Report error Found 27 Enz. Inhib. hit(s) with all data for entry = 50005306
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
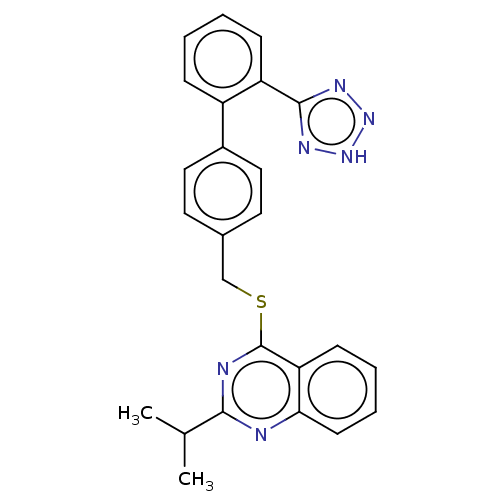
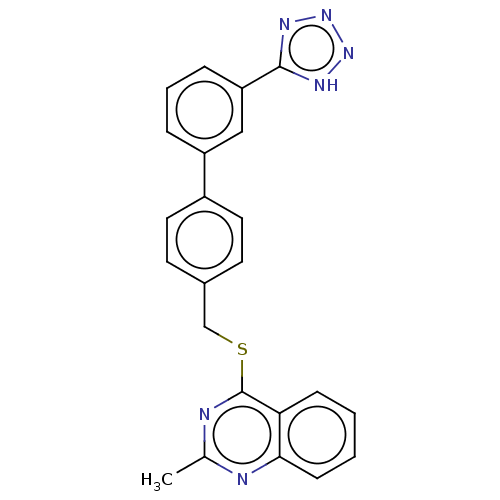
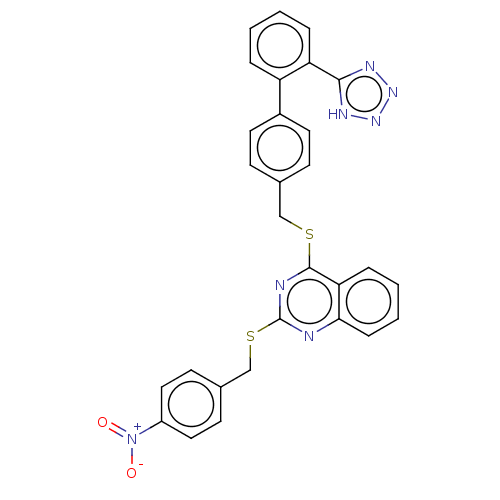
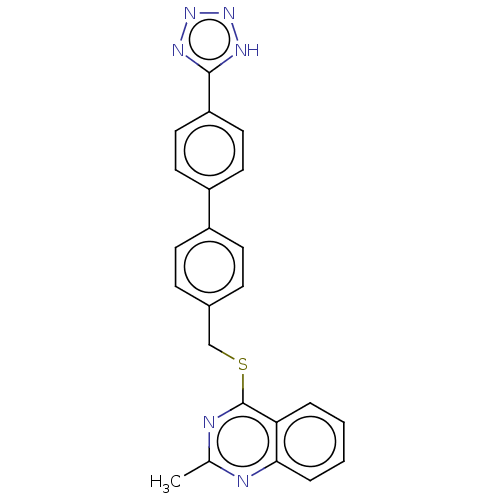
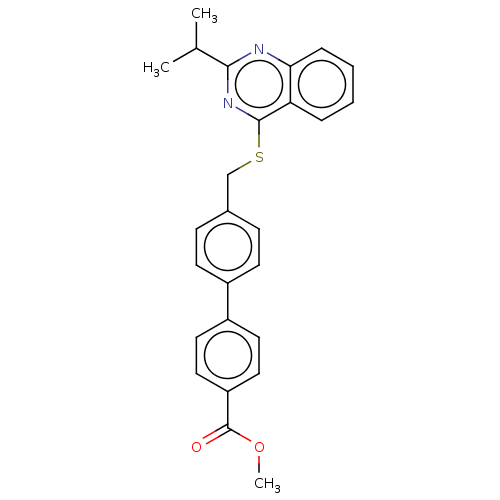
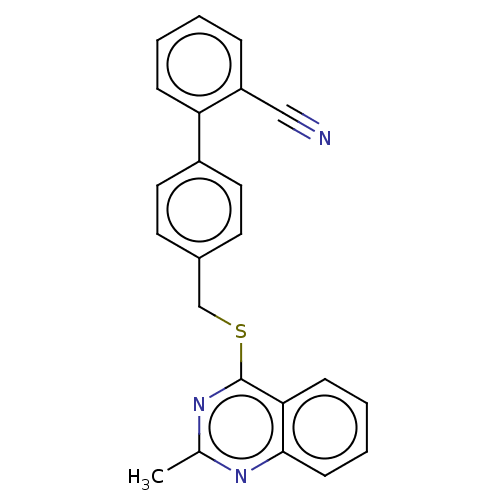
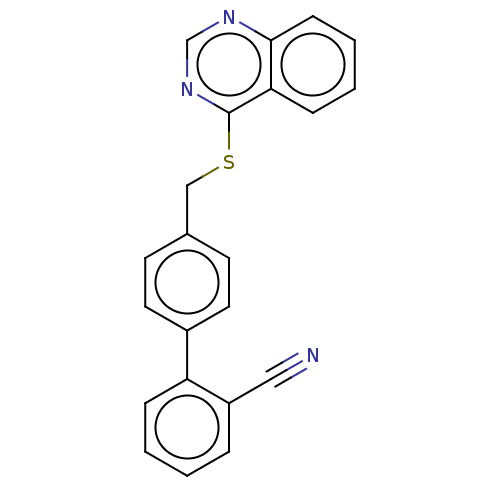
Affinity DataIC50: 770nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
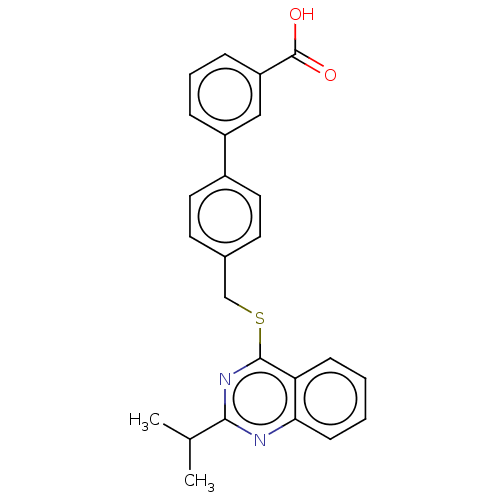
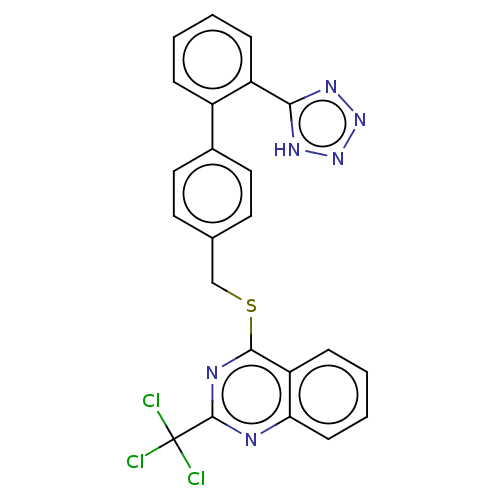
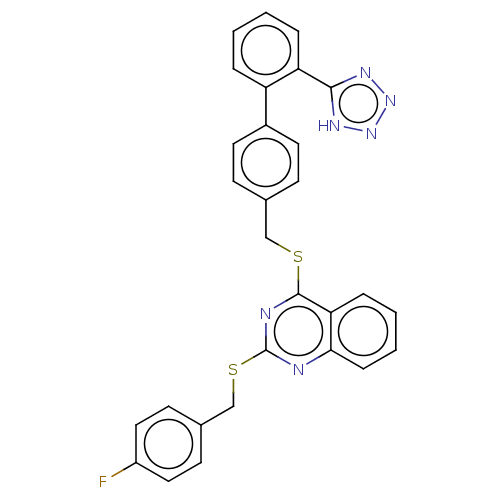
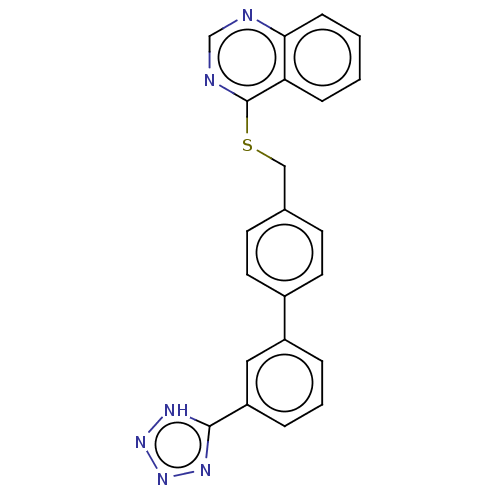
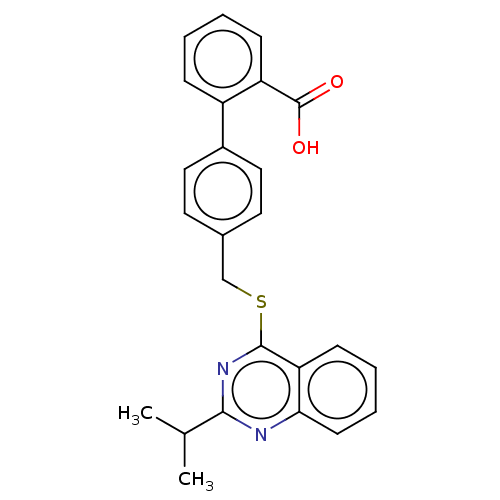
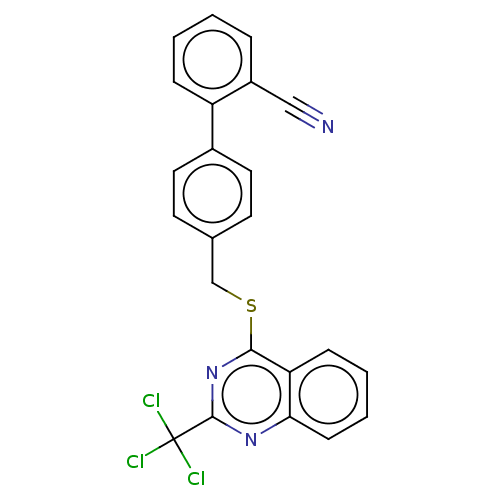
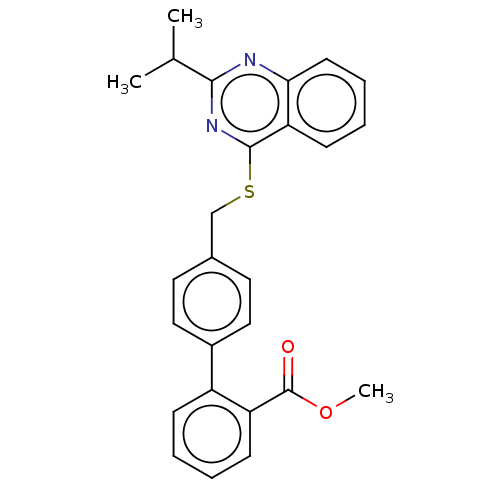
Affinity DataIC50: 850nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
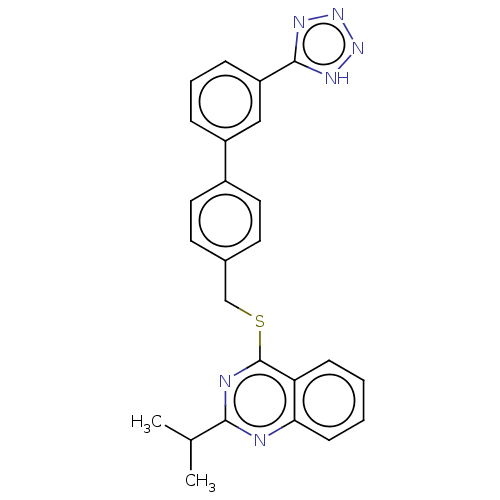
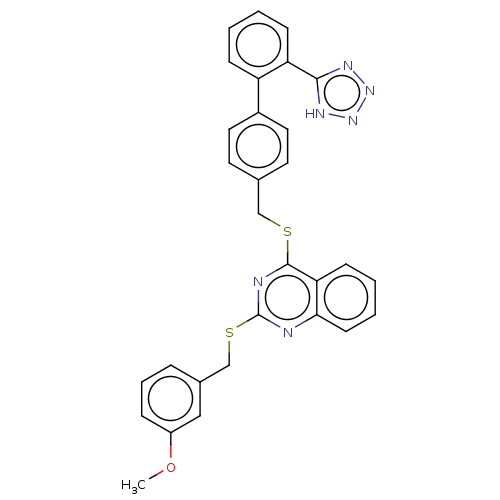
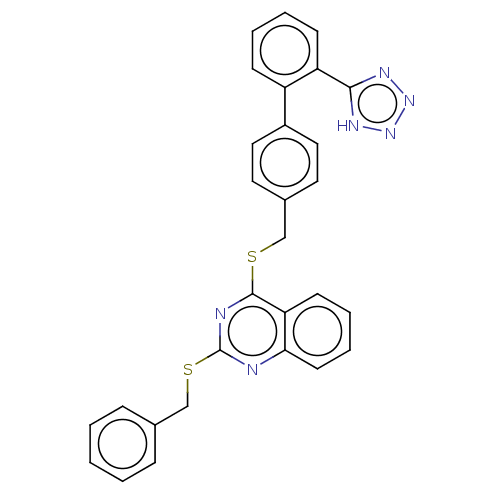
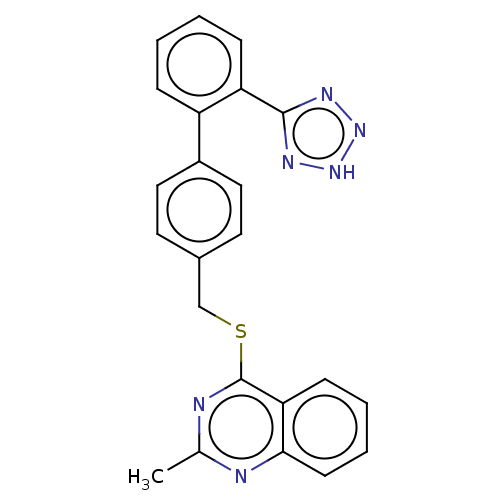
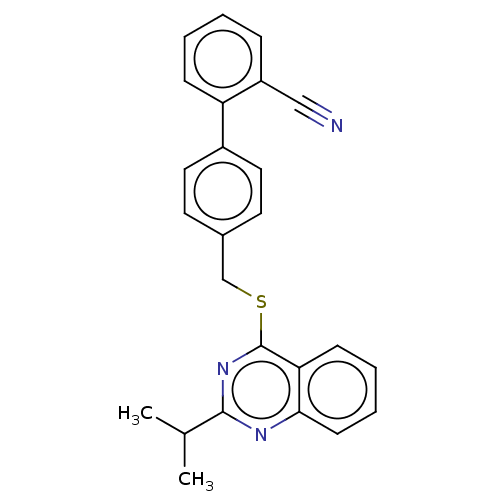
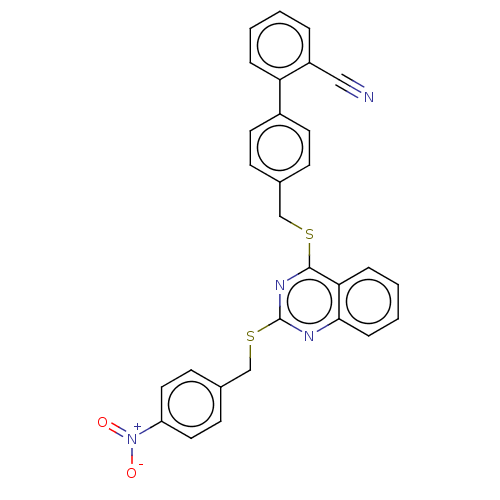
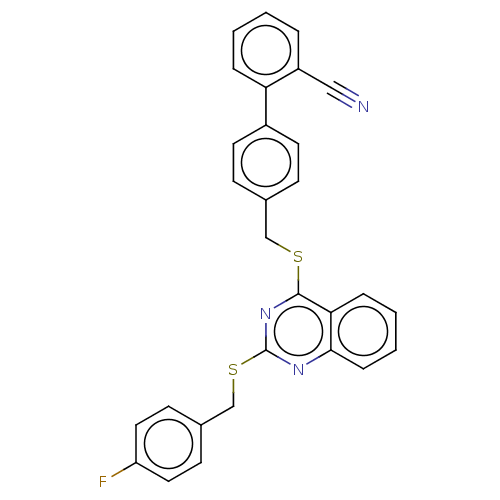
Affinity DataIC50: 890nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
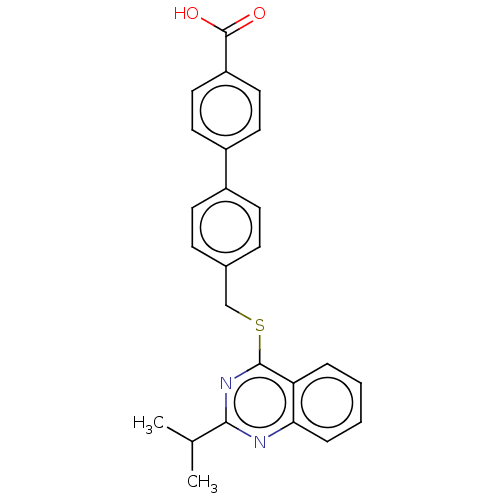
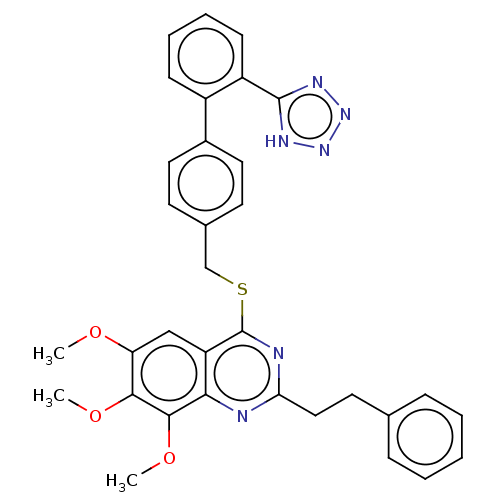
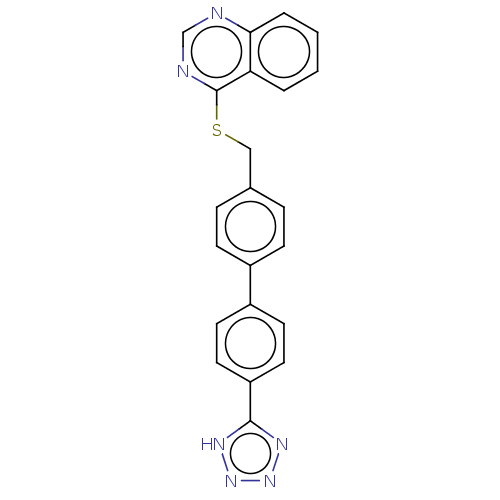
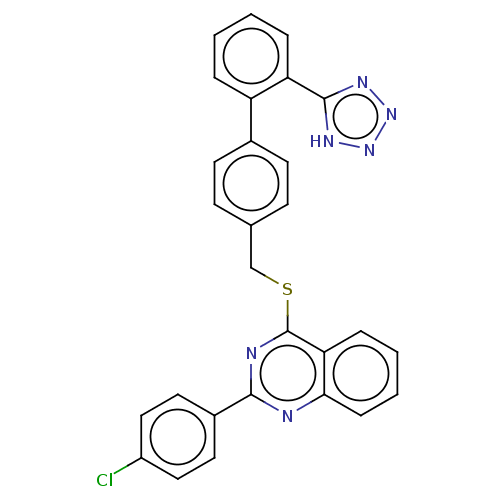
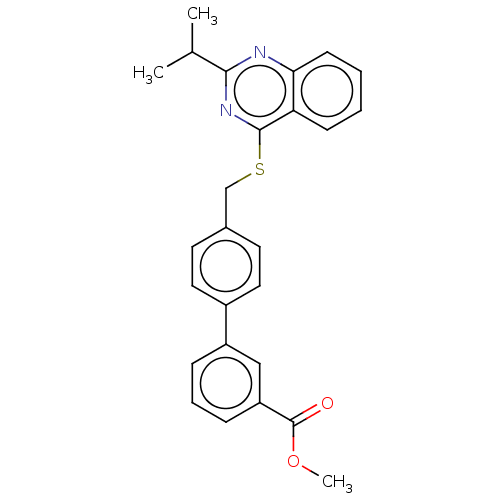
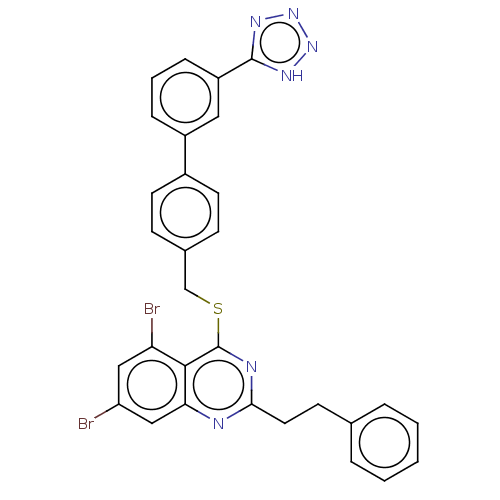
Affinity DataIC50: 930nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 2.50E+3nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 2.60E+3nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 3.30E+3nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 3.60E+3nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 4.90E+3nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 6.80E+3nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 7.10E+3nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 7.20E+3nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 8.00E+3nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 8.20E+3nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 1.10E+4nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 3.30E+4nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 4.40E+4nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair
TargetHigh affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A/cAMP-specific 3',5'-cyclic phosphodiesterase 7B(Human)
Universidad De Alcala
Curated by ChEMBL
Universidad De Alcala
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of PDE7 catalytic domain (unknown origin) using [3H]-cAMP as substrate after 1 hr by scintillation proximity assayMore data for this Ligand-Target Pair