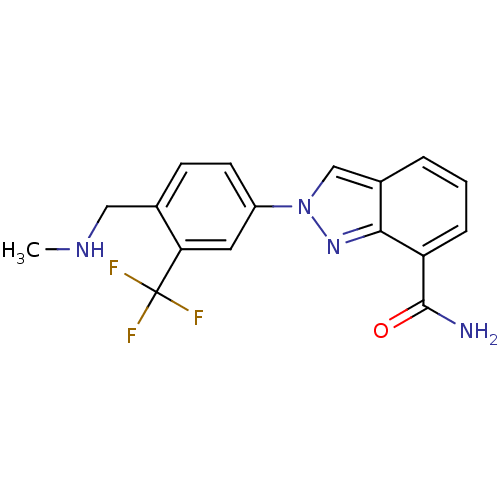
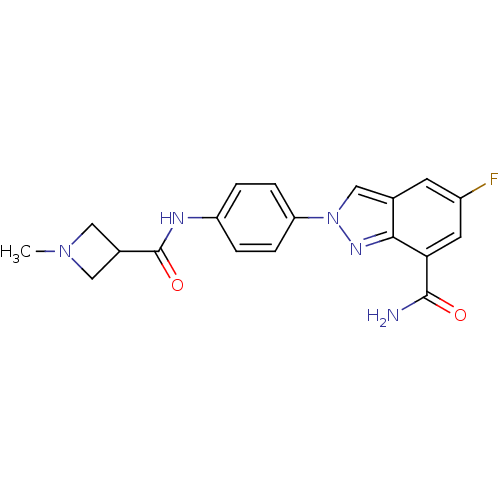
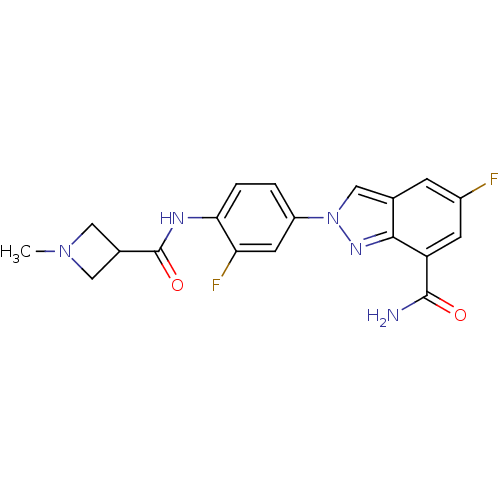
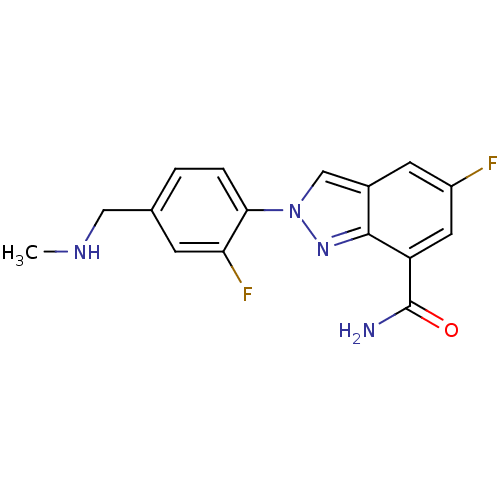
Report error Found 45 Enz. Inhib. hit(s) with all data for entry = 50031057
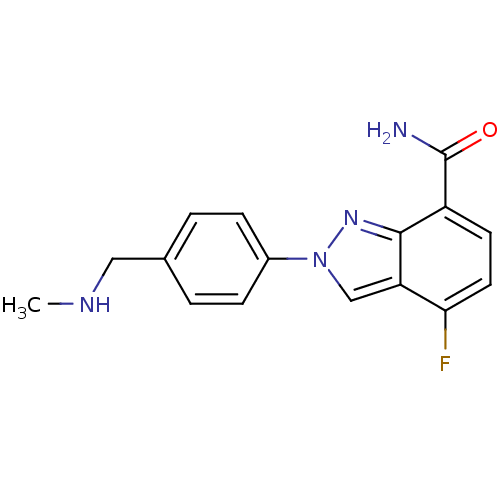
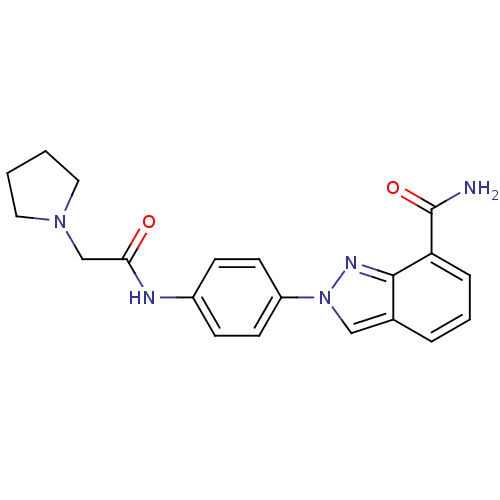
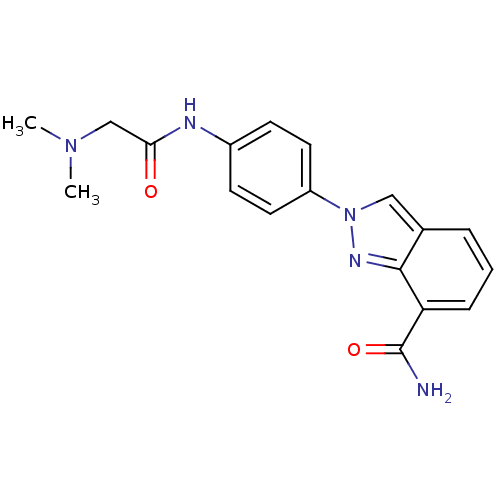
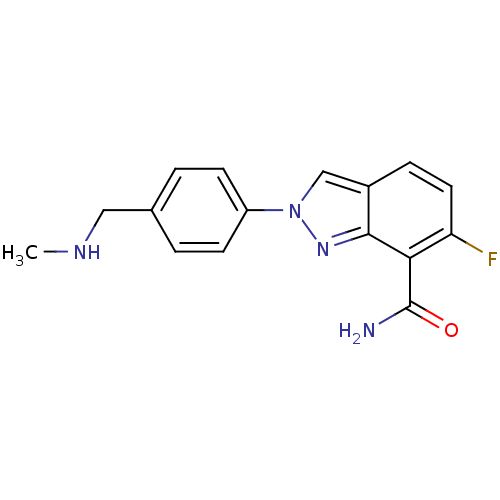
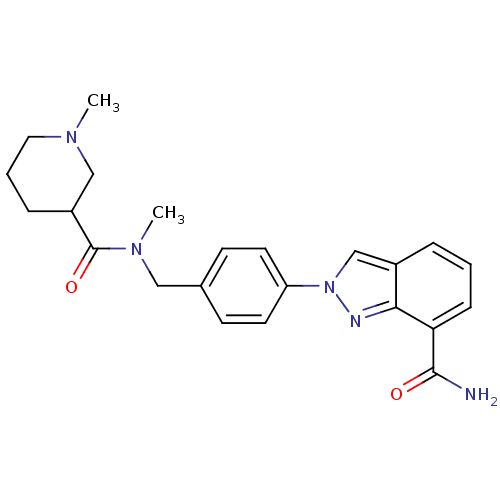
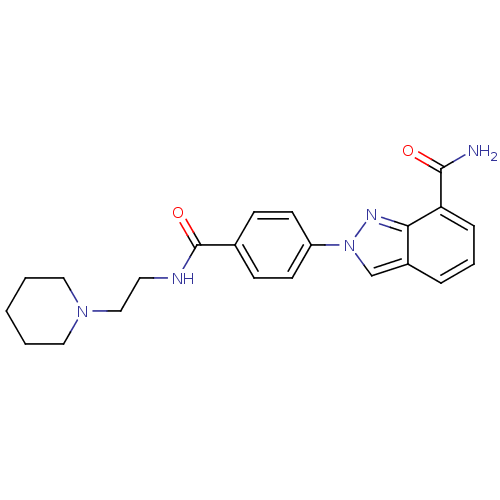
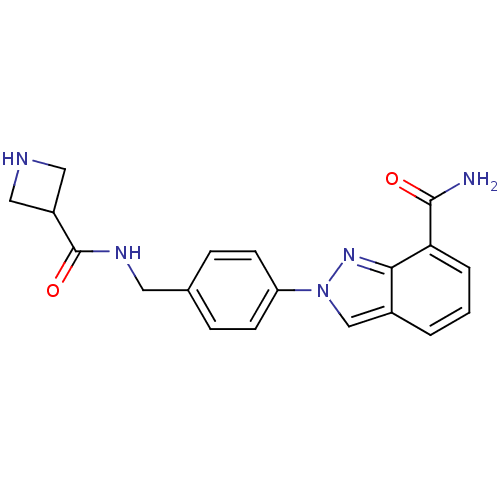
Affinity DataIC50: 1.40nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
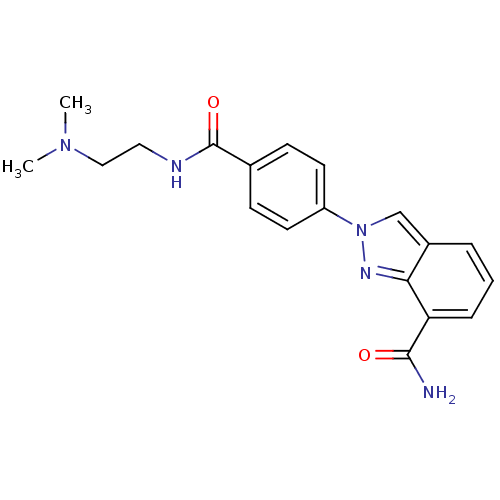
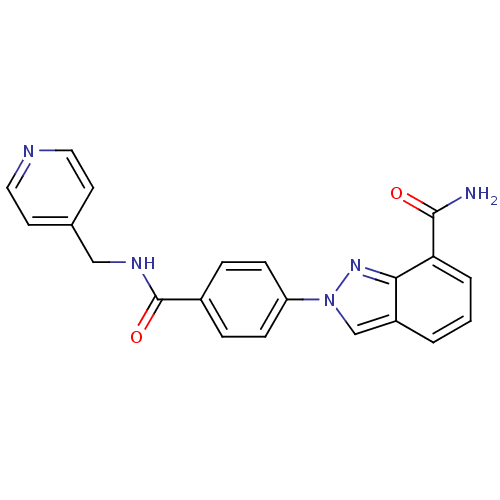
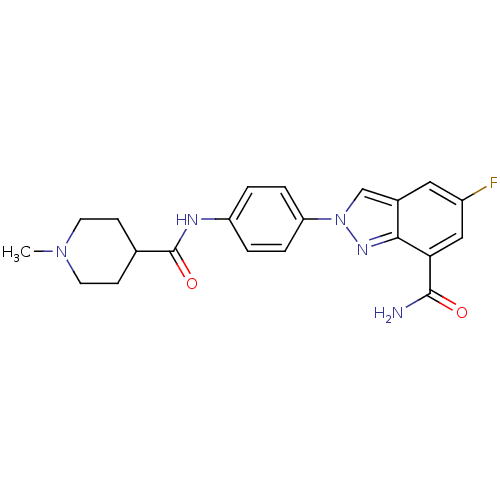
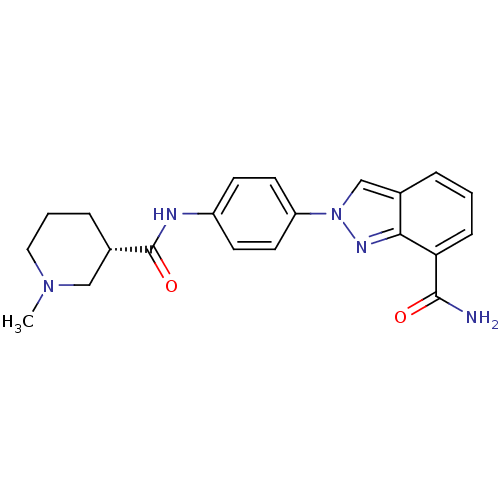
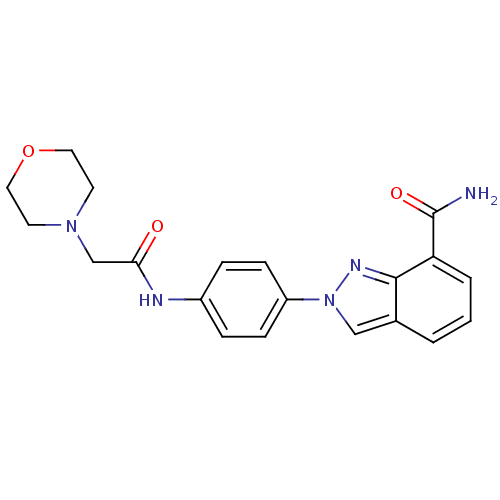
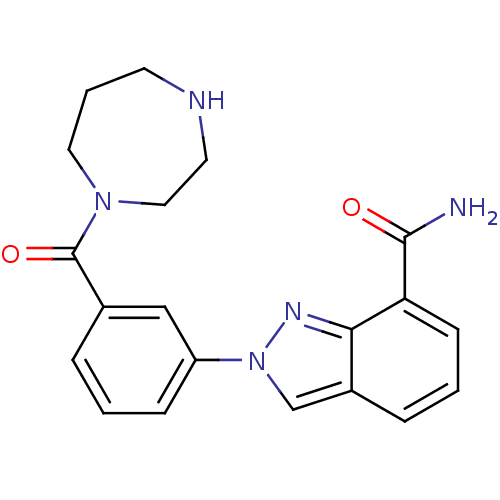
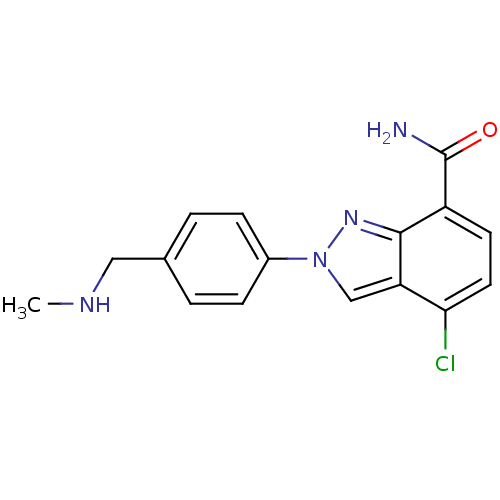
Affinity DataIC50: 2nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
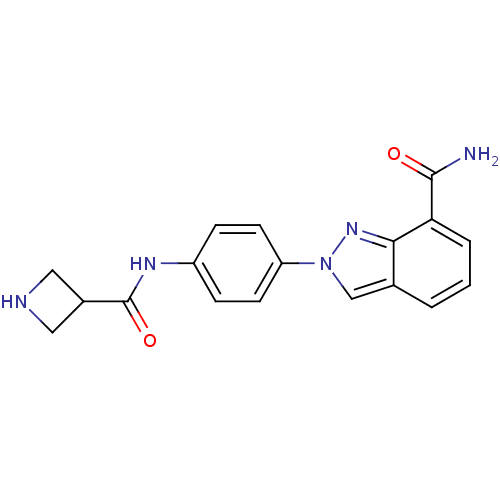
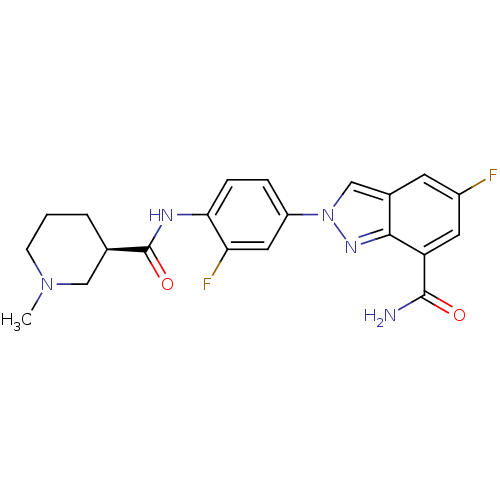
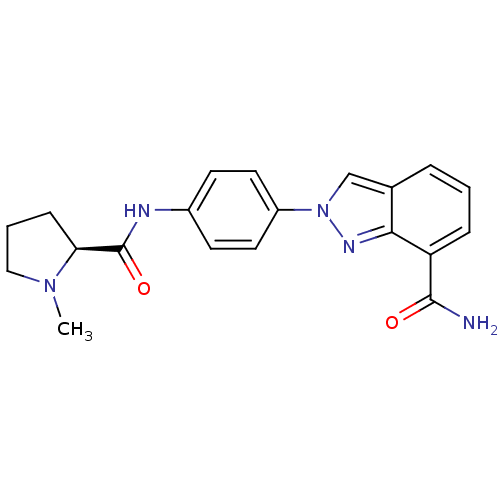
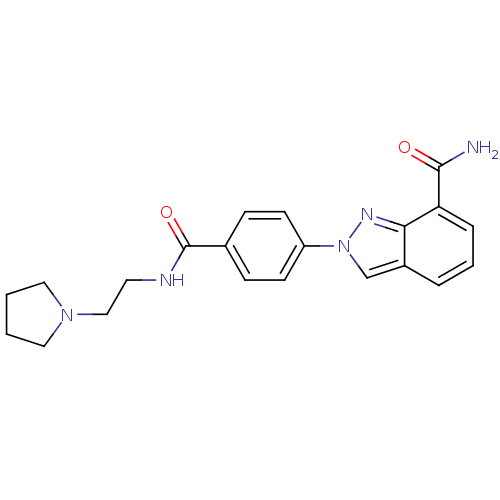
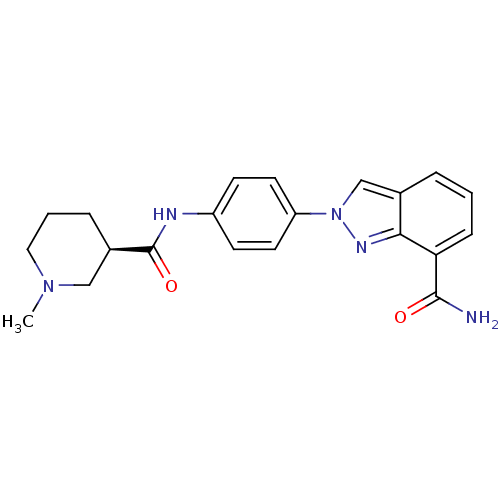
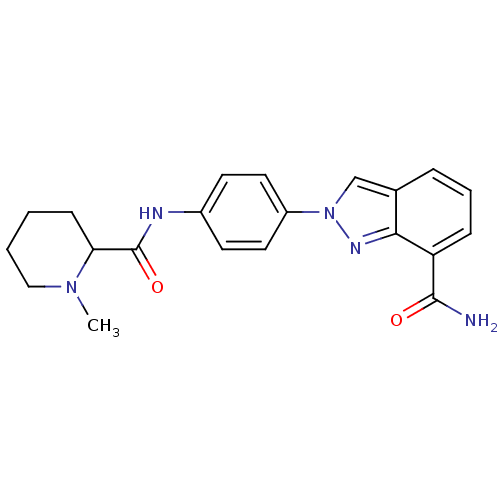
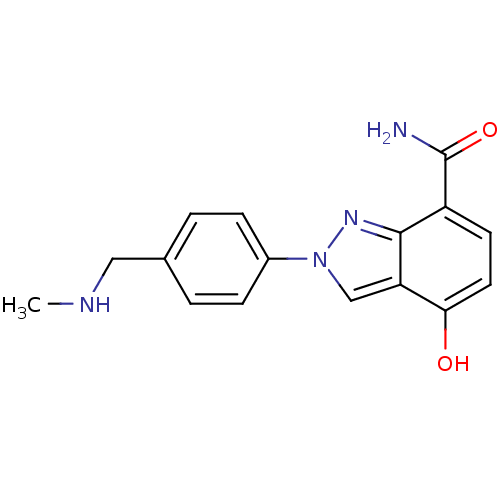
Affinity DataIC50: 4nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
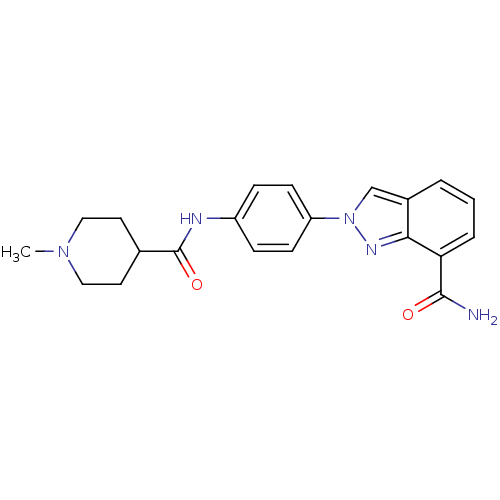
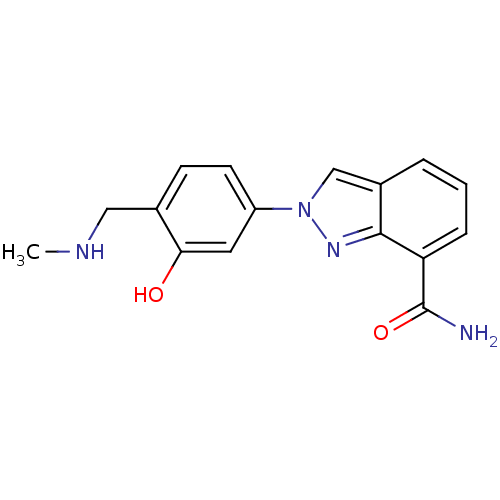
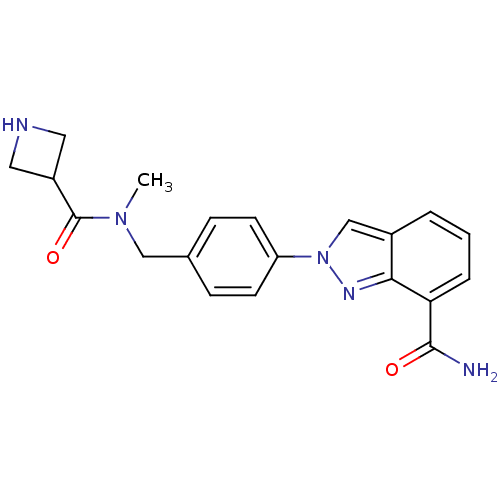
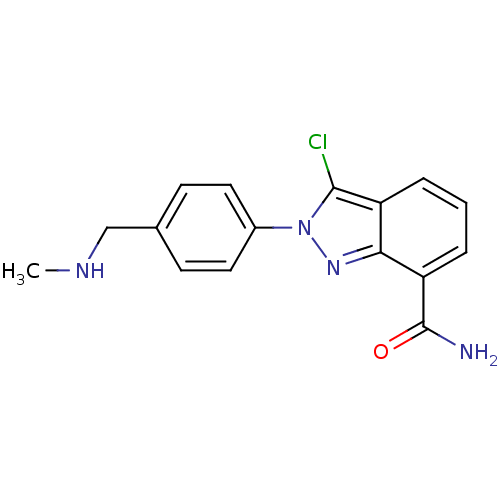
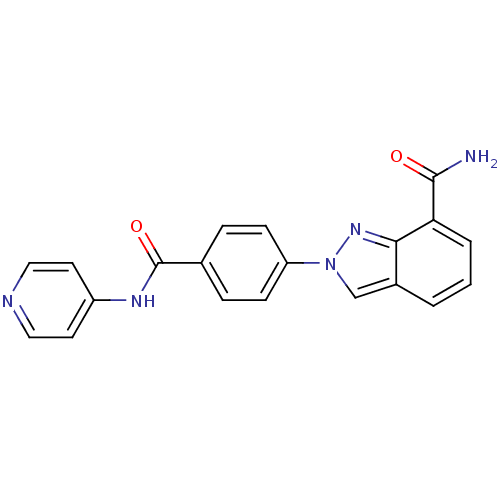
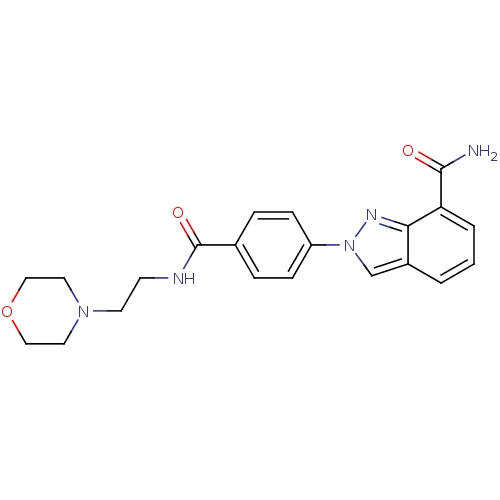
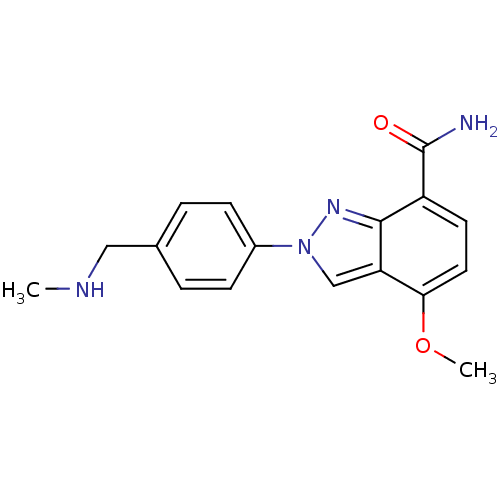
Affinity DataIC50: 4nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 15nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 16nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 17nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 25nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 25nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 25nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 38nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 46nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 54nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 65nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 180nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 540nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+3nMAssay Description:Inhibition of human PARP1 after 3 hrs using [3H]NAD+ by scintillation proximity assayMore data for this Ligand-Target Pair