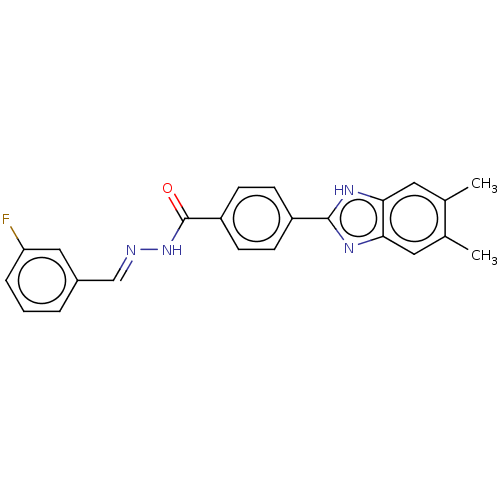
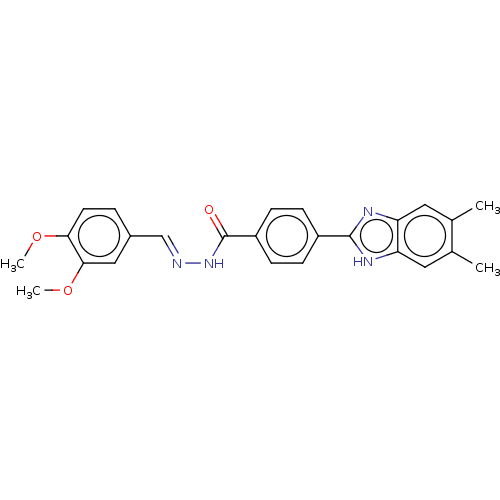
Report error Found 27 Enz. Inhib. hit(s) with all data for entry = 7120
Affinity DataIC50: 8.40E+3nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.44E+3nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.44E+3nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.74E+3nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 8.74E+3nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 9.99E+3nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.25E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.51E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.82E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.83E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.94E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.24E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.35E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.48E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.48E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.57E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.60E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 2.90E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.21E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.25E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.26E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 3.45E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 4.60E+4nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.48E+5nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.58E+5nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 1.80E+5nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair
Affinity DataIC50: 7.75E+5nMpH: 6.8Assay Description:The enzyme inhibition was evaluated according to the method previously reported by Rahim et al. [25] with slight modification. Various concentration ...More data for this Ligand-Target Pair