Report error Found 37 Enz. Inhib. hit(s) with all data for entry = 2967
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
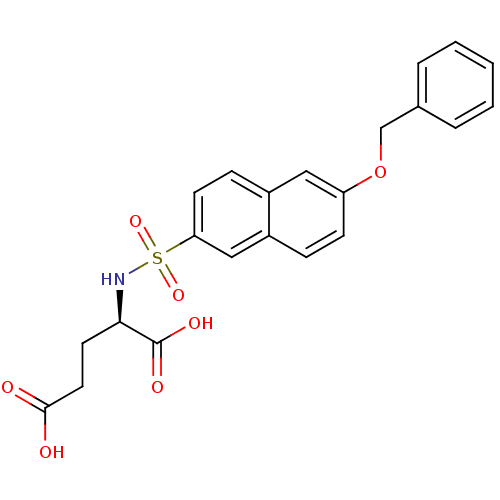
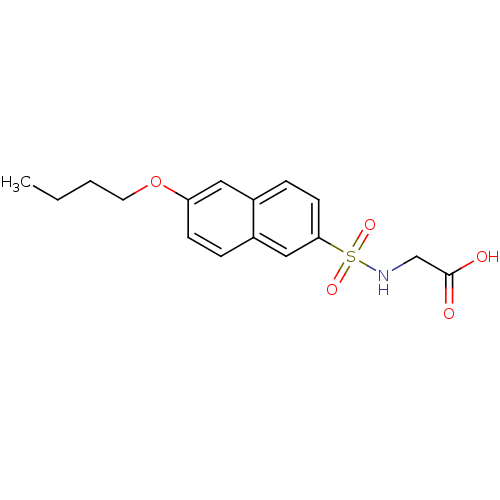
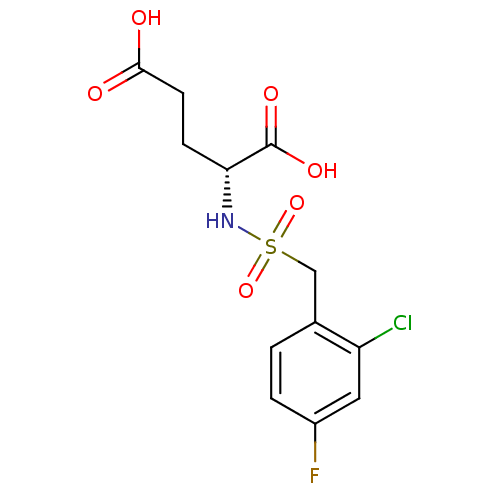
Affinity DataIC50: 8.50E+4nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
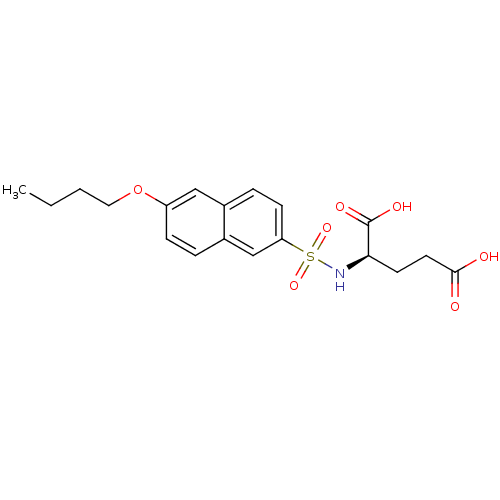
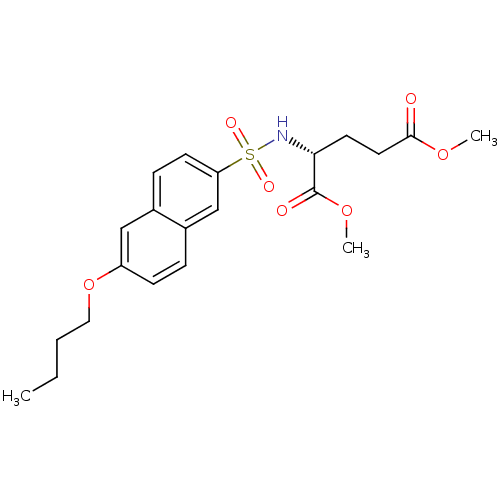
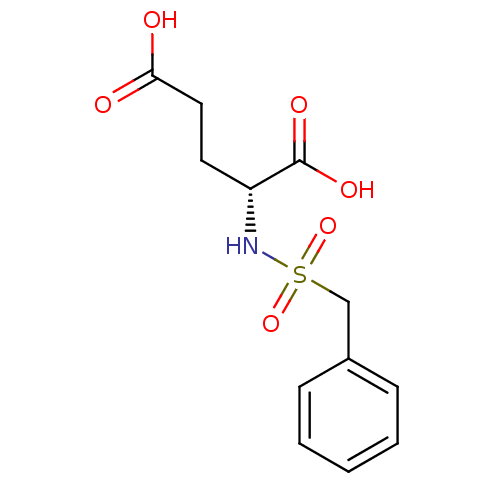
Affinity DataIC50: 1.00E+5nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
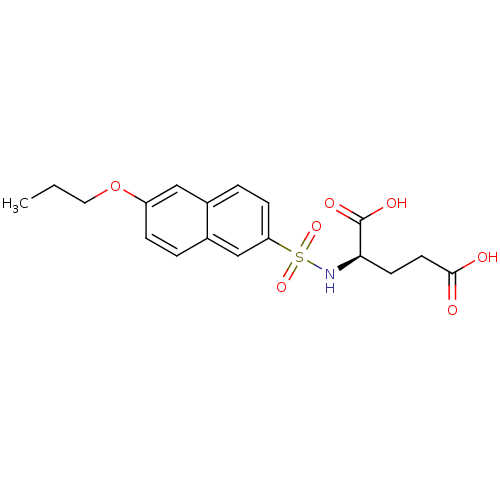
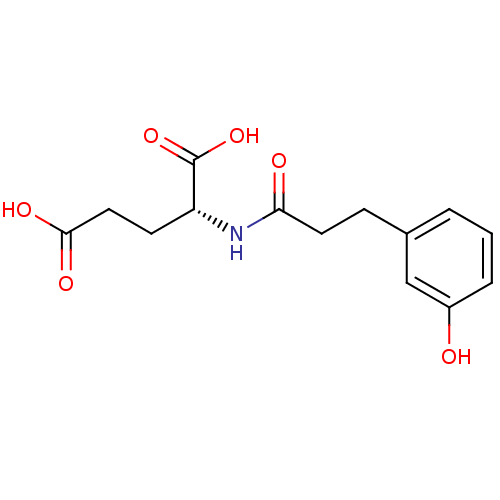
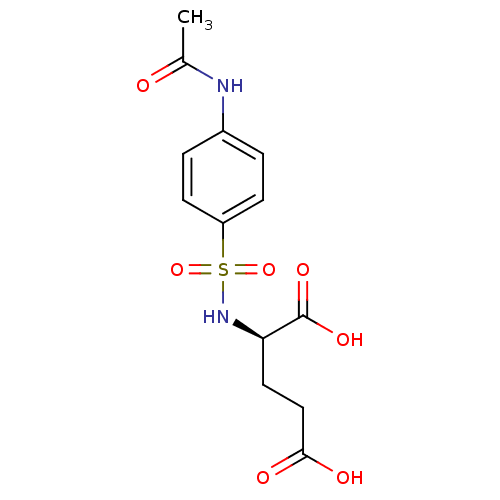
Affinity DataIC50: 1.05E+5nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
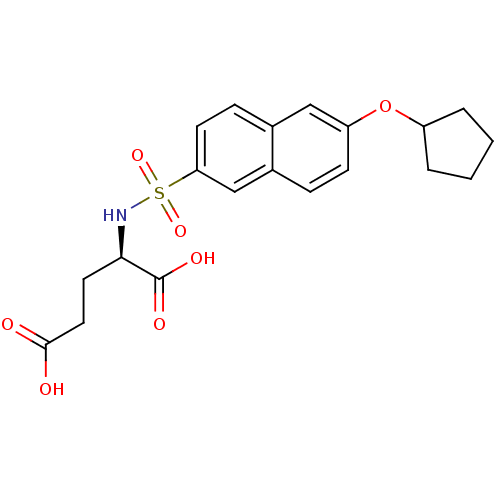
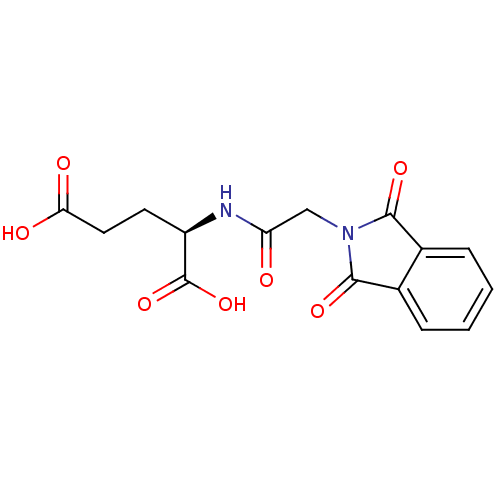
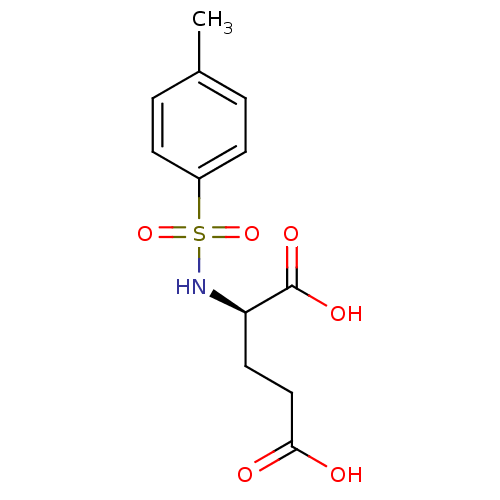
Affinity DataIC50: 1.22E+5nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 1.32E+5nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 1.70E+5nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 1.76E+5nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 1.80E+5nMAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 1.92E+5nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 2.39E+5nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 2.80E+5nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 3.05E+5nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 4.00E+5nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 5.90E+5nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 6.30E+5nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 7.10E+5nMAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 8.10E+5nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 1.00E+6nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 1.00E+6nMAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 1.00E+6nMAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 1.00E+6nMAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 1.00E+6nMAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 1.00E+6nMAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 1.72E+6nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 2.00E+6nMpH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 2.00E+6nMAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 2.00E+6nMAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DataIC50: 2.00E+6nMAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DatapH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DatapH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DatapH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DatapH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DatapH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DatapH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DatapH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DatapH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
TargetUDP-N-acetylmuramoylalanine--D-glutamate ligase(Escherichia coli (strain K12))
Lek Pharmaceuticals
Lek Pharmaceuticals
Affinity DatapH: 8.6 T: 2°CAssay Description:The compounds were tested for their ability to inhibit the addition of D-[14C]Glu to UMA. The radioactive substrate and product were separated by re...More data for this Ligand-Target Pair
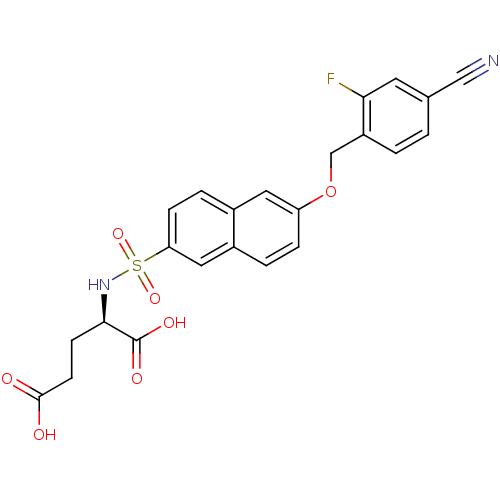
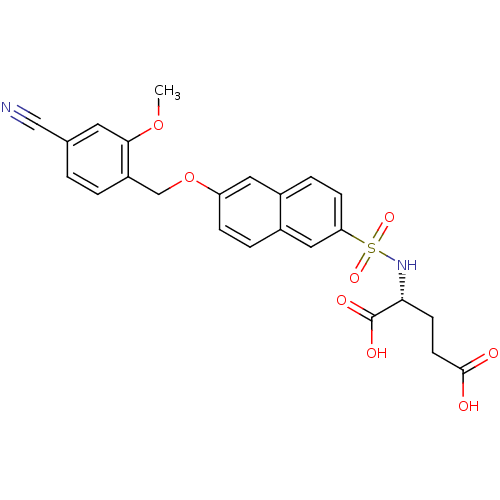

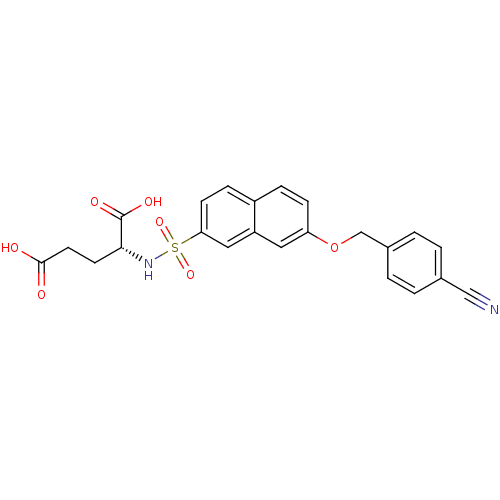
 3D Structure (crystal)
3D Structure (crystal)