Report error Found 11 Enz. Inhib. hit(s) with all data for entry = 8110
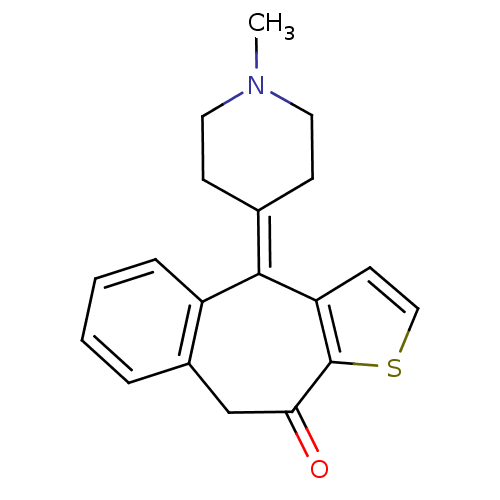
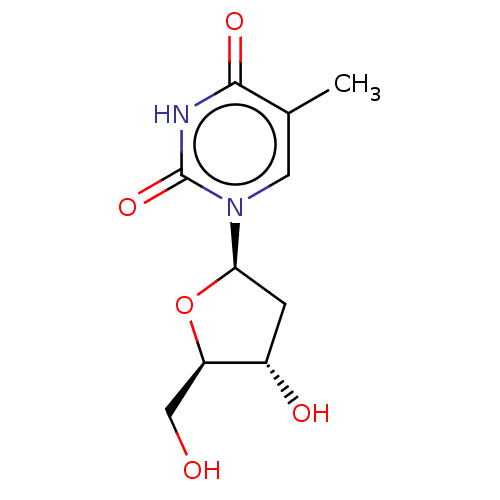
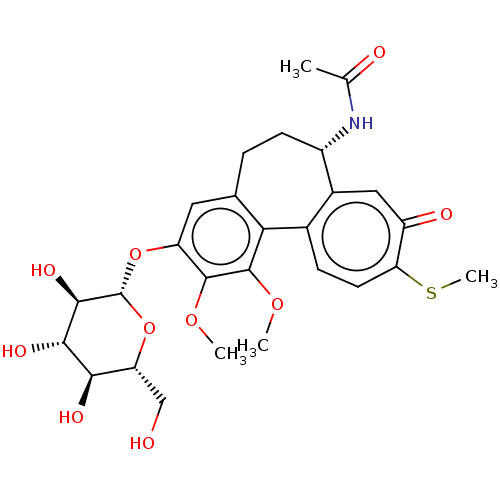
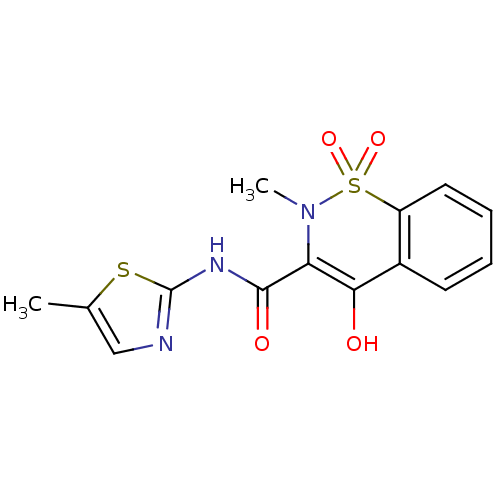
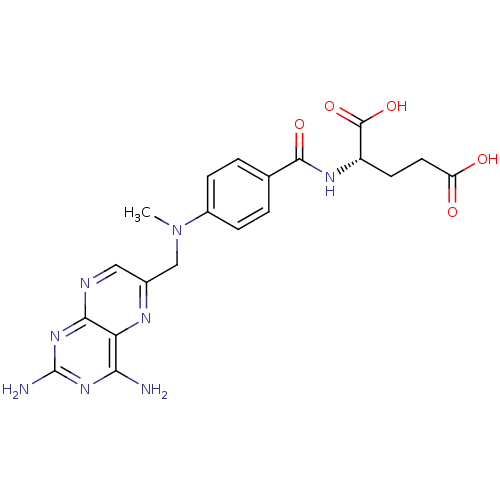
Affinity DataKi: 5.20E+3nM IC50: 8.00E+3nMAssay Description:Ketotifen, dacarbazine, thiocolchicoside, meloxicam, methotrexate, furosemide, olanzapine, methylprednizolone acetate, paricalcitol, ritodrine hydroc...More data for this Ligand-Target Pair
Affinity DataKi: 1.54E+4nM IC50: 2.10E+4nMAssay Description:Ketotifen, dacarbazine, thiocolchicoside, meloxicam, methotrexate, furosemide, olanzapine, methylprednizolone acetate, paricalcitol, ritodrine hydroc...More data for this Ligand-Target Pair
Affinity DataKi: 3.97E+4nM IC50: 5.60E+4nMAssay Description:Ketotifen, dacarbazine, thiocolchicoside, meloxicam, methotrexate, furosemide, olanzapine, methylprednizolone acetate, paricalcitol, ritodrine hydroc...More data for this Ligand-Target Pair
Affinity DataKi: 5.79E+4nM IC50: 6.70E+4nMAssay Description:Ketotifen, dacarbazine, thiocolchicoside, meloxicam, methotrexate, furosemide, olanzapine, methylprednizolone acetate, paricalcitol, ritodrine hydroc...More data for this Ligand-Target Pair
Affinity DataKi: 9.10E+4nM IC50: 1.14E+5nMAssay Description:Ketotifen, dacarbazine, thiocolchicoside, meloxicam, methotrexate, furosemide, olanzapine, methylprednizolone acetate, paricalcitol, ritodrine hydroc...More data for this Ligand-Target Pair
Affinity DataKi: 1.87E+5nM IC50: 3.15E+5nMAssay Description:Ketotifen, dacarbazine, thiocolchicoside, meloxicam, methotrexate, furosemide, olanzapine, methylprednizolone acetate, paricalcitol, ritodrine hydroc...More data for this Ligand-Target Pair
Affinity DataKi: 4.97E+5nM IC50: 5.30E+5nMAssay Description:Ketotifen, dacarbazine, thiocolchicoside, meloxicam, methotrexate, furosemide, olanzapine, methylprednizolone acetate, paricalcitol, ritodrine hydroc...More data for this Ligand-Target Pair
Affinity DataKi: 5.41E+5nM IC50: 2.23E+6nMAssay Description:Ketotifen, dacarbazine, thiocolchicoside, meloxicam, methotrexate, furosemide, olanzapine, methylprednizolone acetate, paricalcitol, ritodrine hydroc...More data for this Ligand-Target Pair
Affinity DataKi: 4.29E+6nM IC50: 2.28E+6nMAssay Description:Ketotifen, dacarbazine, thiocolchicoside, meloxicam, methotrexate, furosemide, olanzapine, methylprednizolone acetate, paricalcitol, ritodrine hydroc...More data for this Ligand-Target Pair
Affinity DataKi: 8.21E+6nM IC50: 8.11E+6nMAssay Description:Ketotifen, dacarbazine, thiocolchicoside, meloxicam, methotrexate, furosemide, olanzapine, methylprednizolone acetate, paricalcitol, ritodrine hydroc...More data for this Ligand-Target Pair
Affinity DataKi: 4.84E+7nM IC50: 1.15E+8nMAssay Description:Ketotifen, dacarbazine, thiocolchicoside, meloxicam, methotrexate, furosemide, olanzapine, methylprednizolone acetate, paricalcitol, ritodrine hydroc...More data for this Ligand-Target Pair