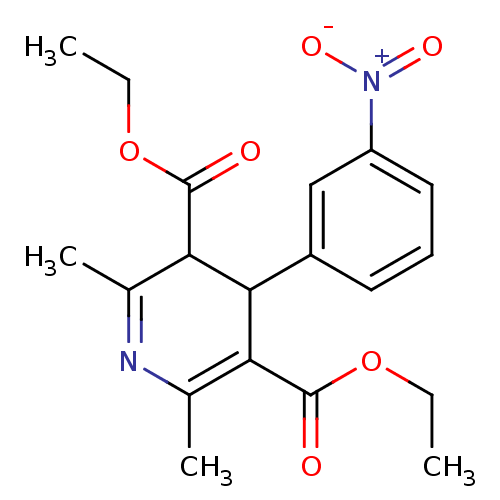
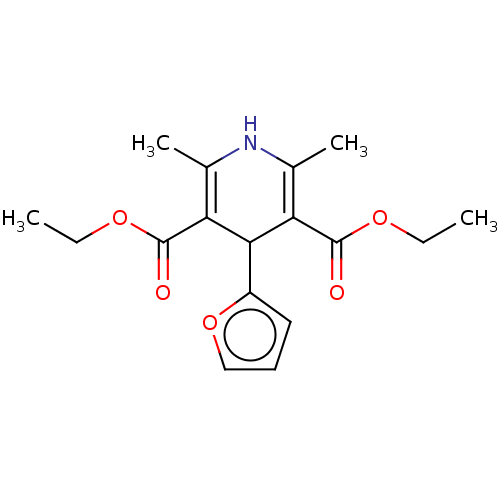
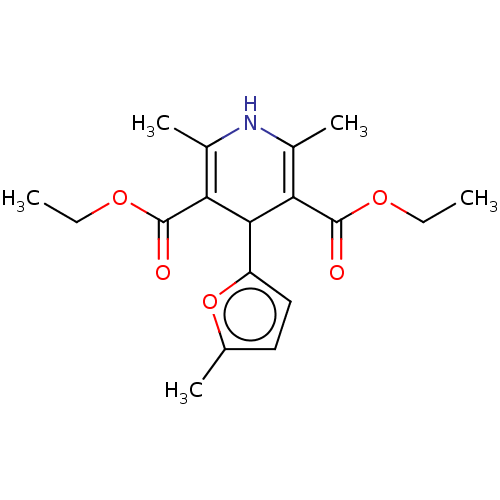
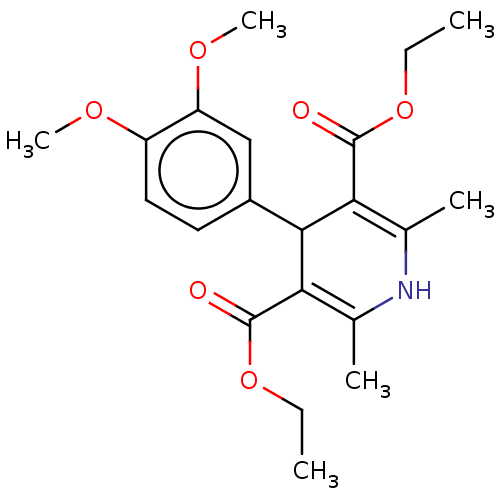
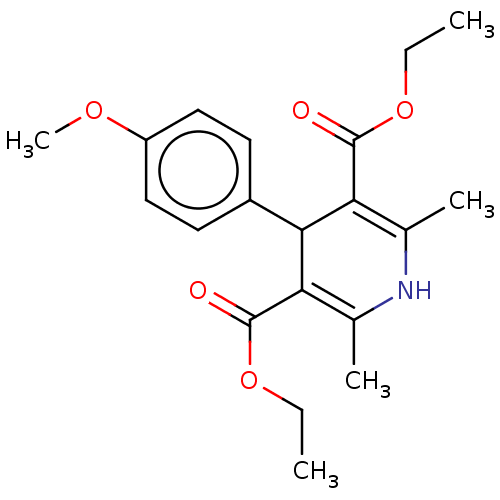
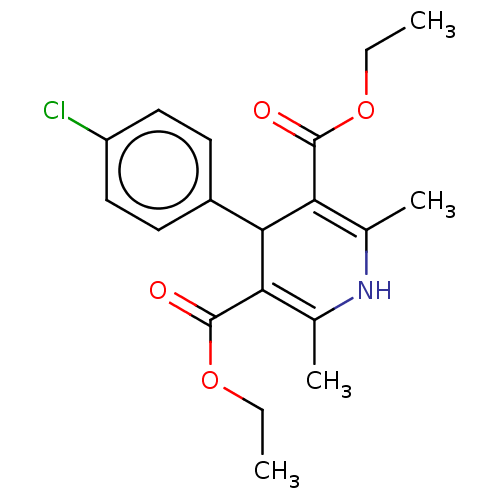
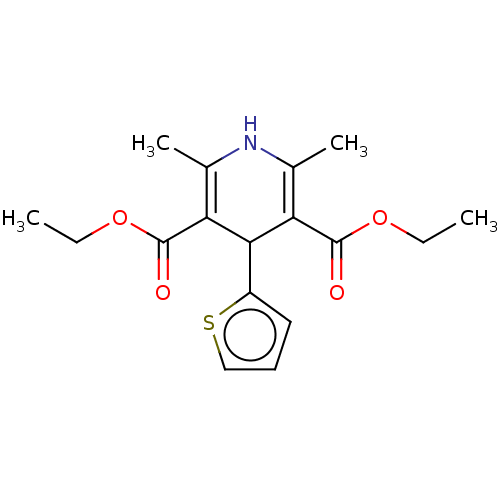
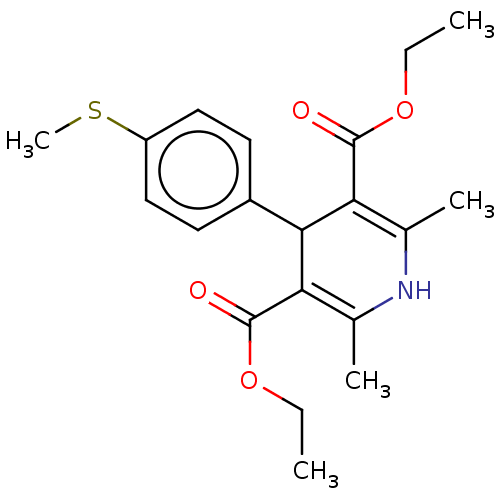
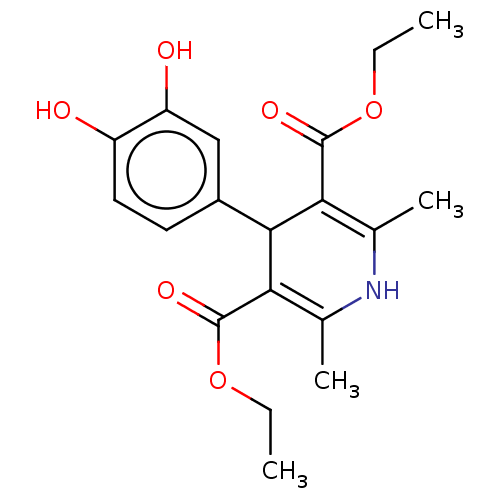
Report error Found 27 Enz. Inhib. hit(s) with all data for entry = 50045675
Affinity DataKi: 2.50E+4nMAssay Description:Non-competitive inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15...More data for this Ligand-Target Pair
Affinity DataIC50: 3.50E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 5.00E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 5.10E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 6.02E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 6.51E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataKi: 6.80E+4nMAssay Description:Competitive inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 min...More data for this Ligand-Target Pair
Affinity DataIC50: 7.24E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 7.43E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 9.74E+4nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 1.12E+5nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 1.18E+5nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 1.22E+5nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 1.24E+5nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair
Affinity DataIC50: 2.74E+5nMAssay Description:Inhibition of Saccharomyces cerevisiae alpha-glucosidase using p-nitrophenyl alpha-D-glucopyranoside as substrate preincubated for 15 mins measured u...More data for this Ligand-Target Pair