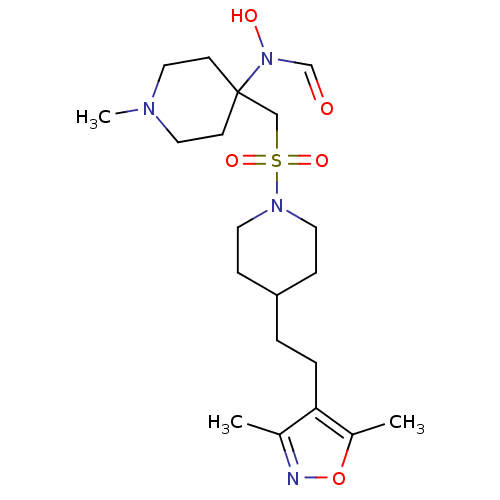
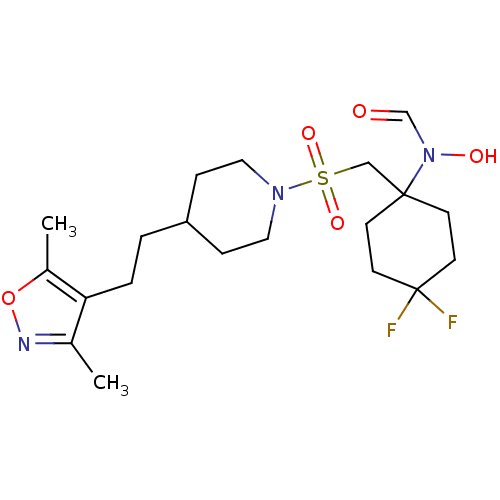
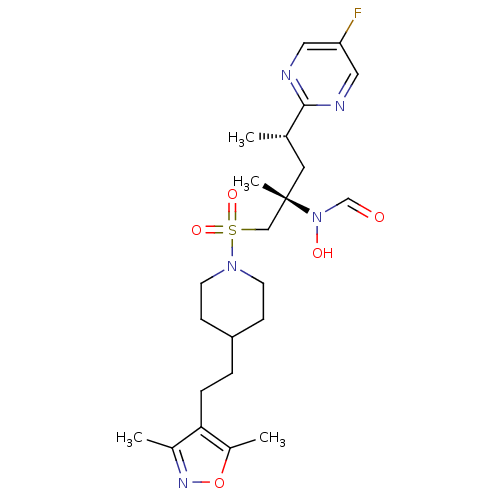
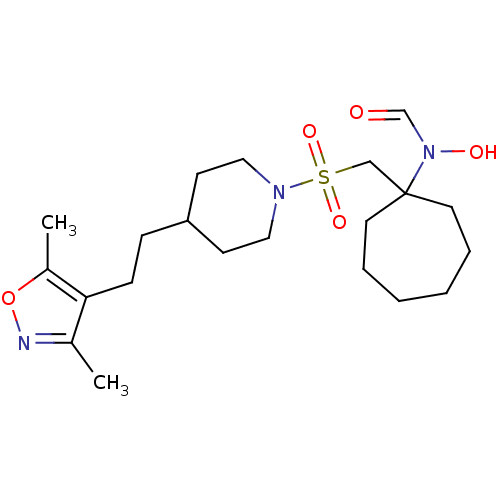
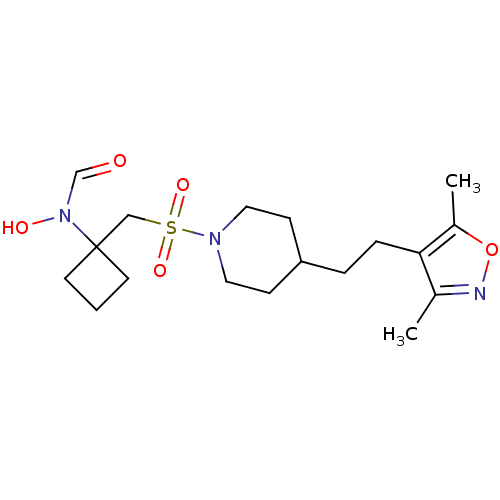
Report error Found 219 Enz. Inhib. hit(s) with all data for entry = 50033446
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
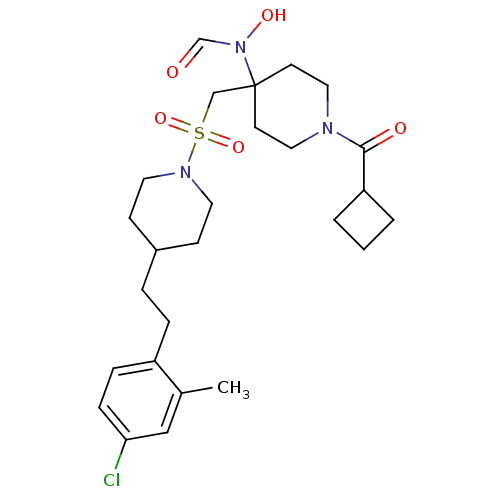
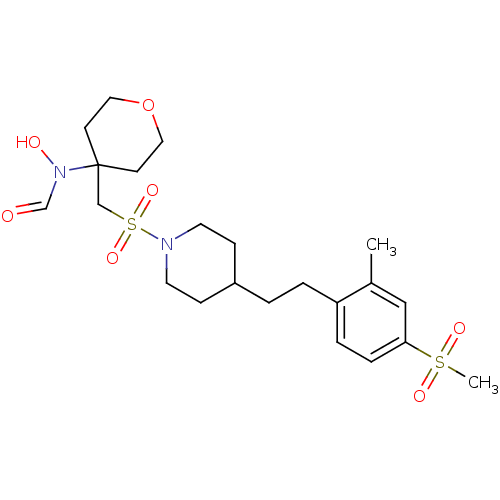
Affinity DataIC50: 1.20nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
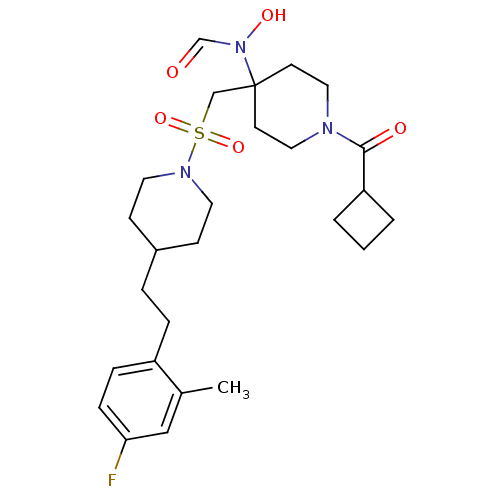
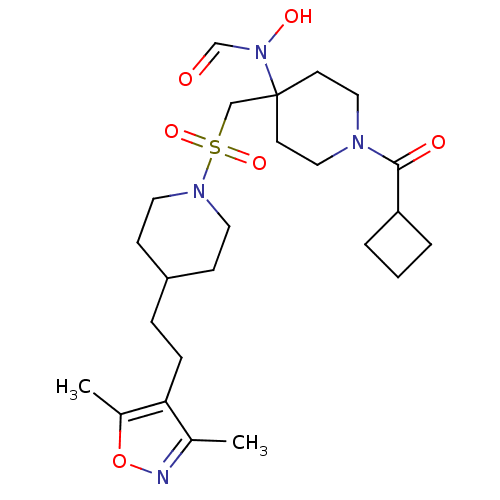
Affinity DataIC50: 1.20nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
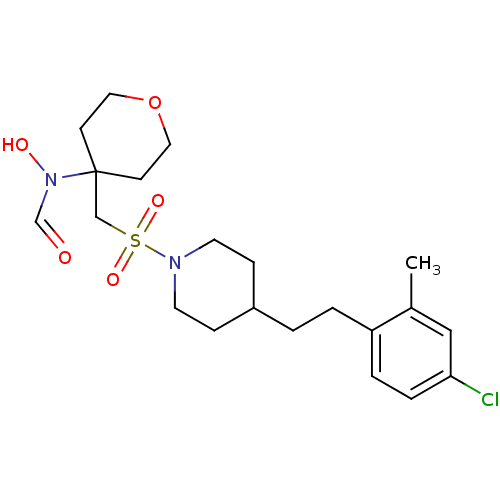
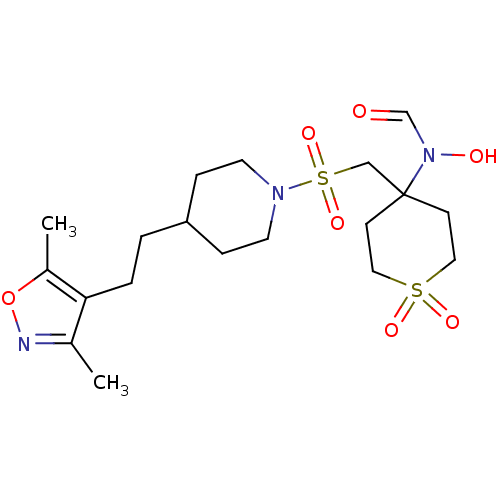
Affinity DataIC50: 1.40nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
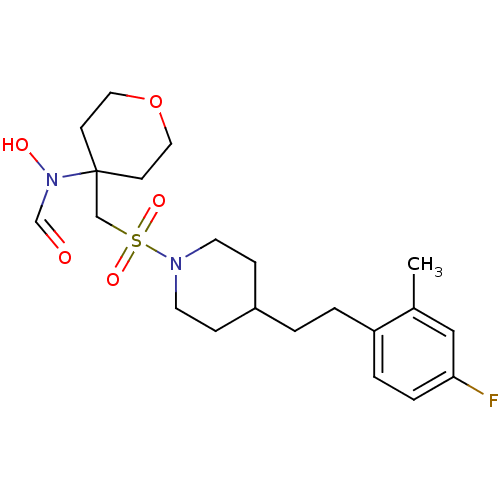
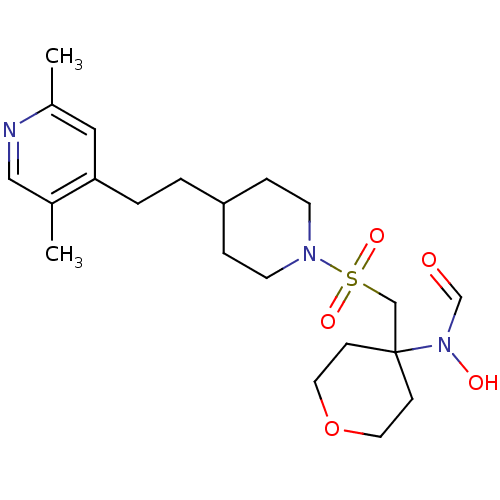
Affinity DataIC50: 1.90nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 2.70nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 4.10nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 4.5nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 5.60nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 5.80nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 8.70nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 9.70nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 10nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 5(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of ADAMTS-5 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 11nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 13nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 5(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 13nMAssay Description:Inhibition of ADAMTS-5 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 16nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 18nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 19nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 20nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 20nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 5(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 20nMAssay Description:Inhibition of ADAMTS-5 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 21nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 5(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 23nMAssay Description:Inhibition of ADAMTS-5 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 24nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 26nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 29nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 5(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 31nMAssay Description:Inhibition of ADAMTS-5 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 48nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 51nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 52nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 5(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 61nMAssay Description:Inhibition of ADAMTS-5 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 5(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 65nMAssay Description:Inhibition of ADAMTS-5 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 5(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 69nMAssay Description:Inhibition of ADAMTS-5 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 5(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 79nMAssay Description:Inhibition of ADAMTS-5 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 91nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 92nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 100nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 110nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 130nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Inhibition of recombinant human MMP-2 assessed as substrate cleavage after 20 hrs by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 154nMAssay Description:Inhibition of recombinant human MMP-13 assessed as substrate cleavage after 20 hrs by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 226nMAssay Description:Inhibition of recombinant human MMP-14 assessed as substrate cleavage after 20 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 5(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 250nMAssay Description:Inhibition of ADAMTS-5 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataIC50: 260nMAssay Description:Inhibition of recombinant human MMP-13 assessed as substrate cleavage after 20 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 5(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 270nMAssay Description:Inhibition of ADAMTS-5 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 5(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 270nMAssay Description:Inhibition of ADAMTS-5 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 5(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 310nMAssay Description:Inhibition of ADAMTS-5 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 380nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair
TargetA disintegrin and metalloproteinase with thrombospondin motifs 4(Human)
Astrazeneca
Curated by ChEMBL
Astrazeneca
Curated by ChEMBL
Affinity DataIC50: 400nMAssay Description:Inhibition of ADAMTS-4 assessed as substrate cleavage after 16 hrs by fluorescence assayMore data for this Ligand-Target Pair