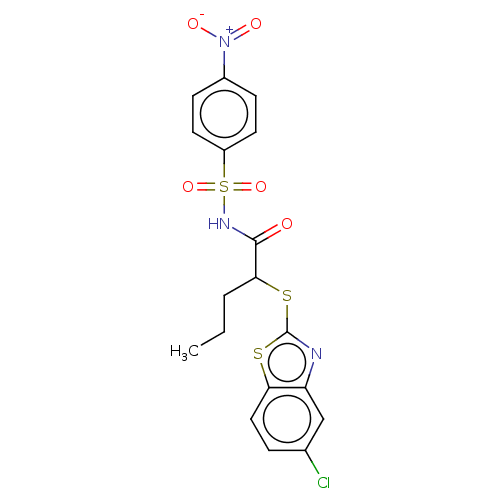
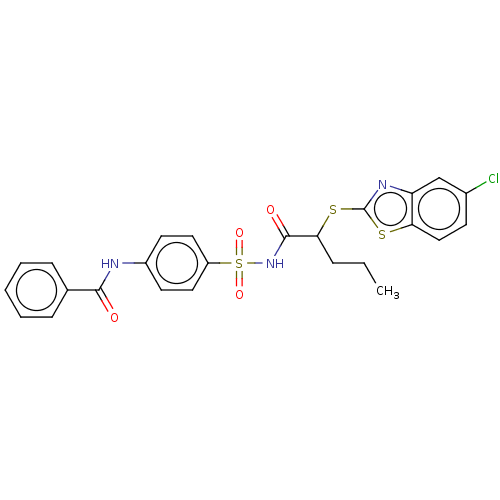
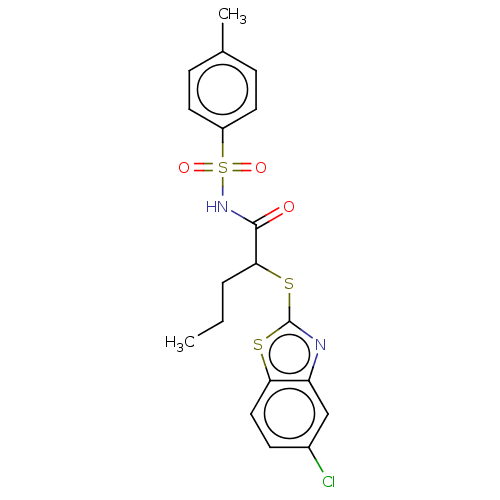
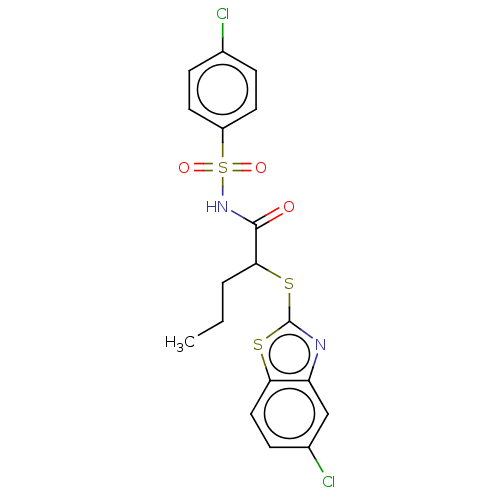
Report error Found 11 Enz. Inhib. hit(s) with all data for entry = 50047232
TargetPeroxisome proliferator-activated receptor alpha(Human)
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
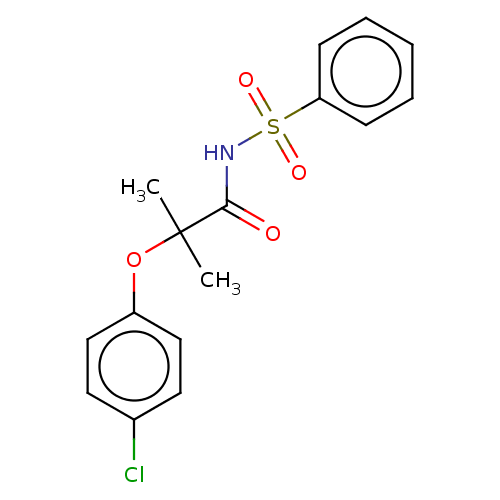
Affinity DataIC50: 650nMAssay Description:Antagonist activity at PPARalpha (unknown origin) expressed in HEK293A cells assessed as inhibition of GW7647-induced transactivation after 16 to 18 ...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor alpha(Human)
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
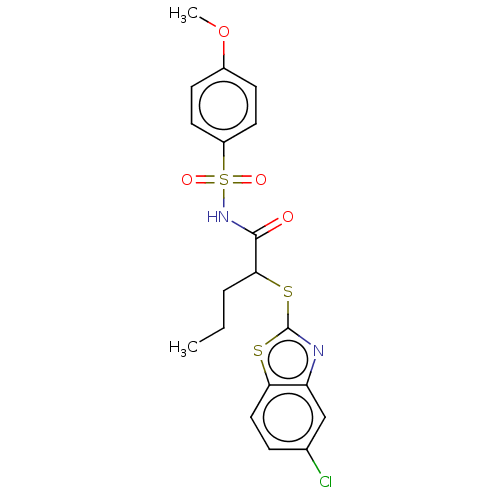
Affinity DataIC50: 680nMAssay Description:Antagonist activity at PPARalpha (unknown origin) expressed in HEK293A cells assessed as inhibition of GW7647-induced transactivation after 16 to 18 ...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor alpha(Human)
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
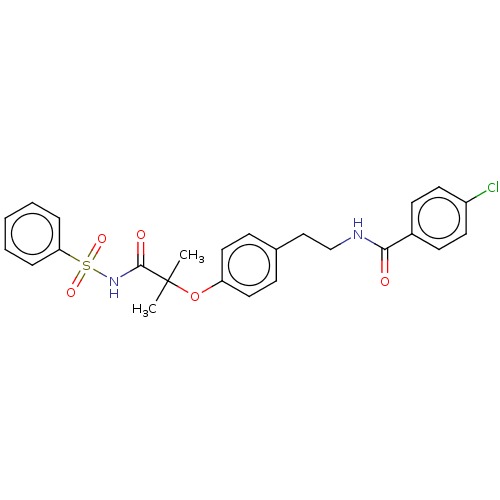
Affinity DataIC50: 800nMAssay Description:Antagonist activity at PPARalpha (unknown origin) expressed in HEK293A cells assessed as inhibition of GW7647-induced transactivation after 16 to 18 ...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor alpha(Human)
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
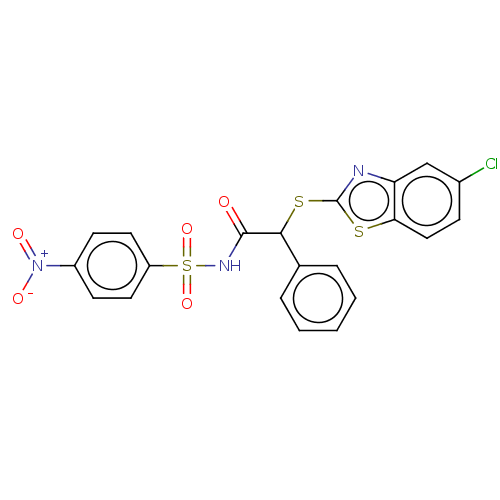
Affinity DataIC50: 820nMAssay Description:Antagonist activity at PPARalpha (unknown origin) expressed in HEK293A cells assessed as inhibition of GW7647-induced transactivation after 16 to 18 ...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor alpha(Human)
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
Affinity DataIC50: 850nMAssay Description:Antagonist activity at PPARalpha (unknown origin) expressed in HEK293A cells assessed as inhibition of GW7647-induced transactivation after 16 to 18 ...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor alpha(Human)
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
Affinity DataIC50: 4.50E+3nMAssay Description:Antagonist activity at PPARalpha (unknown origin) expressed in HEK293A cells assessed as inhibition of GW7647-induced transactivation after 16 to 18 ...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor alpha(Human)
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
Affinity DataIC50: 4.60E+3nMAssay Description:Antagonist activity at PPARalpha (unknown origin) expressed in HEK293A cells assessed as inhibition of GW7647-induced transactivation after 16 to 18 ...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor alpha(Human)
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
Affinity DataIC50: 6.86E+3nMAssay Description:Antagonist activity at PPARalpha (unknown origin) expressed in HEK293A cells assessed as inhibition of GW7647-induced transactivation after 16 to 18 ...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor alpha(Human)
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
Affinity DataIC50: 7.65E+3nMAssay Description:Antagonist activity at PPARalpha (unknown origin) expressed in HEK293A cells assessed as inhibition of GW7647-induced transactivation after 16 to 18 ...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor alpha(Human)
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
Affinity DataIC50: 8.80E+3nMAssay Description:Antagonist activity at PPARalpha (unknown origin) expressed in HEK293A cells assessed as inhibition of GW7647-induced transactivation after 16 to 18 ...More data for this Ligand-Target Pair
TargetPeroxisome proliferator-activated receptor alpha(Human)
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
Universit£&Quot;G. D'Annunzio&Quot
Curated by ChEMBL
Affinity DataIC50: 1.08E+5nMAssay Description:Antagonist activity at PPARalpha (unknown origin) expressed in HEK293A cells assessed as inhibition of GW7647-induced transactivation after 16 to 18 ...More data for this Ligand-Target Pair