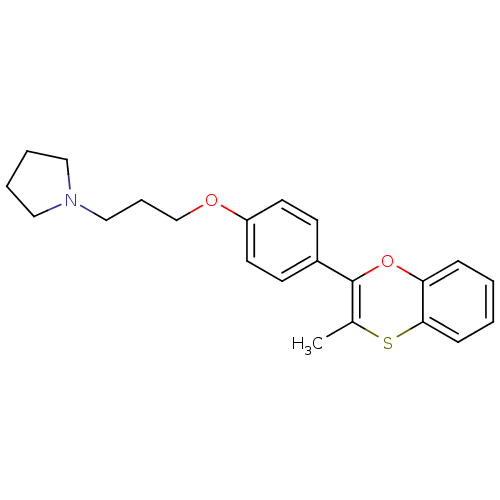
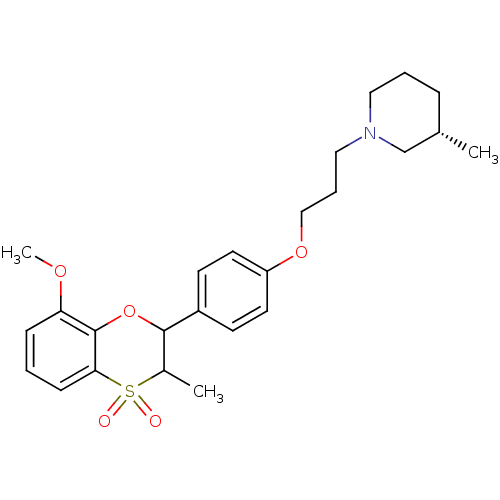
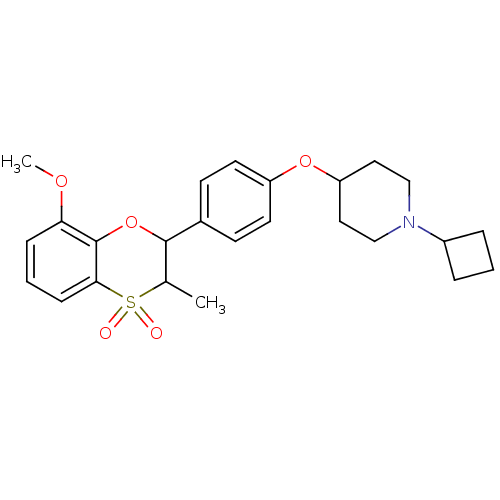
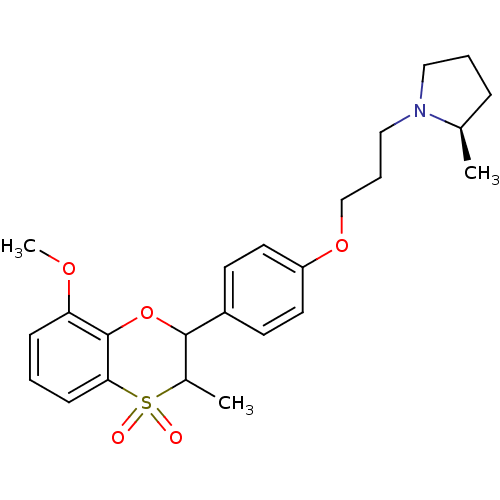
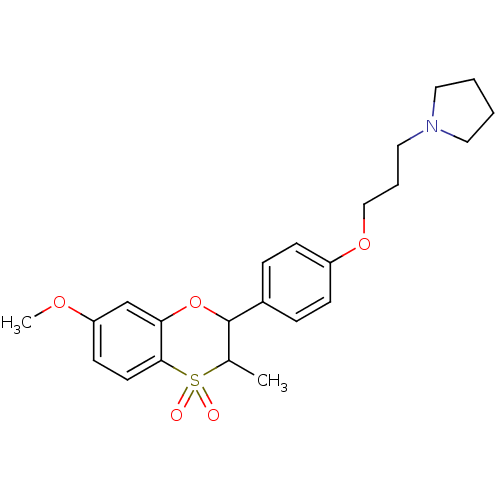
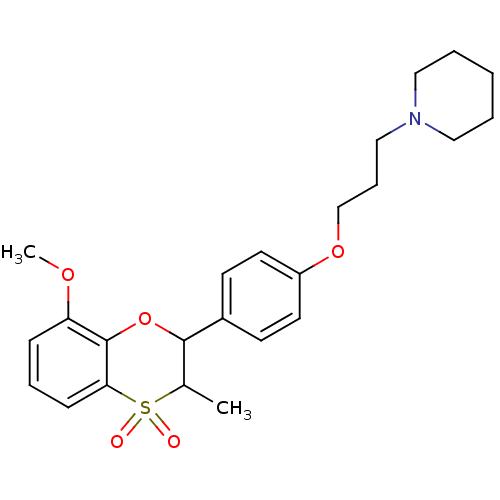
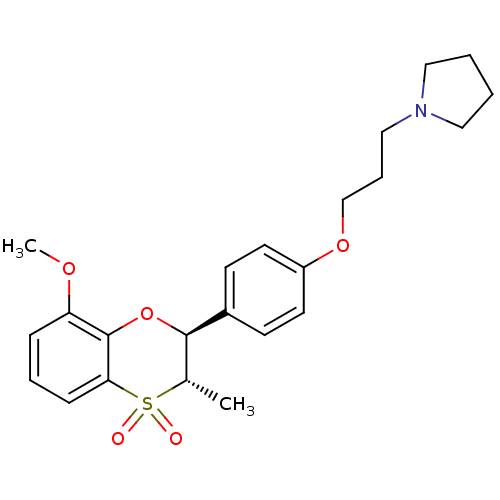
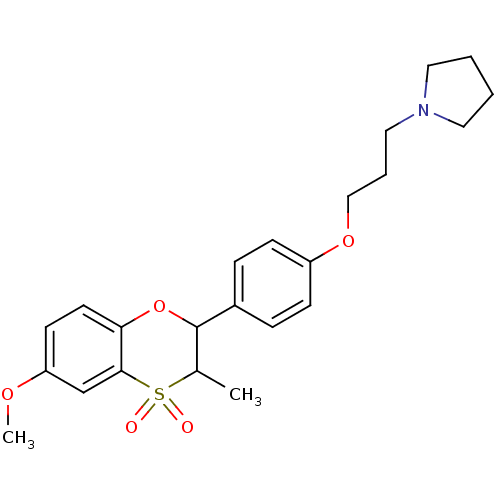
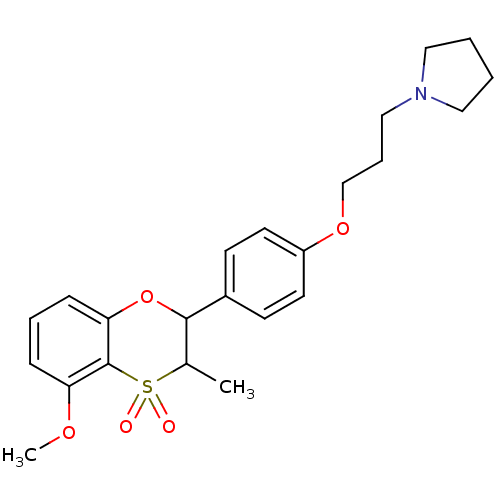
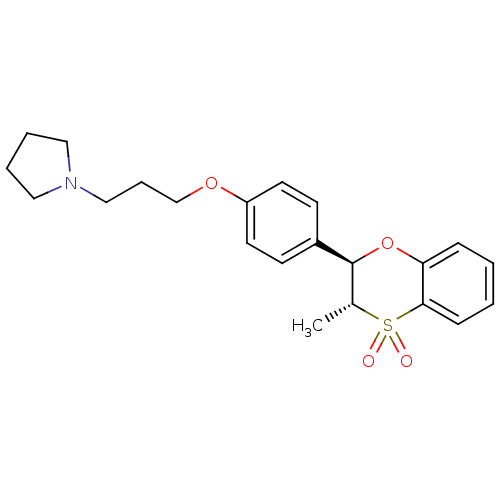
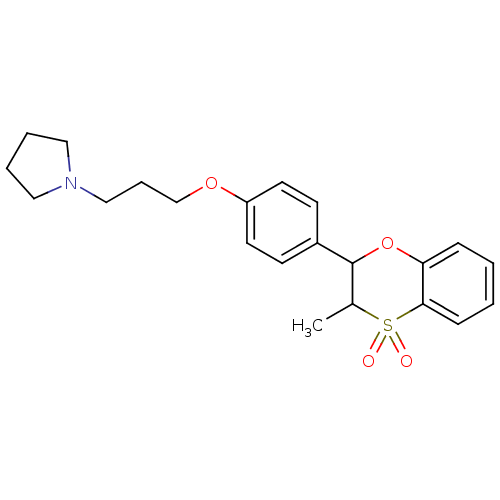
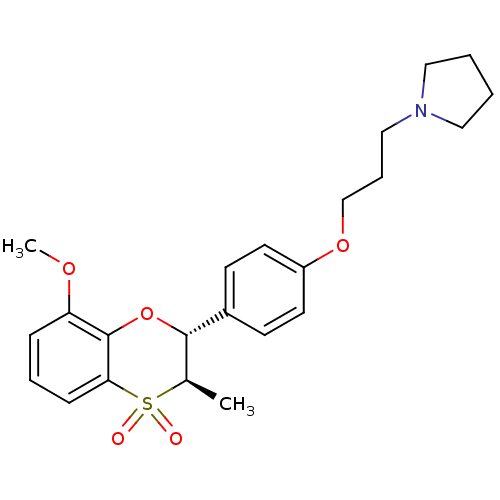
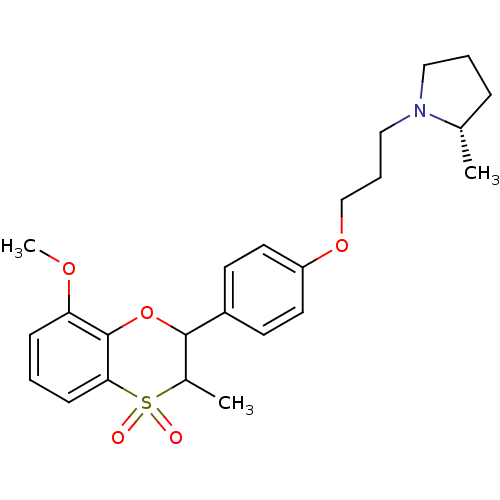
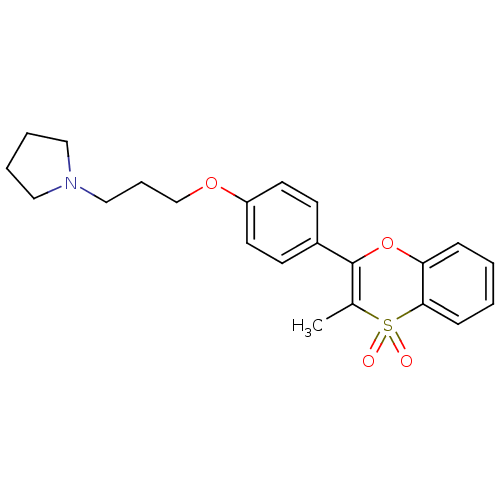
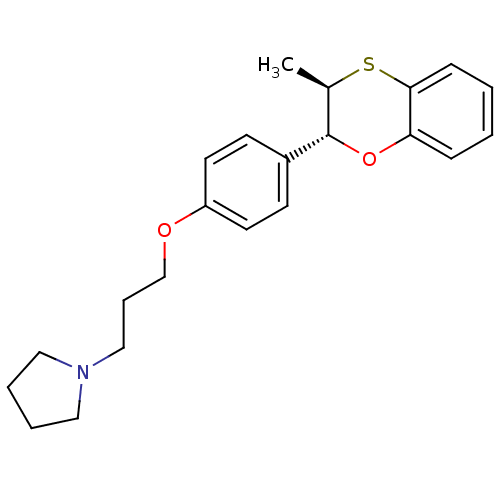
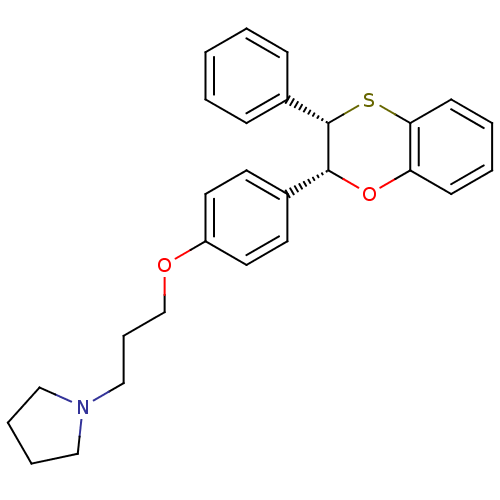
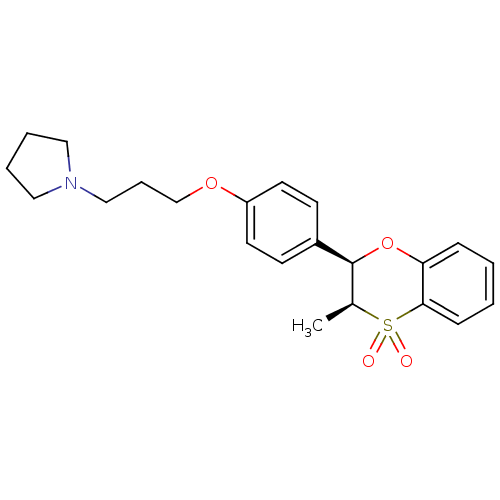
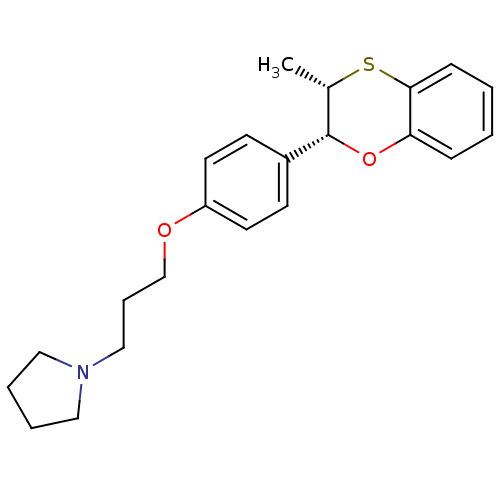
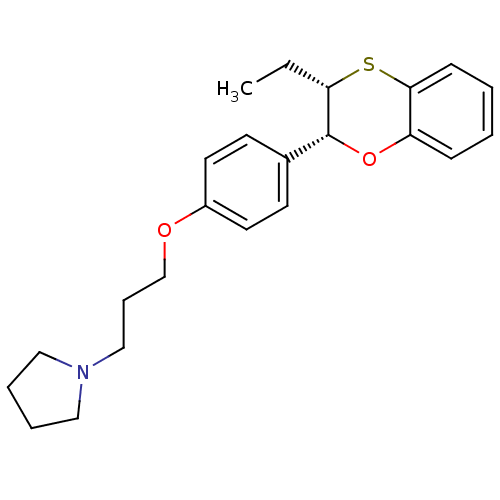
Report error Found 46 Enz. Inhib. hit(s) with all data for entry = 50030438
Affinity DataIC50: 0.160nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
Affinity DataIC50: 0.160nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
Affinity DataIC50: 0.210nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
Affinity DataIC50: 0.320nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
Affinity DataIC50: 0.340nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
Affinity DataIC50: 0.390nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
Affinity DataIC50: 0.430nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
Affinity DataIC50: 0.440nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
Affinity DataIC50: 0.600nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
Affinity DataIC50: 0.600nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
Affinity DataIC50: 0.607nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
Affinity DataIC50: 0.820nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
Affinity DataIC50: 0.840nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
Affinity DataIC50: 3.90nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 4.40nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 5nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 5.90nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
Affinity DataIC50: 6.70nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
Affinity DataIC50: 8.60nMAssay Description:Inverse agonist activity at human cloned histamine H3 receptor assessed as inhibition of R-alpha-methylhistamine-induced [35S]GTPgammaS binding by ce...More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 12nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 25nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 440nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 470nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.30E+3nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.30E+3nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.50E+3nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 2.90E+3nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 3.80E+3nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 4.80E+3nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 4.90E+3nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 6.20E+3nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 6.30E+3nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 7.00E+3nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human histamine H1 receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of CYP3A4More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of CYP2D6More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human histamine H2 receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of human histamine H4 receptorMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of CYP2C9More data for this Ligand-Target Pair
TargetPotassium voltage-gated channel subfamily H member 2(Human)
Tsukuba Research Institute
Curated by ChEMBL
Tsukuba Research Institute
Curated by ChEMBL
Affinity DataIC50: 1.40E+4nMAssay Description:Inhibition of [35S]MK499 binding to human ERG transfected in HEK293 cells by microscintillation countingMore data for this Ligand-Target Pair