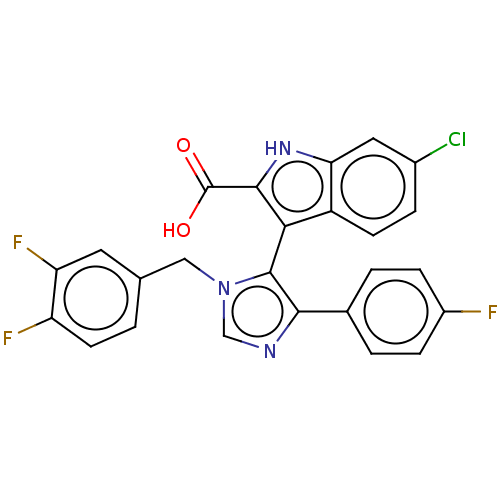
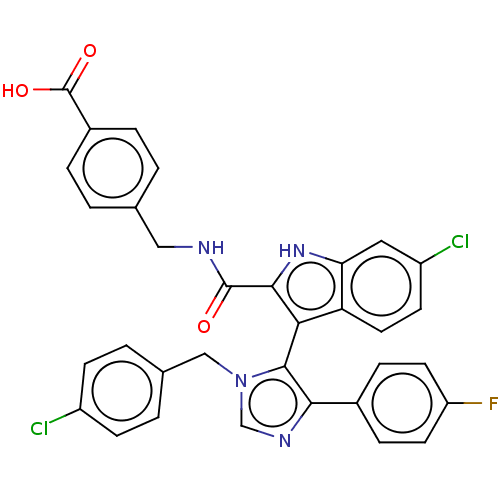
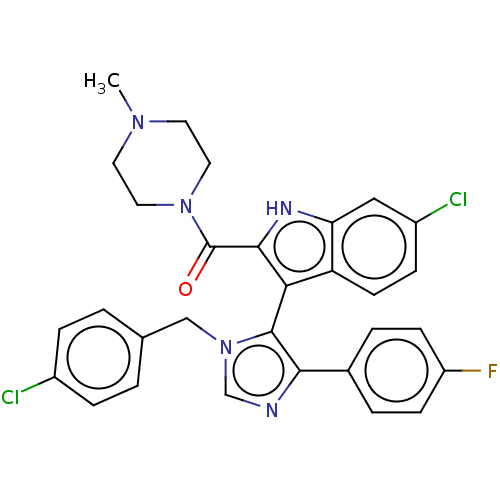
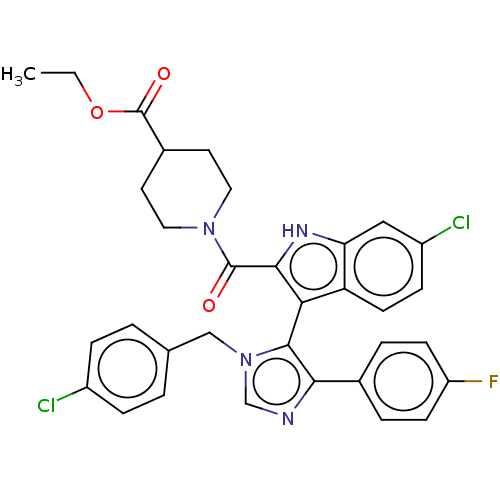
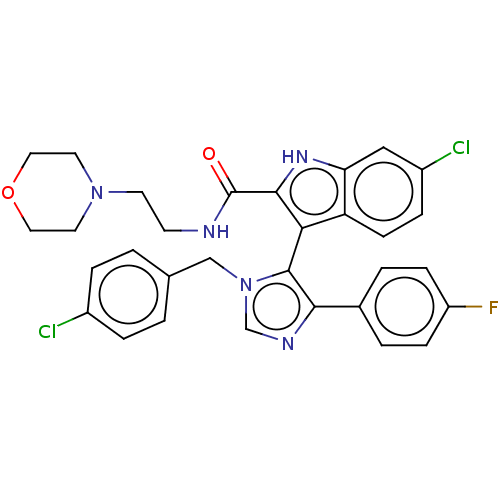
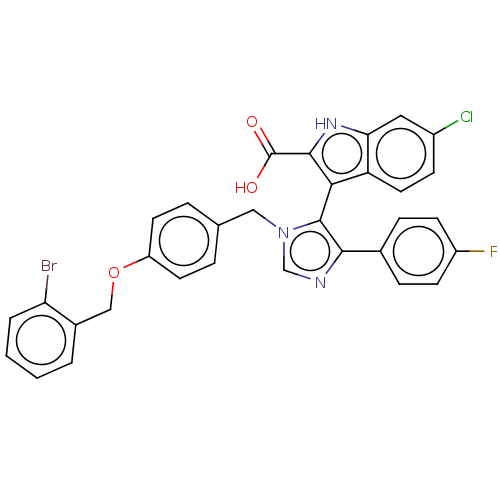
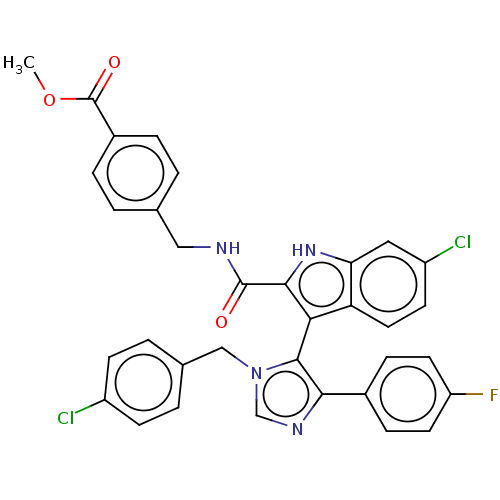
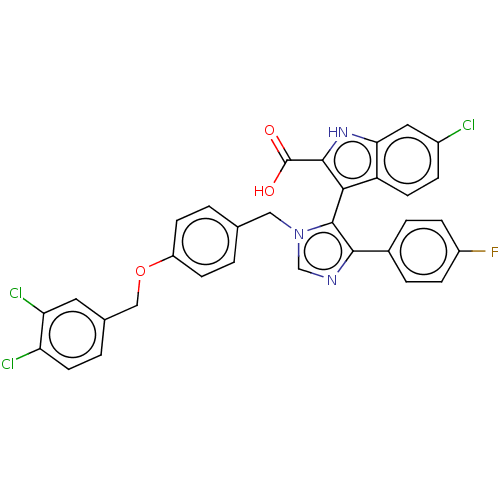
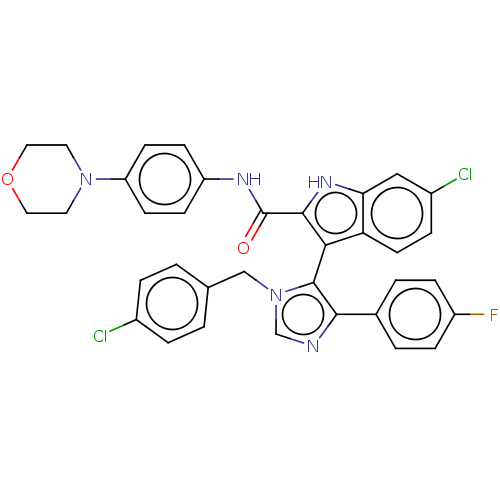
Report error Found 37 Enz. Inhib. hit(s) with all data for entry = 50000979
Affinity DataKi: 6nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 8nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 21nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 23nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 38nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 38nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 54nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 58nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 59nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 60nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 98nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 121nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 157nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 225nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 322nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 433nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 656nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKd: <1.00E+3nMAssay Description:Binding affinity to N-terminal domain of human 15N-labeled MDM2 expressed in Escherichia coli BL21 (DE3) by HSQC methodMore data for this Ligand-Target Pair
Affinity DataKi: 2.31E+3nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 3.15E+3nMAssay Description:Binding affinity to human N-terminal MDMx (1 to 134 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 4.17E+3nMAssay Description:Binding affinity to human N-terminal MDMx (1 to 134 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 4.40E+3nMAssay Description:Binding affinity to human N-terminal MDMx (1 to 134 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 4.71E+3nMAssay Description:Binding affinity to human N-terminal MDMx (1 to 134 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 4.91E+3nMAssay Description:Binding affinity to human N-terminal MDMx (1 to 134 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 6.02E+3nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 6.76E+3nMAssay Description:Binding affinity to human N-terminal MDMx (1 to 134 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 9.75E+3nMAssay Description:Binding affinity to human N-terminal MDMx (1 to 134 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 1.24E+4nMAssay Description:Binding affinity to human N-terminal MDMx (1 to 134 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 1.47E+4nMAssay Description:Binding affinity to human N-terminal MDMx (1 to 134 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 2.63E+4nMAssay Description:Binding affinity to human N-terminal MDMx (1 to 134 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 2.74E+4nMAssay Description:Binding affinity to human N-terminal MDMx (1 to 134 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 2.80E+4nMAssay Description:Binding affinity to human N-terminal MDM2 (1 to 118 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 2.96E+4nMAssay Description:Binding affinity to human N-terminal MDMx (1 to 134 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 3.53E+4nMAssay Description:Binding affinity to human N-terminal MDMx (1 to 134 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair
Affinity DataKi: 4.02E+4nMAssay Description:Binding affinity to human N-terminal MDMx (1 to 134 residues) expressed in Escherichia coli BL21 (DE3) after 15 mins in presence of 5-FAM-LTFEHYWAQLT...More data for this Ligand-Target Pair