Report error Found 104 Enz. Inhib. hit(s) with all data for entry = 8229
Affinity DataIC50: 3.00E+3nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+3nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+3nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 7.00E+3nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+3nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+3nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 9.00E+3nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 9.00E+3nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 9.00E+3nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 1.40E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 2.20E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 3.00E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 3.20E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 3.30E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 3.50E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 4.00E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 4.50E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 5.20E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
Affinity DataIC50: 5.40E+4nMpH: 8.0Assay Description:The rate of hydrolysis of the substrates (0.3 mM) by trypsin (18 U/ml), thrombin (12.5 NIH U/ml), urokinase (62500 IU/ml) and elastase (8.4 U/ml) was...More data for this Ligand-Target Pair
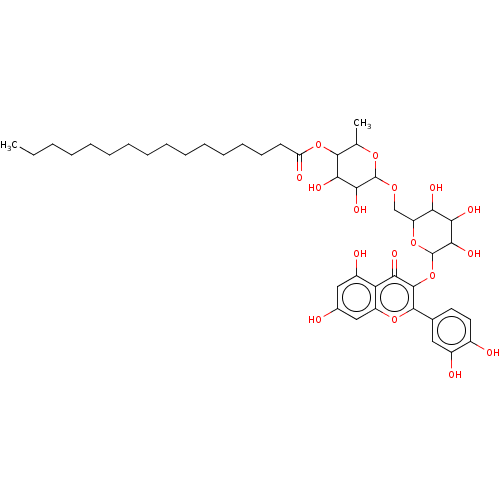
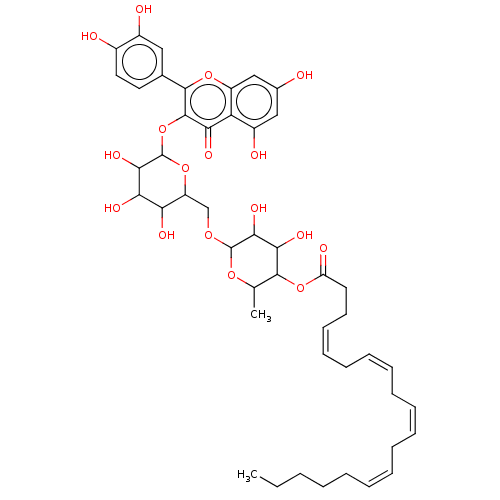
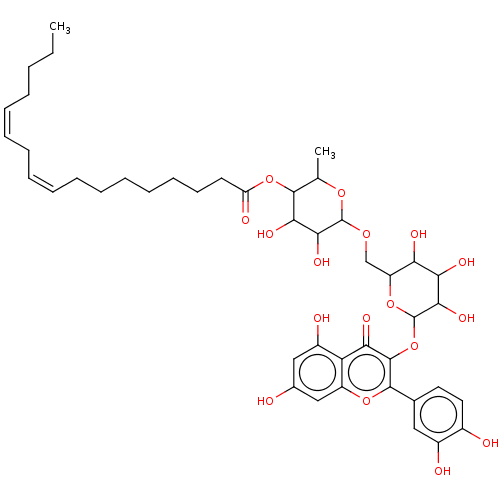
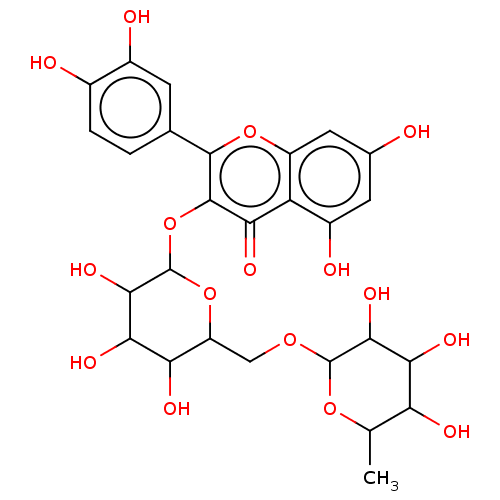
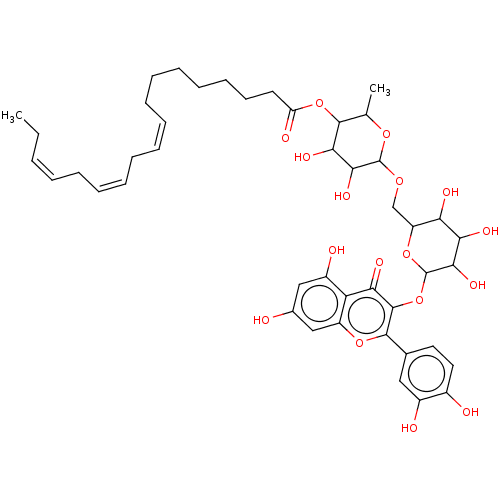
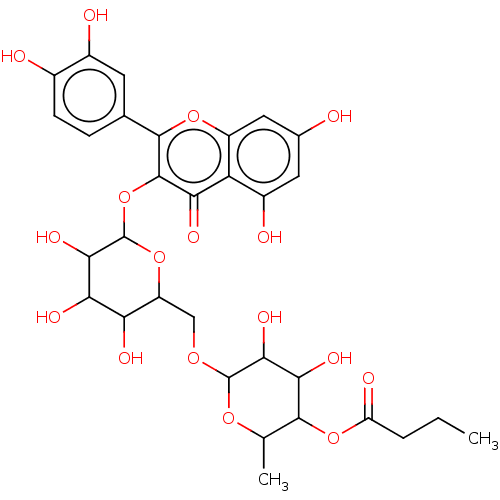
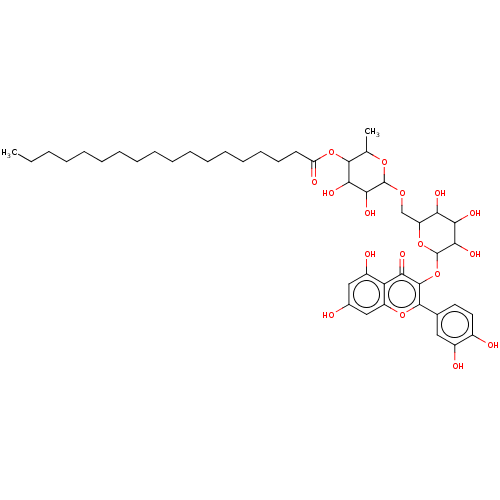
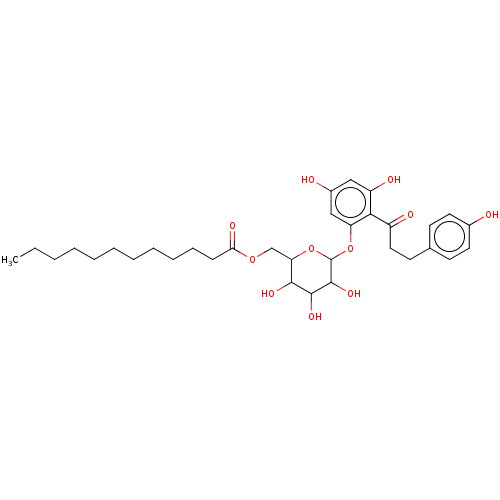
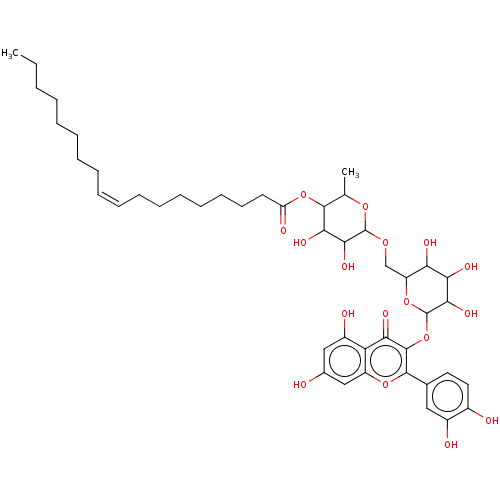
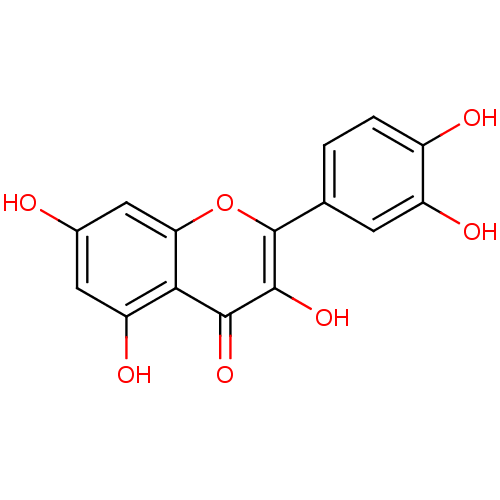
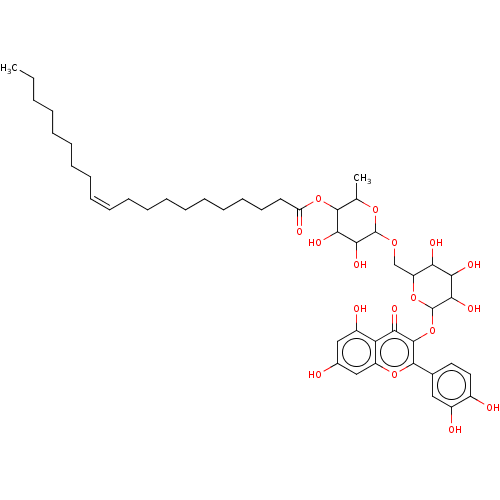
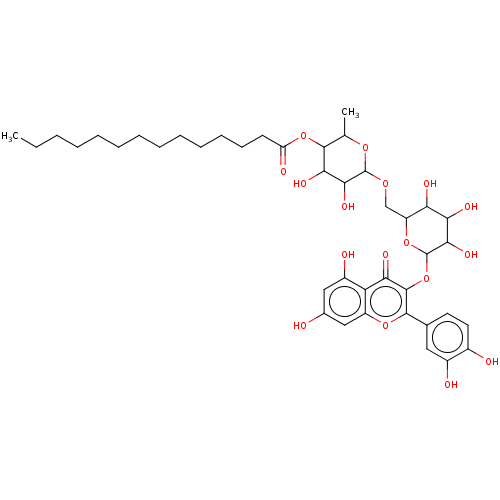
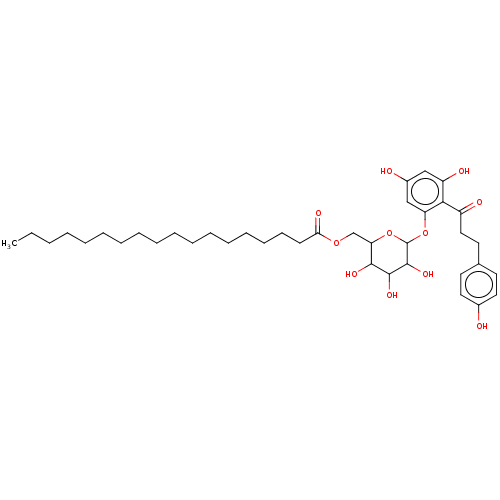
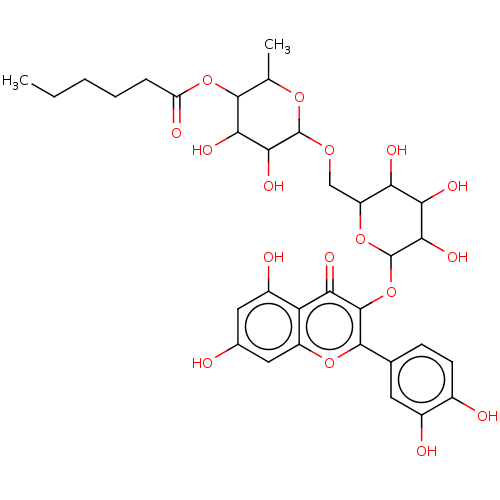


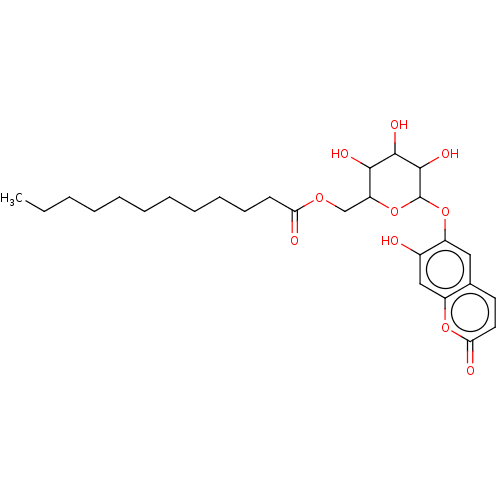
 3D Structure (crystal)
3D Structure (crystal)