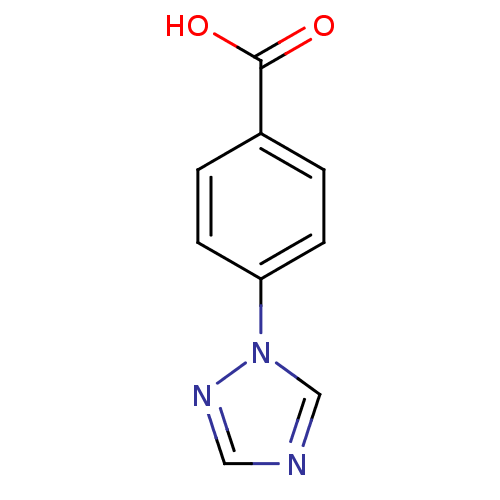
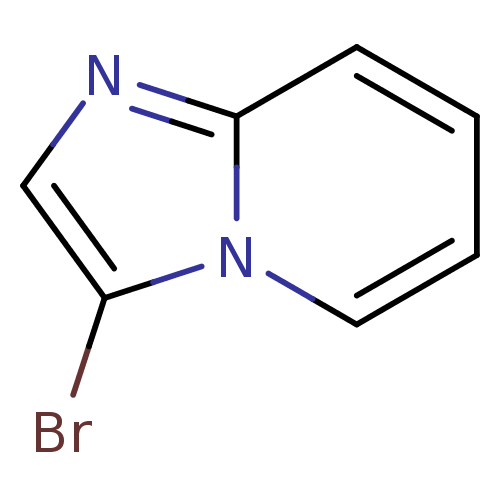
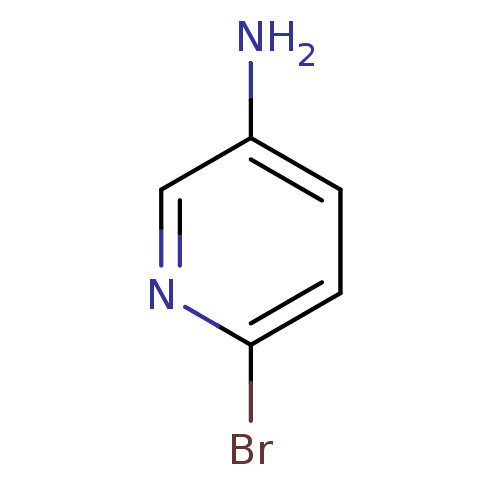
Report error Found 24 Enz. Inhib. hit(s) with all data for entry = 6037
Affinity DataIC50: 11nMAssay Description:Cocktails of 4-8 compounds were used for the initial screening with single compound soaks to verify the bound fragment identity and to rule out coope...More data for this Ligand-Target Pair
Affinity DataIC50: 41nMAssay Description:Cocktails of 4-8 compounds were used for the initial screening with single compound soaks to verify the bound fragment identity and to rule out coope...More data for this Ligand-Target Pair
Affinity DataIC50: 380nMAssay Description:Cocktails of 4-8 compounds were used for the initial screening with single compound soaks to verify the bound fragment identity and to rule out coope...More data for this Ligand-Target Pair
Affinity DataIC50: 450nMAssay Description:Cocktails of 4-8 compounds were used for the initial screening with single compound soaks to verify the bound fragment identity and to rule out coope...More data for this Ligand-Target Pair
Affinity DataIC50: 730nMAssay Description:Cocktails of 4-8 compounds were used for the initial screening with single compound soaks to verify the bound fragment identity and to rule out coope...More data for this Ligand-Target Pair
Affinity DataIC50: 2.00E+3nMAssay Description:Cocktails of 4-8 compounds were used for the initial screening with single compound soaks to verify the bound fragment identity and to rule out coope...More data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+4nMAssay Description:Cocktails of 4-8 compounds were used for the initial screening with single compound soaks to verify the bound fragment identity and to rule out coope...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+4nMAssay Description:Cocktails of 4-8 compounds were used for the initial screening with single compound soaks to verify the bound fragment identity and to rule out coope...More data for this Ligand-Target Pair
Affinity DataIC50: 2.50E+5nMAssay Description:Cocktails of 4-8 compounds were used for the initial screening with single compound soaks to verify the bound fragment identity and to rule out coope...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Cocktails of 4-8 compounds were used for the initial screening with single compound soaks to verify the bound fragment identity and to rule out coope...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Cocktails of 4-8 compounds were used for the initial screening with single compound soaks to verify the bound fragment identity and to rule out coope...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Cocktails of 4-8 compounds were used for the initial screening with single compound soaks to verify the bound fragment identity and to rule out coope...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Cocktails of 4-8 compounds were used for the initial screening with single compound soaks to verify the bound fragment identity and to rule out coope...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Cocktails of 4-8 compounds were used for the initial screening with single compound soaks to verify the bound fragment identity and to rule out coope...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Cocktails of 4-8 compounds were used for the initial screening with single compound soaks to verify the bound fragment identity and to rule out coope...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+6nMAssay Description:Cocktails of 4-8 compounds were used for the initial screening with single compound soaks to verify the bound fragment identity and to rule out coope...More data for this Ligand-Target Pair
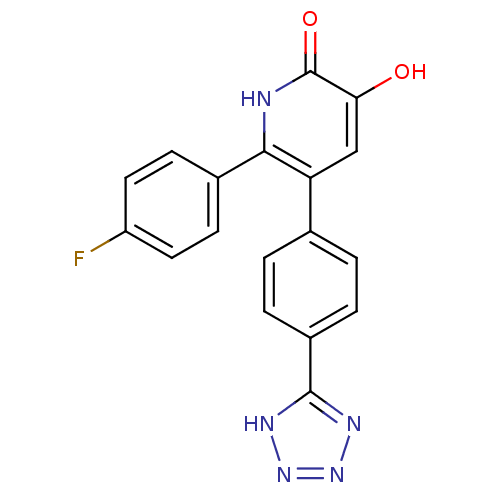

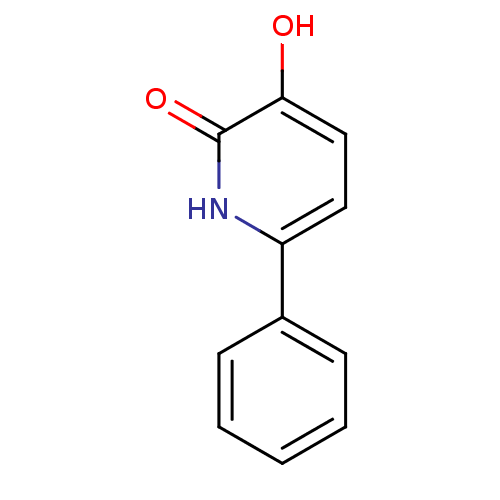
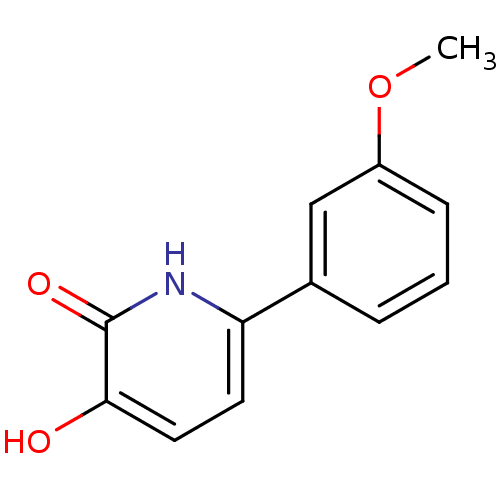
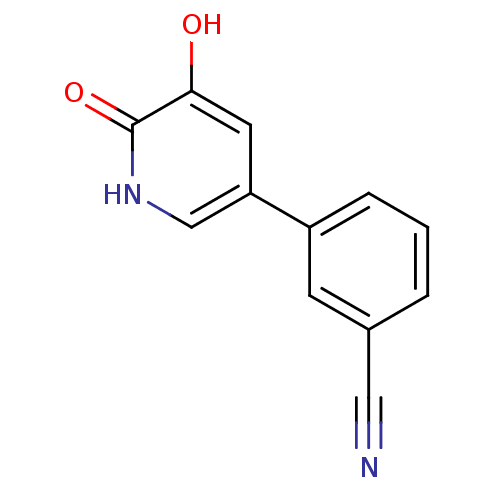
 3D Structure (crystal)
3D Structure (crystal)