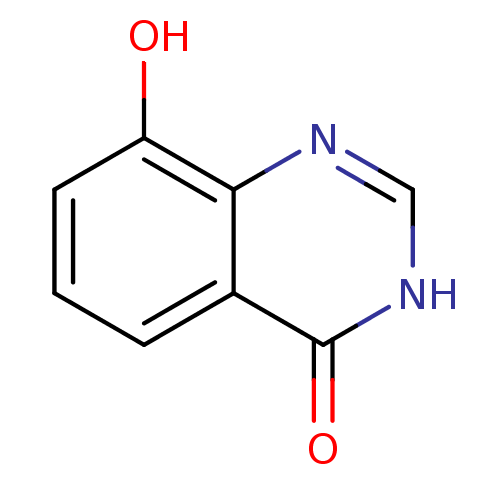
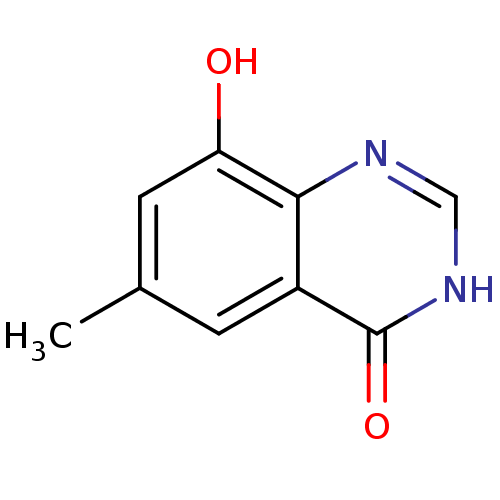
Report error Found 6 Enz. Inhib. hit(s) with all data for entry = 5743
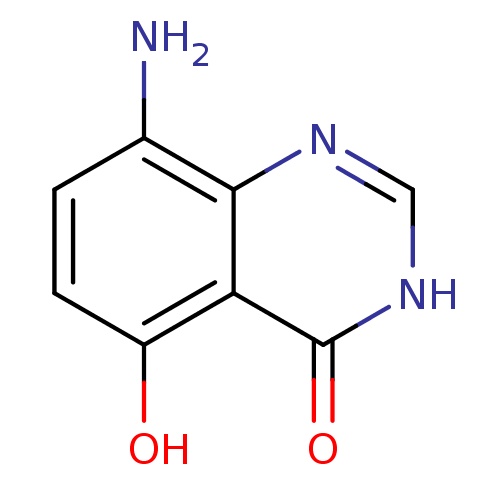
Affinity DataKi: 1.31E+5nMAssay Description:PNPase activity (guanosine to guanine and ribose 1-phosphate) was followed by measuring the decrease in absorbance at 268nm.More data for this Ligand-Target Pair
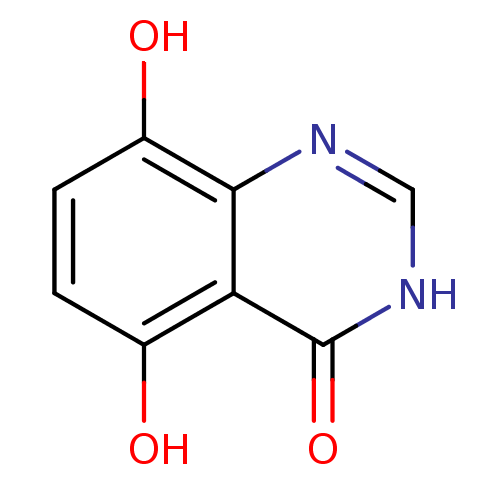
Affinity DataKi: 1.42E+5nMAssay Description:PNPase activity (guanosine to guanine and ribose 1-phosphate) was followed by measuring the decrease in absorbance at 268nm.More data for this Ligand-Target Pair
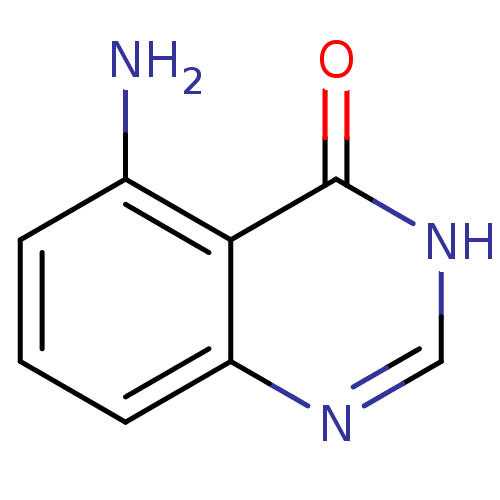
Affinity DataKi: 3.37E+5nMAssay Description:PNPase activity (guanosine to guanine and ribose 1-phosphate) was followed by measuring the decrease in absorbance at 268nm.More data for this Ligand-Target Pair
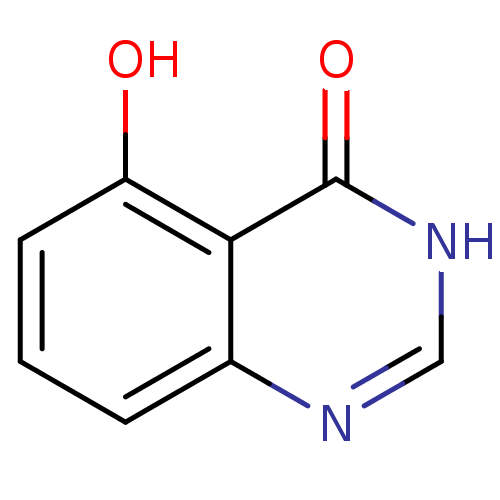
Affinity DataKi: 5.76E+5nMAssay Description:PNPase activity (guanosine to guanine and ribose 1-phosphate) was followed by measuring the decrease in absorbance at 268nm.More data for this Ligand-Target Pair
Affinity DataKi: 5.93E+5nMAssay Description:PNPase activity (guanosine to guanine and ribose 1-phosphate) was followed by measuring the decrease in absorbance at 268nm.More data for this Ligand-Target Pair
Affinity DataKi: 6.18E+5nMAssay Description:PNPase activity (guanosine to guanine and ribose 1-phosphate) was followed by measuring the decrease in absorbance at 268nm.More data for this Ligand-Target Pair