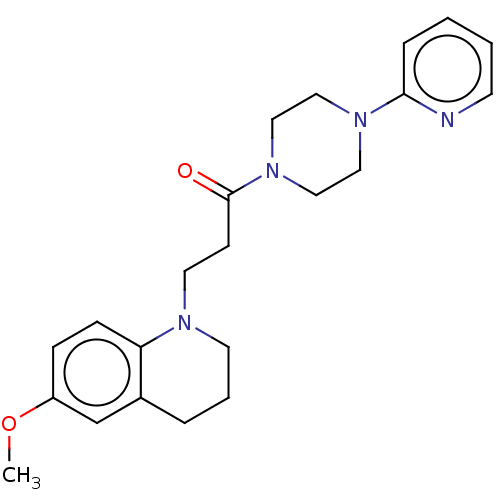
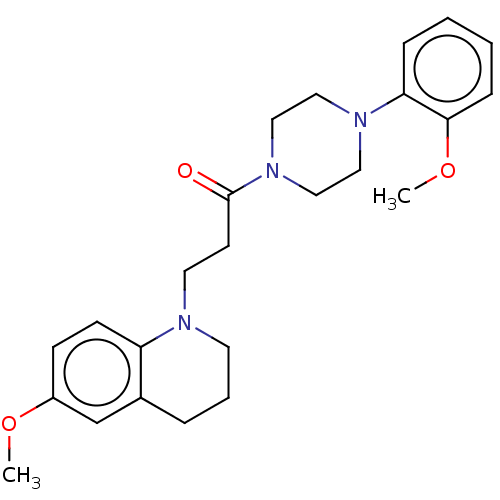
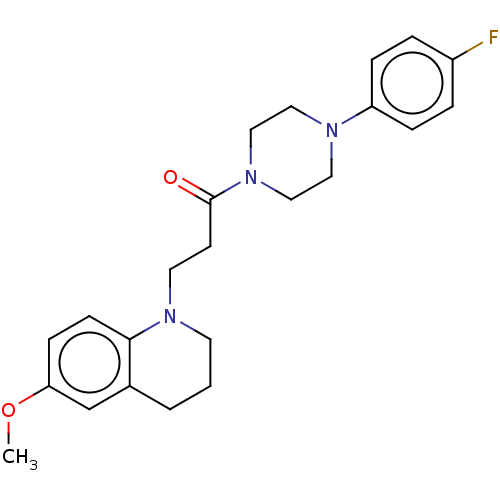
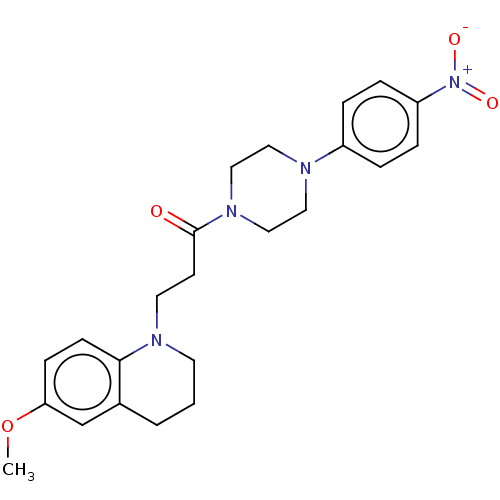
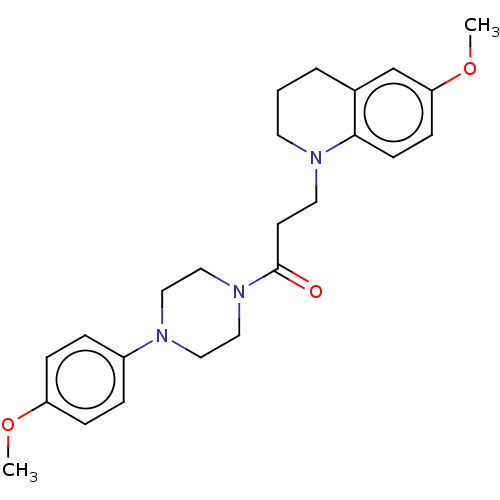
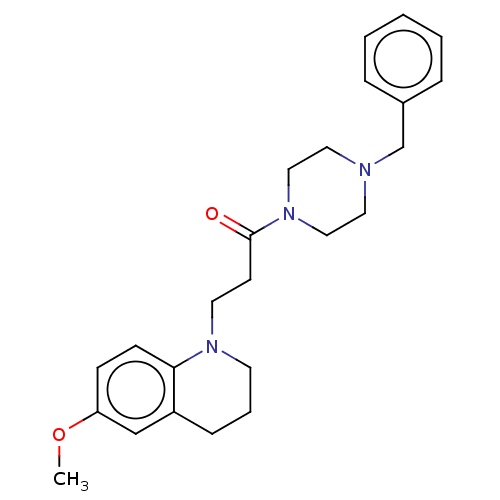
Report error Found 16 Enz. Inhib. hit(s) with all data for entry = 7391
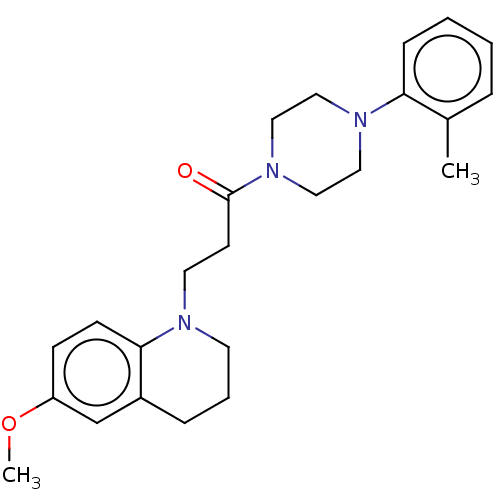
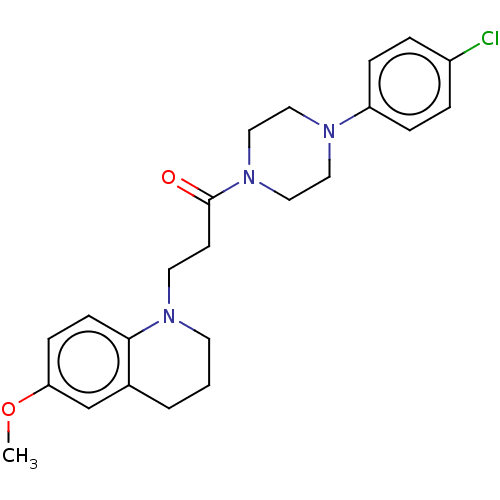
Affinity DataIC50: 25nMAssay Description:The reaction mixture was set with template primer complex, RT enzyme and dNTPs in a lysis buffer with or without inhibitors. The reaction mixture was...More data for this Ligand-Target Pair
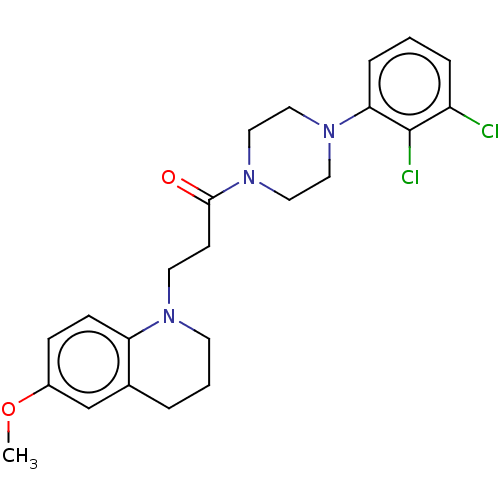
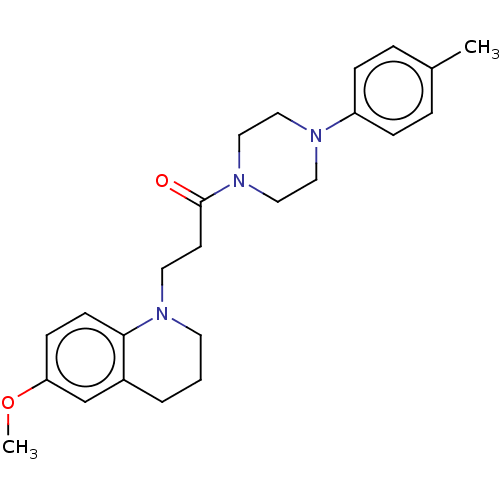
Affinity DataIC50: 1.25E+4nMAssay Description:The reaction mixture was set with template primer complex, RT enzyme and dNTPs in a lysis buffer with or without inhibitors. The reaction mixture was...More data for this Ligand-Target Pair
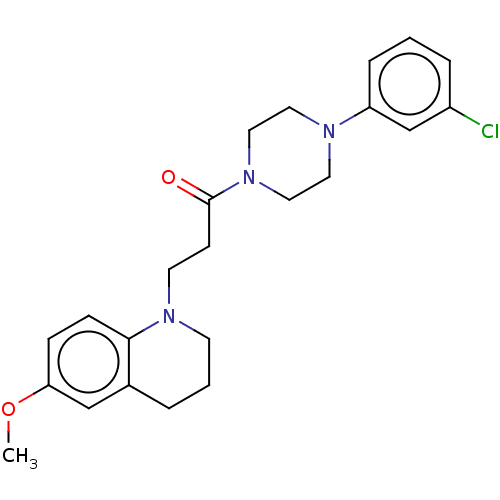
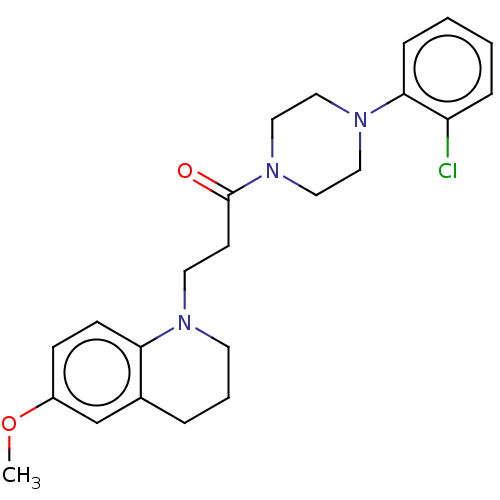
Affinity DataIC50: 1.45E+4nMAssay Description:The reaction mixture was set with template primer complex, RT enzyme and dNTPs in a lysis buffer with or without inhibitors. The reaction mixture was...More data for this Ligand-Target Pair
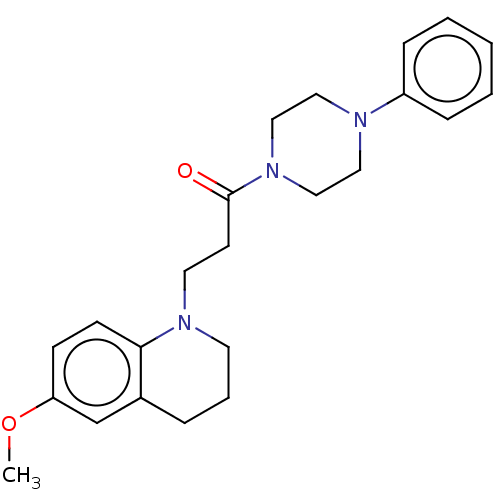
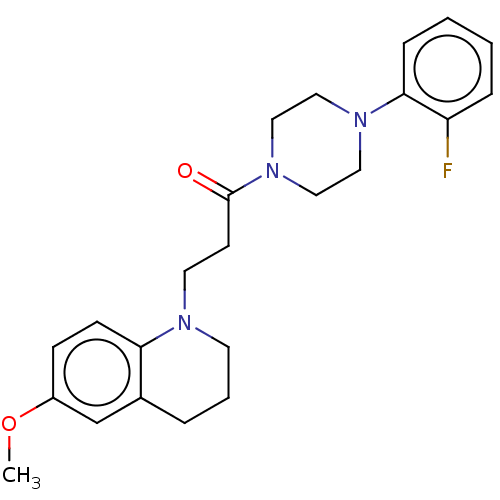
Affinity DataIC50: 1.86E+4nMAssay Description:The reaction mixture was set with template primer complex, RT enzyme and dNTPs in a lysis buffer with or without inhibitors. The reaction mixture was...More data for this Ligand-Target Pair
Affinity DataIC50: 2.43E+4nMAssay Description:The reaction mixture was set with template primer complex, RT enzyme and dNTPs in a lysis buffer with or without inhibitors. The reaction mixture was...More data for this Ligand-Target Pair
Affinity DataIC50: 2.49E+4nMAssay Description:The reaction mixture was set with template primer complex, RT enzyme and dNTPs in a lysis buffer with or without inhibitors. The reaction mixture was...More data for this Ligand-Target Pair
Affinity DataIC50: 2.85E+4nMAssay Description:The reaction mixture was set with template primer complex, RT enzyme and dNTPs in a lysis buffer with or without inhibitors. The reaction mixture was...More data for this Ligand-Target Pair
Affinity DataIC50: 3.29E+4nMAssay Description:The reaction mixture was set with template primer complex, RT enzyme and dNTPs in a lysis buffer with or without inhibitors. The reaction mixture was...More data for this Ligand-Target Pair
Affinity DataIC50: 4.75E+4nMAssay Description:The reaction mixture was set with template primer complex, RT enzyme and dNTPs in a lysis buffer with or without inhibitors. The reaction mixture was...More data for this Ligand-Target Pair
Affinity DataIC50: 4.86E+4nMAssay Description:The reaction mixture was set with template primer complex, RT enzyme and dNTPs in a lysis buffer with or without inhibitors. The reaction mixture was...More data for this Ligand-Target Pair
Affinity DataIC50: 5.28E+4nMAssay Description:The reaction mixture was set with template primer complex, RT enzyme and dNTPs in a lysis buffer with or without inhibitors. The reaction mixture was...More data for this Ligand-Target Pair
Affinity DataIC50: 5.82E+4nMAssay Description:The reaction mixture was set with template primer complex, RT enzyme and dNTPs in a lysis buffer with or without inhibitors. The reaction mixture was...More data for this Ligand-Target Pair
Affinity DataIC50: 7.85E+4nMAssay Description:The reaction mixture was set with template primer complex, RT enzyme and dNTPs in a lysis buffer with or without inhibitors. The reaction mixture was...More data for this Ligand-Target Pair
Affinity DataIC50: 8.27E+4nMAssay Description:The reaction mixture was set with template primer complex, RT enzyme and dNTPs in a lysis buffer with or without inhibitors. The reaction mixture was...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:The reaction mixture was set with template primer complex, RT enzyme and dNTPs in a lysis buffer with or without inhibitors. The reaction mixture was...More data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+5nMAssay Description:The reaction mixture was set with template primer complex, RT enzyme and dNTPs in a lysis buffer with or without inhibitors. The reaction mixture was...More data for this Ligand-Target Pair
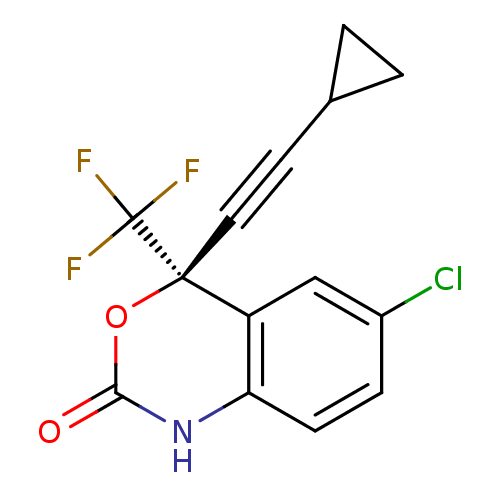
 3D Structure (crystal)
3D Structure (crystal)