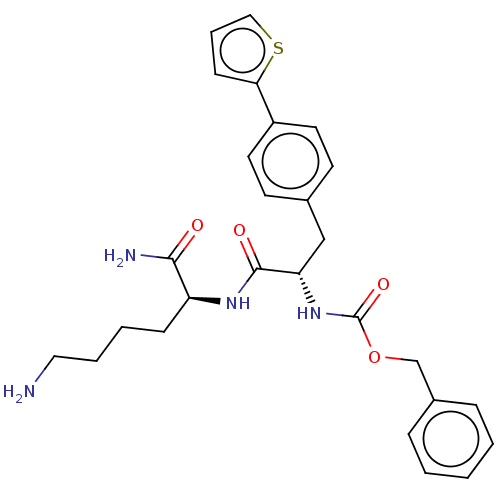
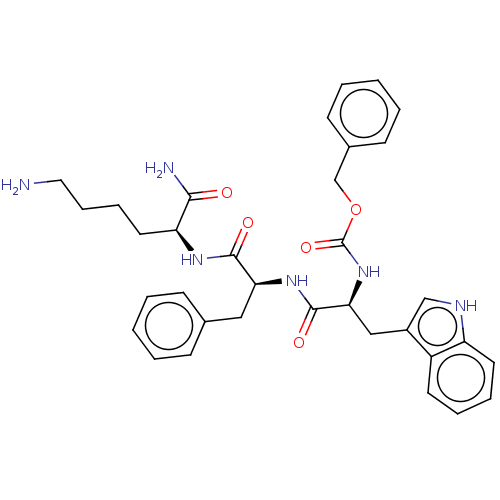
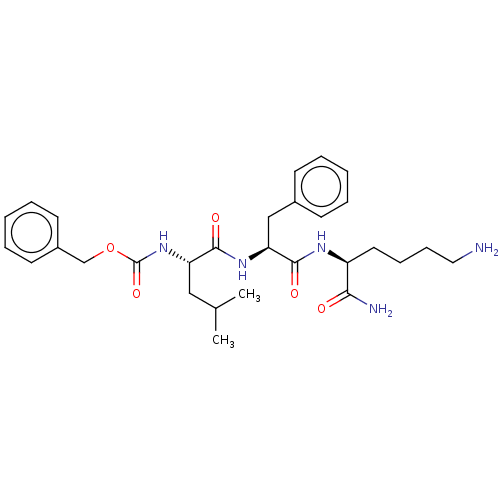
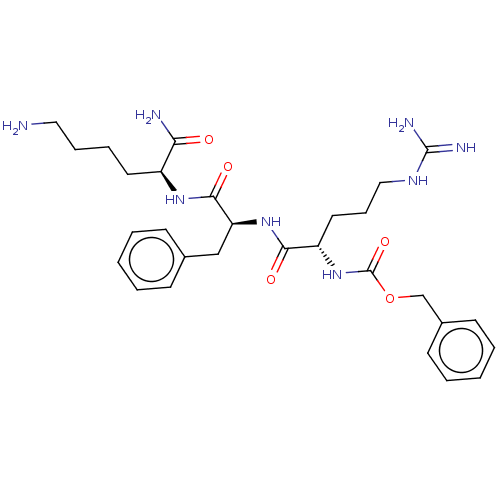
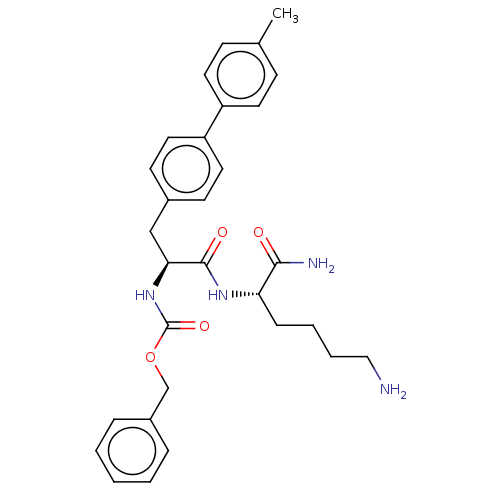
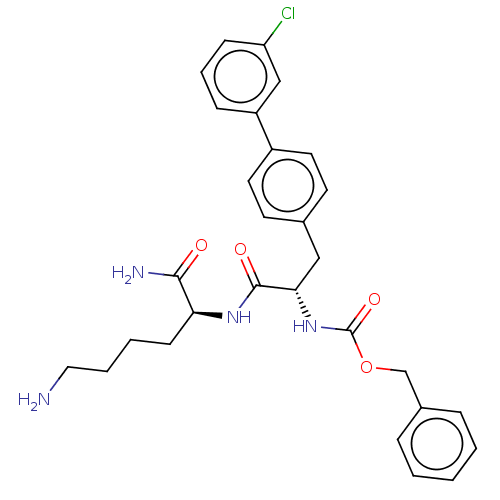
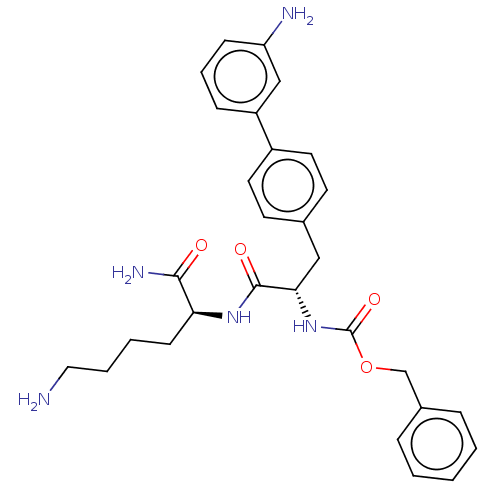
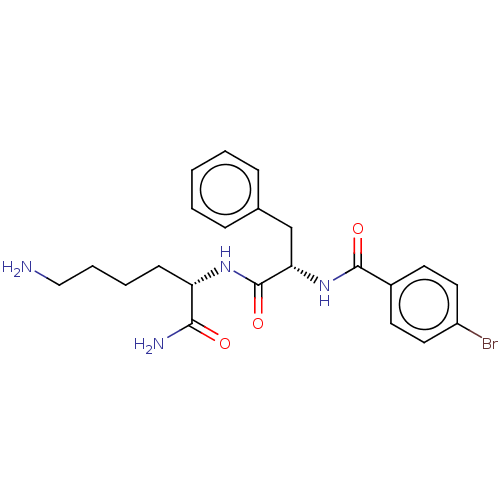
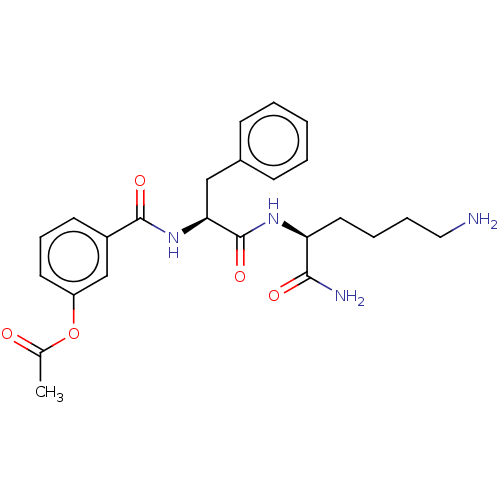
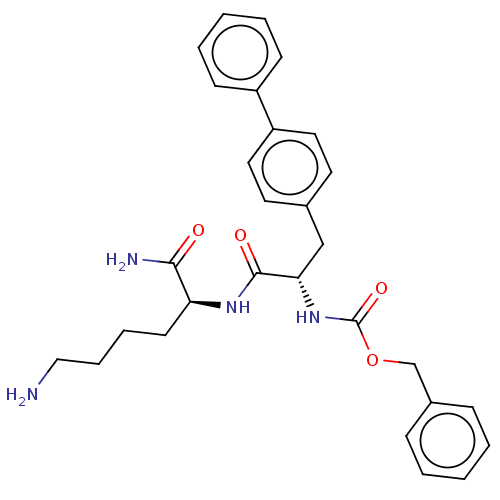
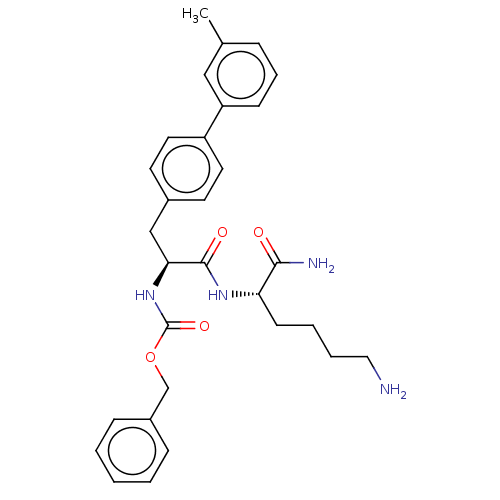
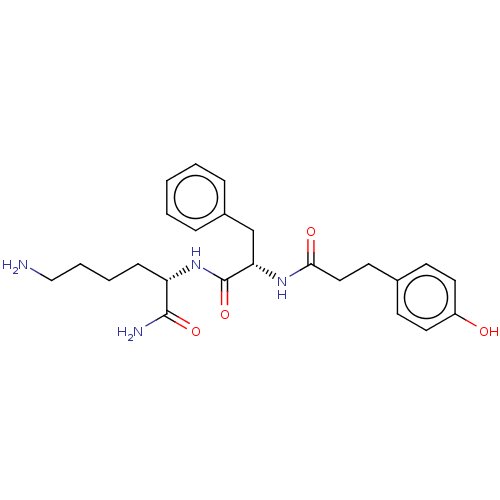
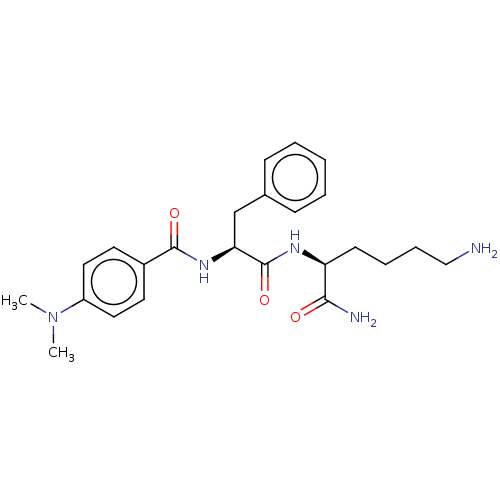
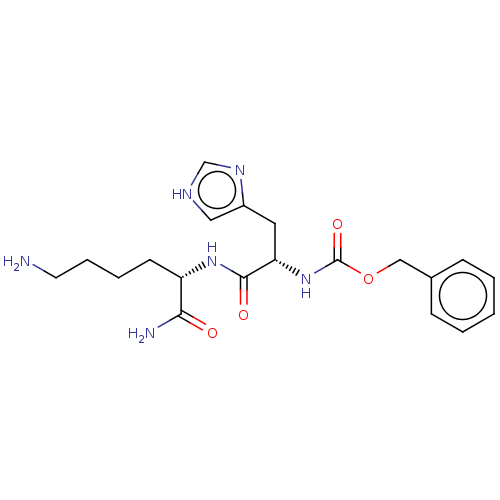
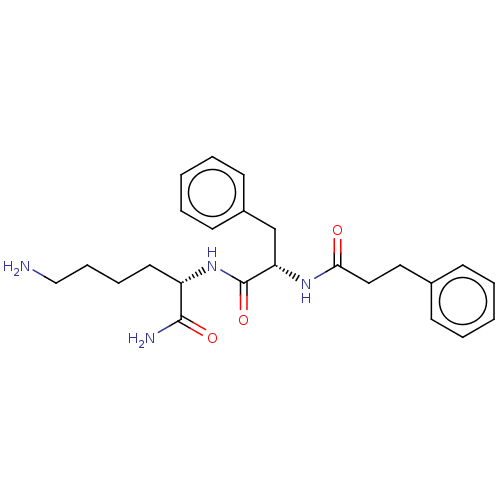
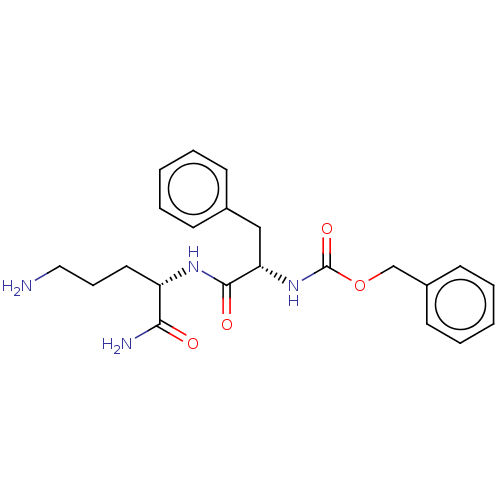
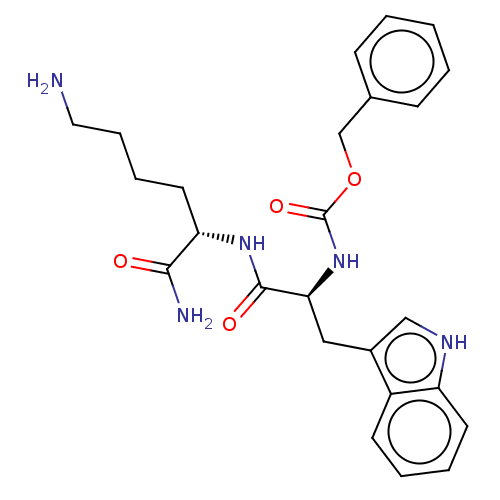
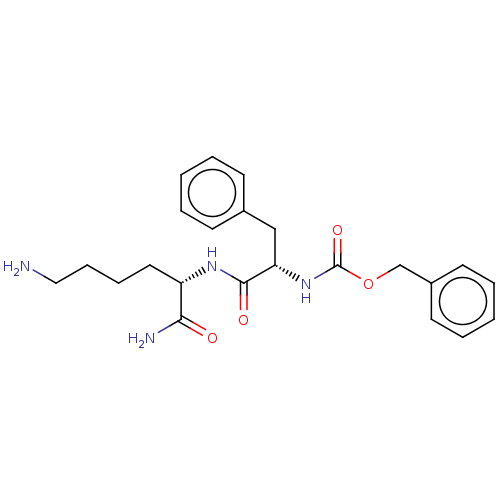
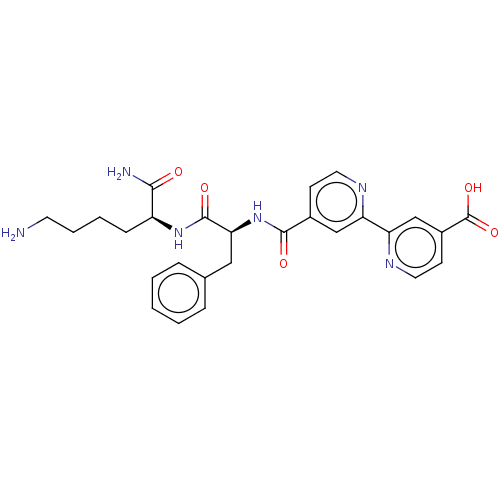
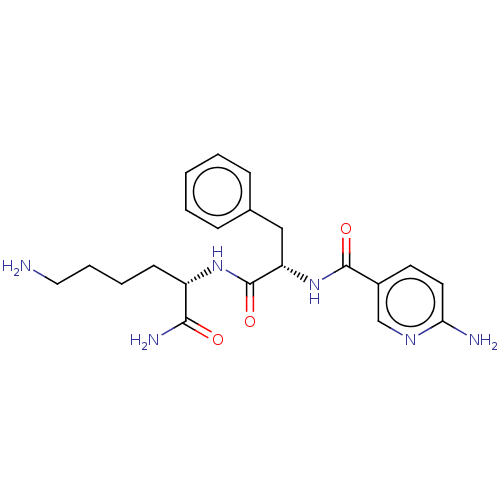
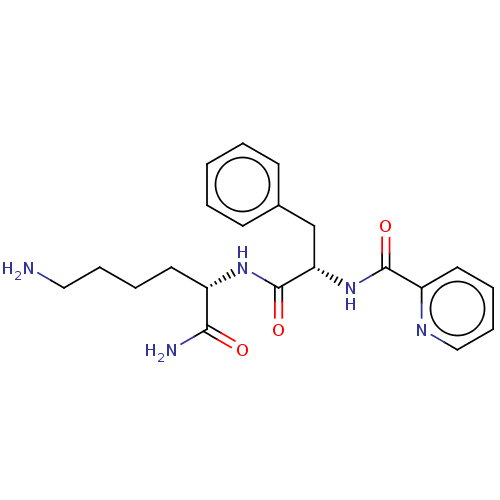
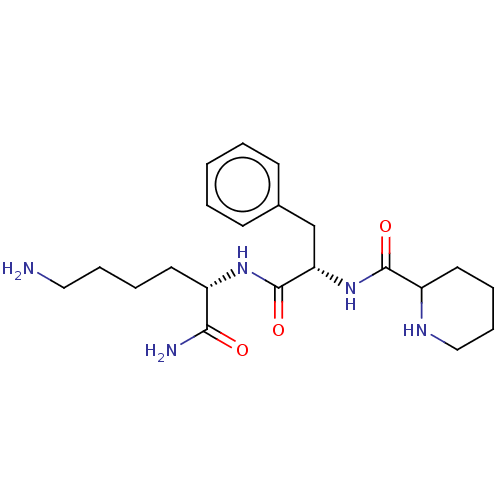
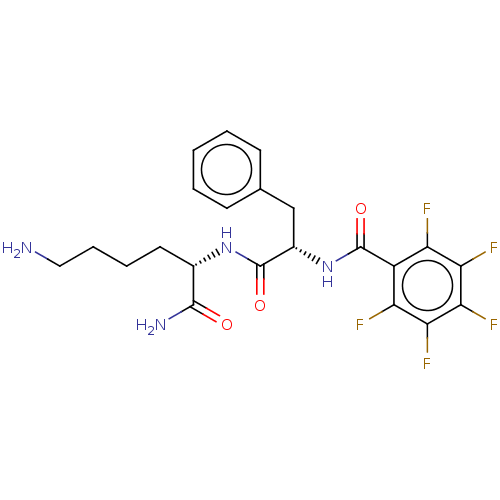
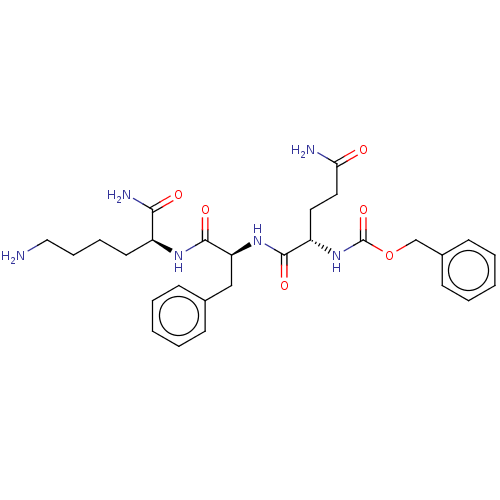
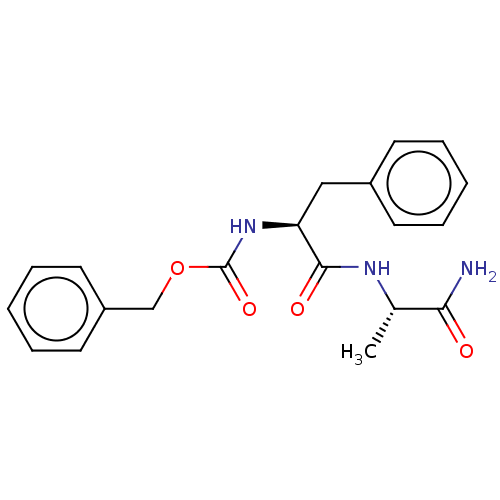
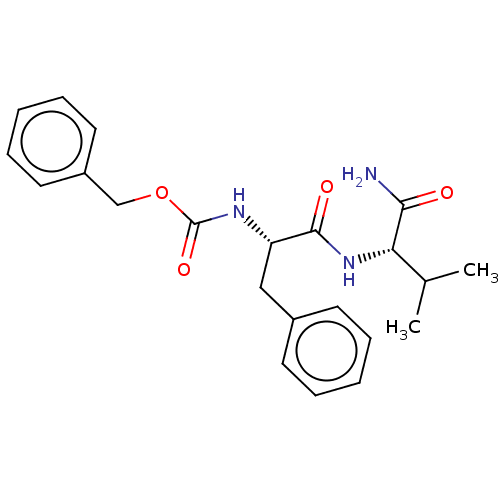
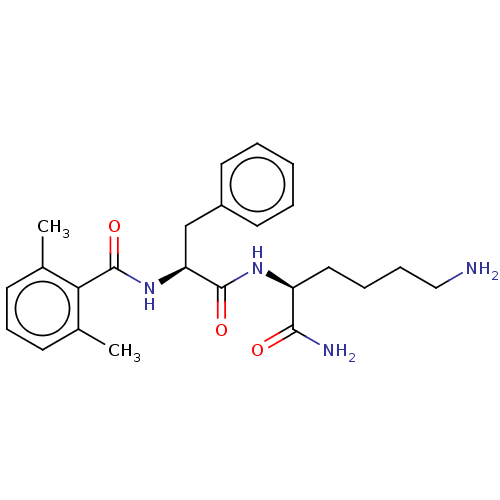
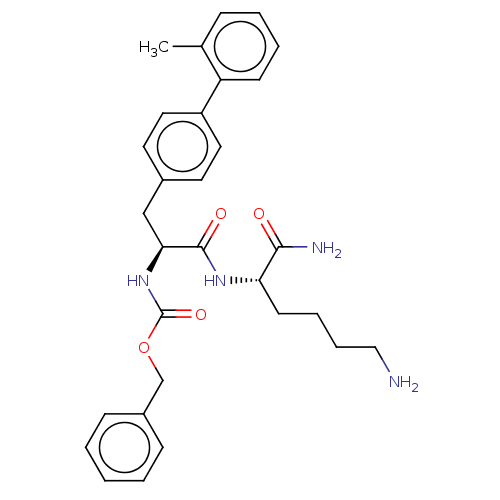
Report error Found 98 Enz. Inhib. hit(s) with all data for entry = 50005347
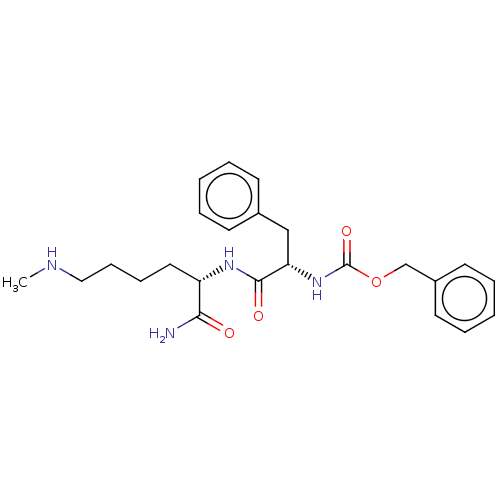
Affinity DataKi: 120nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
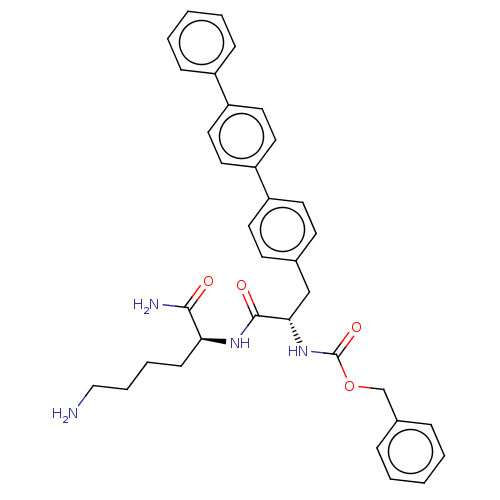
Affinity DataKi: 390nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
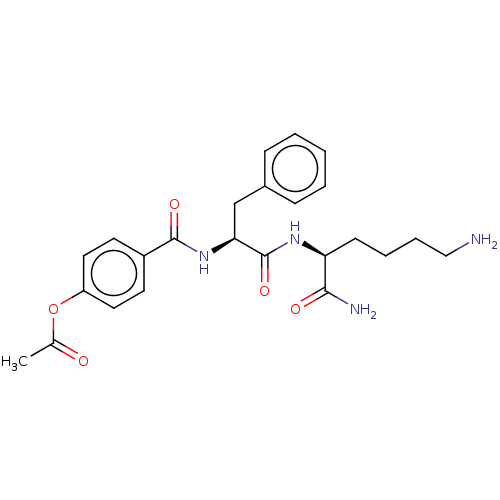
Affinity DataKi: 450nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
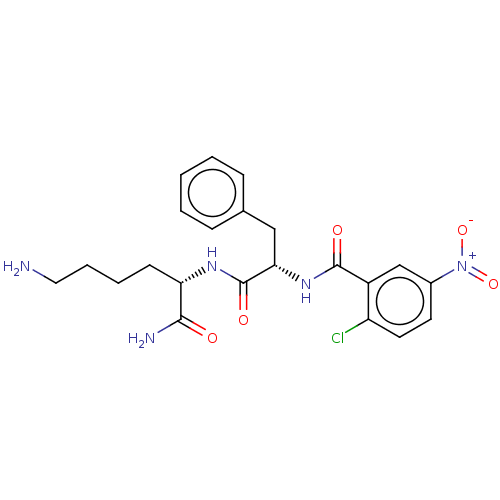
Affinity DataKi: 740nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 890nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.00E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.00E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.40E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.70E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.70E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.80E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.30E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.40E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.40E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.50E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.50E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.60E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.60E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.60E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.70E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.90E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.10E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.20E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.50E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 4.20E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 4.20E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 4.60E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 5.00E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 5.50E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 6.10E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 6.80E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 7.00E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 7.80E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 8.50E+3nMAssay Description:Inhibition of recombinant cathepsin-S (unknown origin) using Z-Val-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 9.00E+3nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.20E+4nMAssay Description:Inhibition of recombinant cathepsin-S (unknown origin) using Z-Val-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.40E+4nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.40E+4nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.60E+4nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.70E+4nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.00E+4nMAssay Description:Inhibition of recombinant cathepsin-S (unknown origin) using Z-Val-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.00E+4nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.30E+4nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.40E+4nMAssay Description:Inhibition of recombinant cathepsin-K (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.60E+4nMAssay Description:Inhibition of recombinant cathepsin-B (unknown origin) using Z-Arg-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.60E+4nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.70E+4nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.80E+4nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.80E+4nMAssay Description:Inhibition of recombinant cathepsin-S (unknown origin) using Z-Val-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.80E+4nMAssay Description:Inhibition of recombinant cathepsin-L (unknown origin) using Z-Phe-Arg-AMC as substrate by fluorescence assayMore data for this Ligand-Target Pair