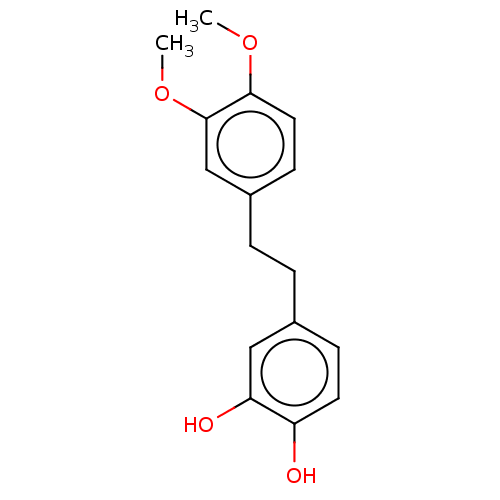
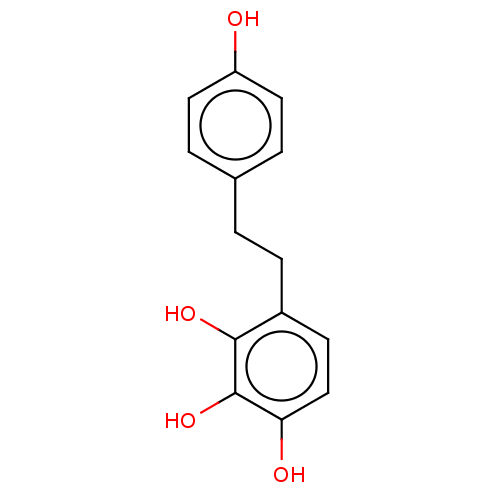
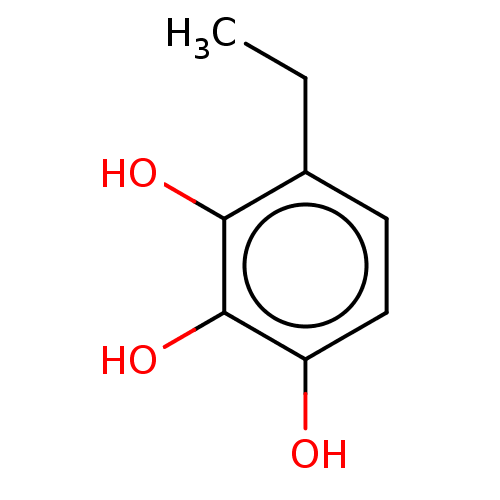
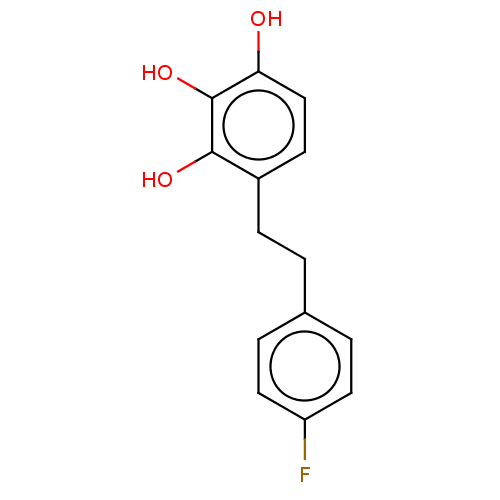
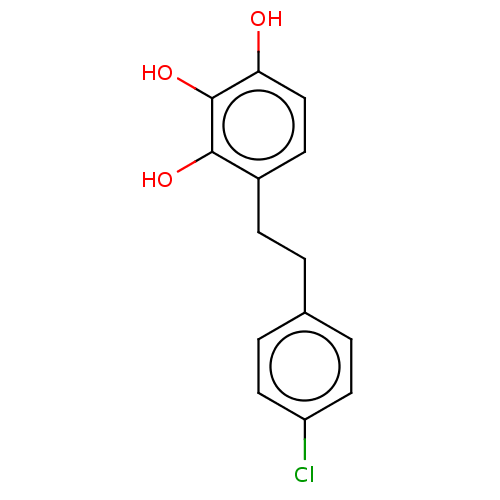
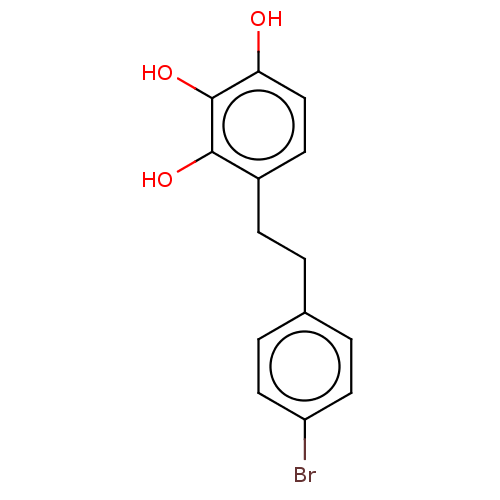
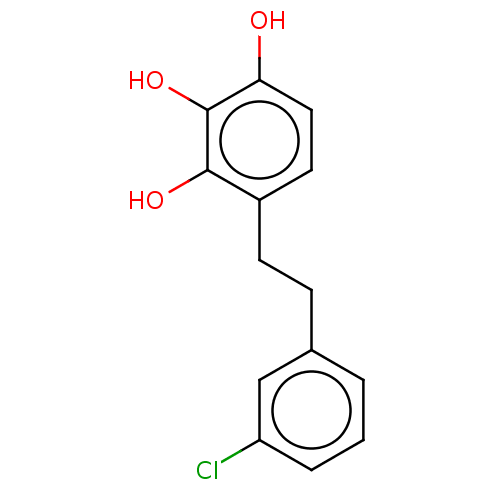
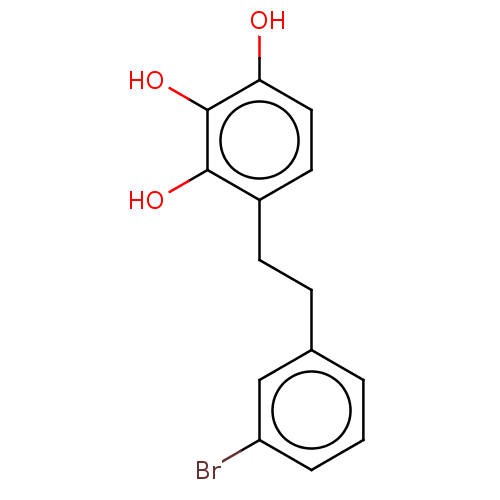
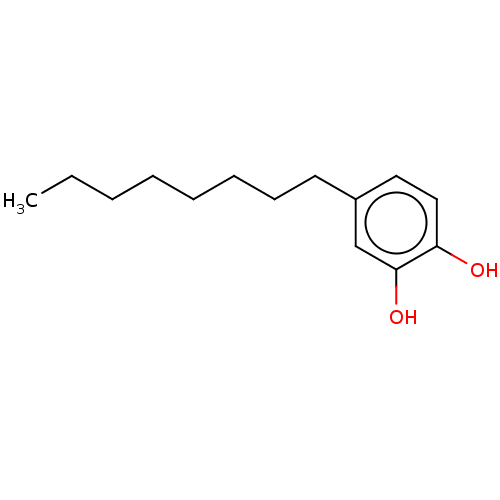
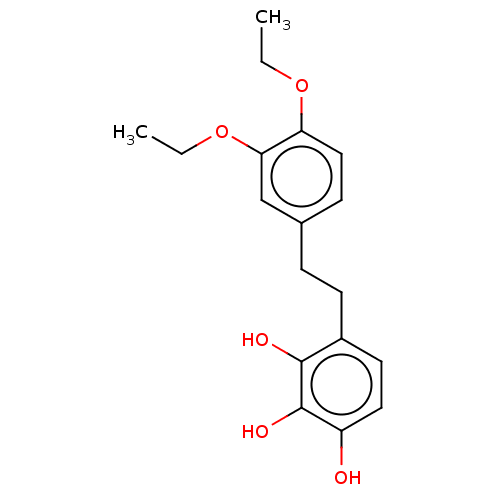
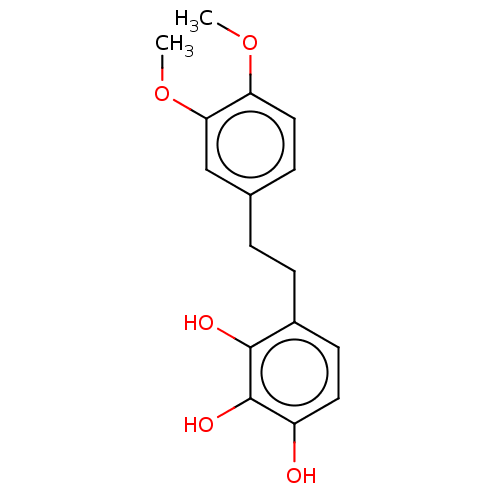
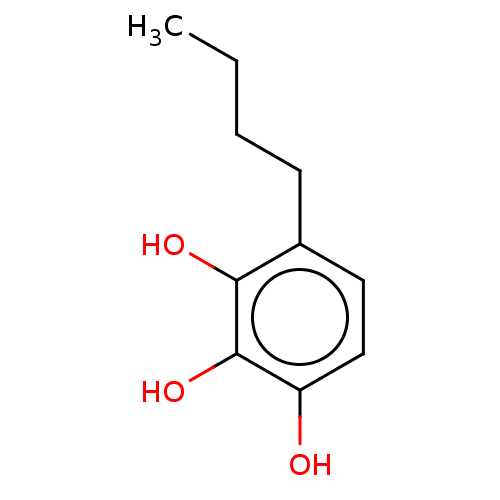
Report error Found 39 Enz. Inhib. hit(s) with all data for entry = 50004449
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
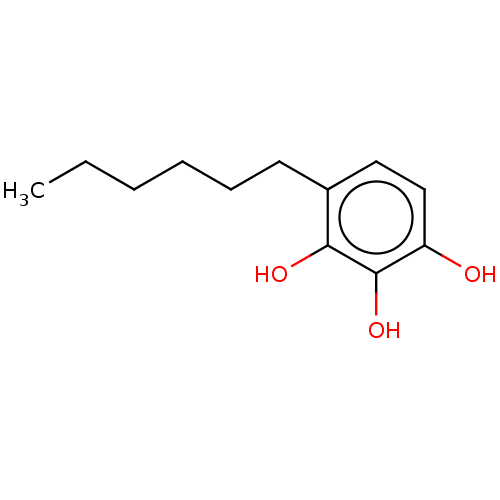
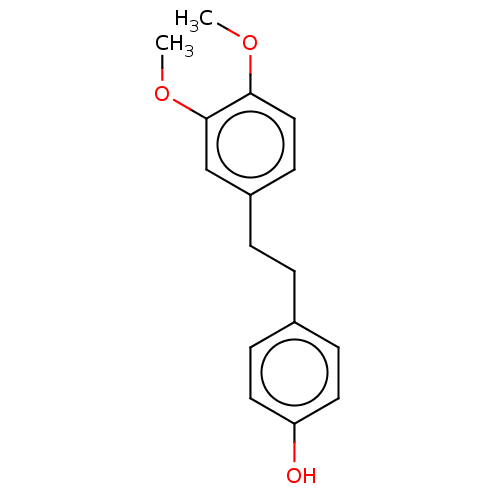
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
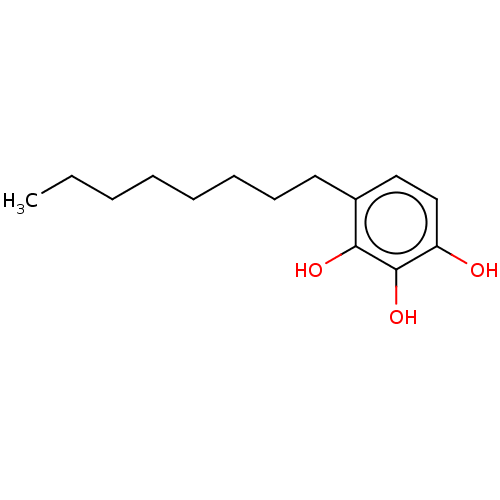
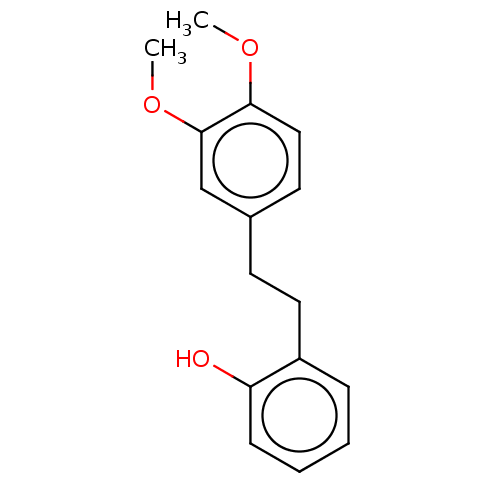
Affinity DataIC50: 4.20E+3nMAssay Description:Inhibition of urease in intact Helicobacter pylori ATCC 43504 assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
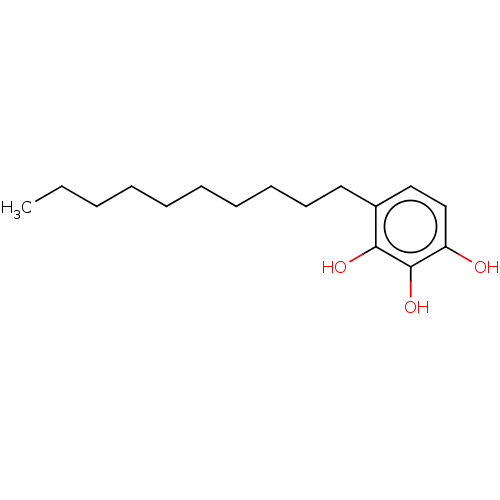
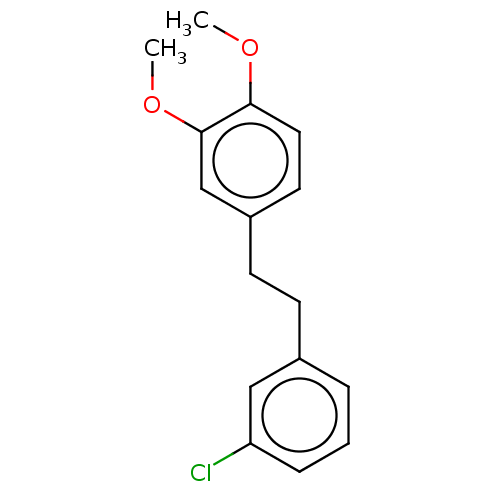
Affinity DataIC50: 4.90E+3nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
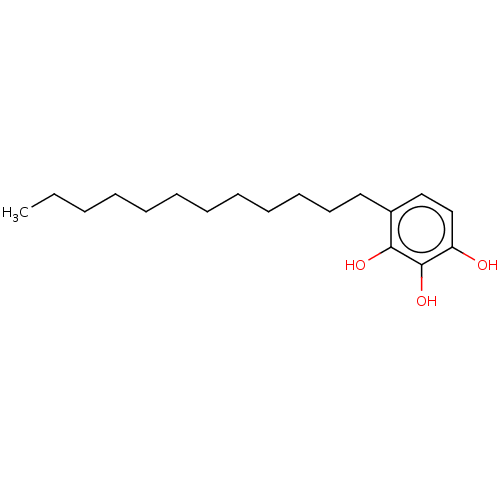
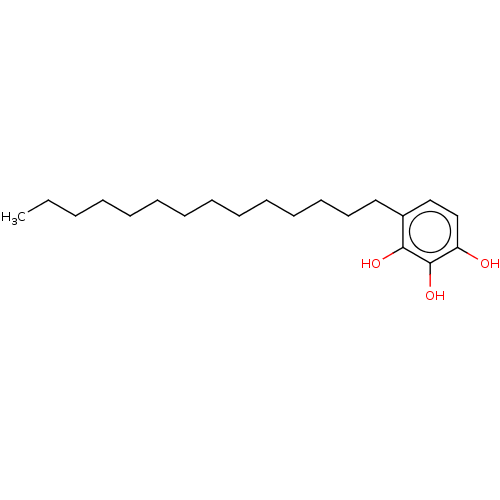
Affinity DataIC50: 6.20E+3nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 7.90E+3nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 8.30E+3nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 1.23E+4nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 1.57E+4nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 1.69E+4nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 1.72E+4nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 1.73E+4nMAssay Description:Inhibition of urease in intact Helicobacter pylori ATCC 43504 assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 2.27E+4nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 3.20E+4nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 3.24E+4nMAssay Description:Inhibition of urease in intact Helicobacter pylori ATCC 43504 assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 3.40E+4nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 3.81E+4nMAssay Description:Inhibition of urease in intact Helicobacter pylori ATCC 43504 assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 4.30E+4nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 4.70E+4nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 4.80E+4nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 5.01E+4nMAssay Description:Inhibition of urease in intact Helicobacter pylori ATCC 43504 assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 5.20E+4nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 5.80E+4nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 6.90E+4nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 7.10E+4nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 7.78E+4nMAssay Description:Inhibition of urease in intact Helicobacter pylori ATCC 43504 assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 7.90E+4nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 8.89E+4nMAssay Description:Inhibition of urease in intact Helicobacter pylori ATCC 43504 assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 9.10E+4nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 9.47E+4nMAssay Description:Inhibition of urease in intact Helicobacter pylori ATCC 43504 assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 1.01E+5nMAssay Description:Inhibition of urease in intact Helicobacter pylori ATCC 43504 assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 1.03E+5nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 1.24E+5nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 1.68E+5nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 2.04E+5nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 2.27E+5nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 2.35E+5nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 2.50E+5nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
TargetUrease subunit beta(Helicobacter pylori (strain ATCC 700392 / 26695) (...)
Jishou University
Curated by ChEMBL
Jishou University
Curated by ChEMBL
Affinity DataIC50: 2.54E+5nMAssay Description:Inhibition of cell free Helicobacter pylori ATCC 43504 urease assessed as reduction in ammonia production by indophenol based Berthelot color reactio...More data for this Ligand-Target Pair
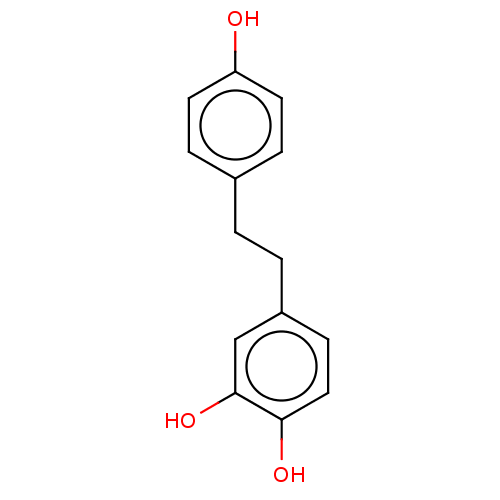
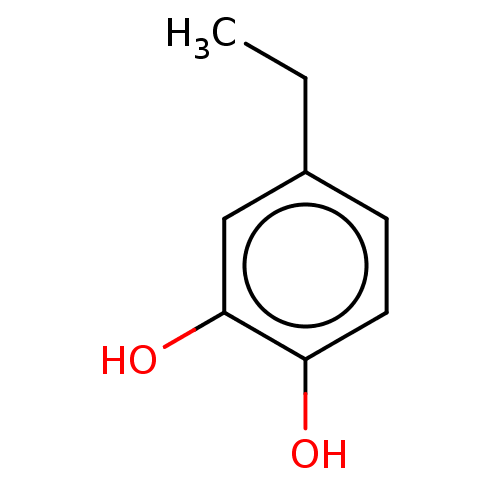
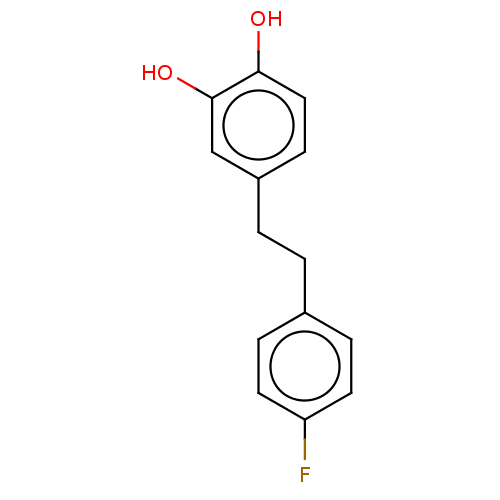
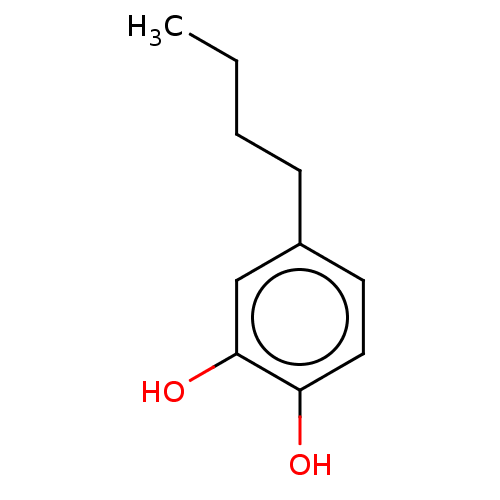
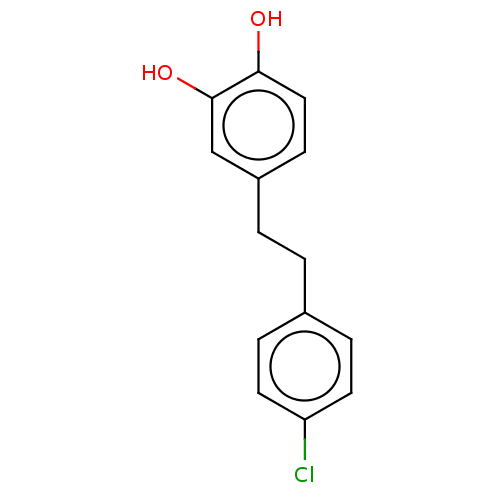
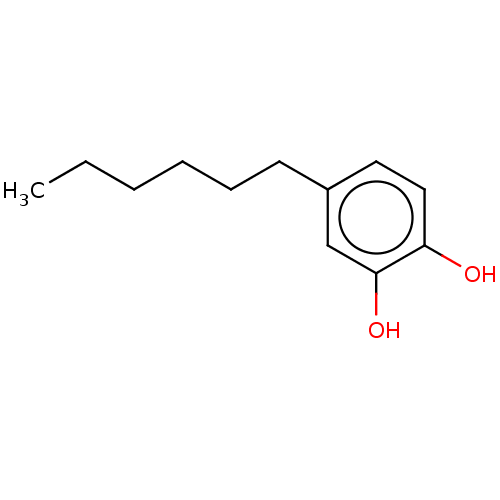
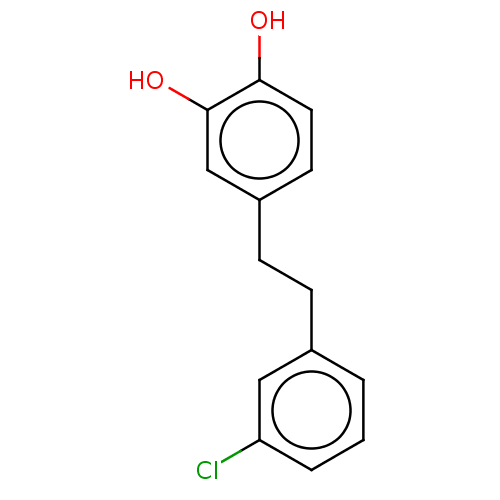
 3D Structure (crystal)
3D Structure (crystal)