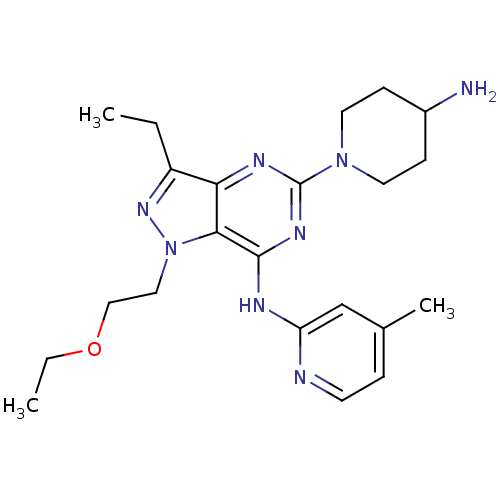
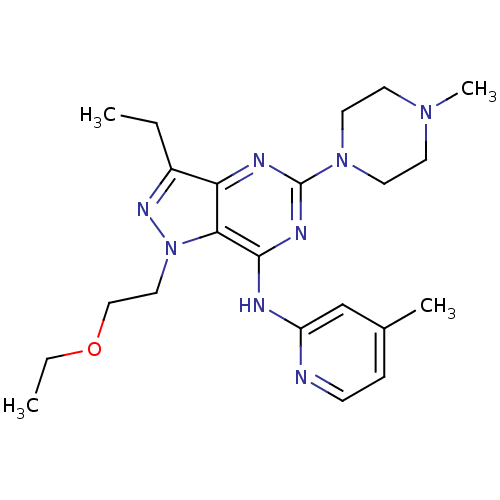
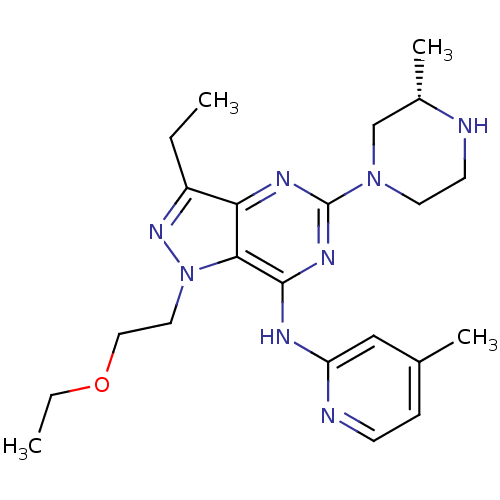
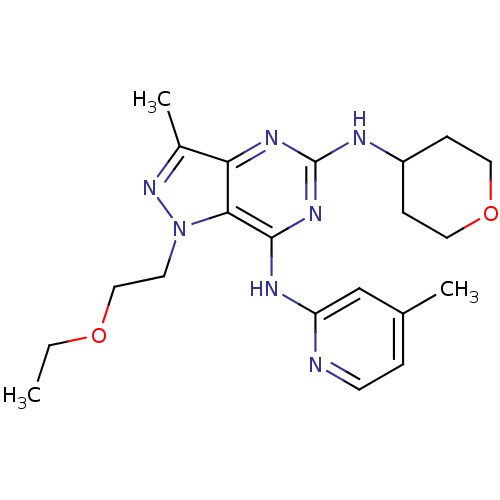
Report error Found 35 Enz. Inhib. hit(s) with all data for entry = 50039138
Affinity DataIC50: 0.00700nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0100nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0400nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0500nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0700nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0700nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0700nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0800nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0800nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.0900nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.190nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.200nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.350nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.380nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.400nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.420nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.5nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.650nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.660nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.720nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.760nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.780nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 0.890nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.20nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.40nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.30nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.30nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.5nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 2.60nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 3.30nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 5.5nMAssay Description:Inhibition of PDE5 in human platelet by assessed as hydrolysis of [3H]cGMP scintillation proximity assayMore data for this Ligand-Target Pair
Affinity DataIC50: 1.10E+3nMAssay Description:Inhibition of human ERG by patch clamp electrophysiologyMore data for this Ligand-Target Pair
Affinity DataIC50: 2.80E+3nMAssay Description:Inhibition of human ERG by patch clamp electrophysiologyMore data for this Ligand-Target Pair