Report error Found 133 Enz. Inhib. hit(s) with all data for entry = 50000661
Affinity DataIC50: 2nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 3nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 7nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 10nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 21nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 21nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 26nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of CYP3A4 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 31nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 31nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 34nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 36nMAssay Description:Inhibition of CYP2C9 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 37nMAssay Description:Inhibition of CYP2C9 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 39nMAssay Description:Inhibition of CYP2C9 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 46nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 57nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 83nMAssay Description:Inhibition of CYP2C9 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 100nMAssay Description:Inhibition of CYP2C9 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 119nMAssay Description:Inhibition of human NaV1.2 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 124nMAssay Description:Inhibition of rat NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp methodMore data for this Ligand-Target Pair
Affinity DataIC50: 149nMAssay Description:Inhibition of human NaV1.2 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 153nMAssay Description:Inhibition of rat NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp methodMore data for this Ligand-Target Pair
Affinity DataIC50: 173nMAssay Description:Inhibition of human NaV1.6 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 180nMAssay Description:Inhibition of CYP2C9 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 184nMAssay Description:Inhibition of CYP2C9 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 199nMAssay Description:Inhibition of human NaV1.6 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 205nMAssay Description:Inhibition of CYP2C9 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 205nMAssay Description:Inhibition of CYP3A4 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 216nMAssay Description:Inhibition of human NaV1.1 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 244nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 264nMAssay Description:Inhibition of CYP2C9 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 276nMAssay Description:Inhibition of human NaV1.2 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 279nMAssay Description:Inhibition of CYP2C9 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 292nMAssay Description:Inhibition of human NaV1.3 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 299nMAssay Description:Inhibition of CYP2C9 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 321nMAssay Description:Inhibition of CYP2C9 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 324nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 325nMAssay Description:Inhibition of human NaV1.7 expressed in HEK cells assessed as half inactivation potential at -120 mV holding potential by automated patch clamp metho...More data for this Ligand-Target Pair
Affinity DataIC50: 372nMAssay Description:Inhibition of CYP2C9 (unknown origin)More data for this Ligand-Target Pair
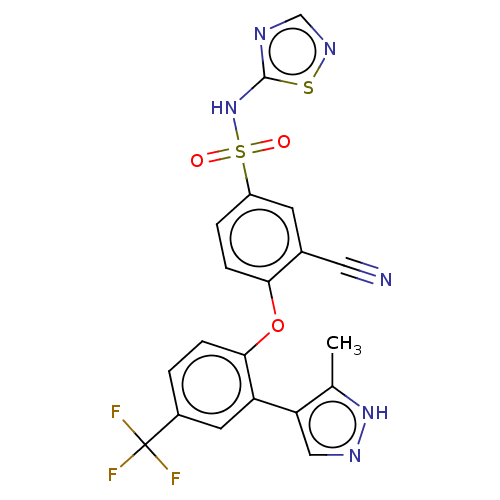
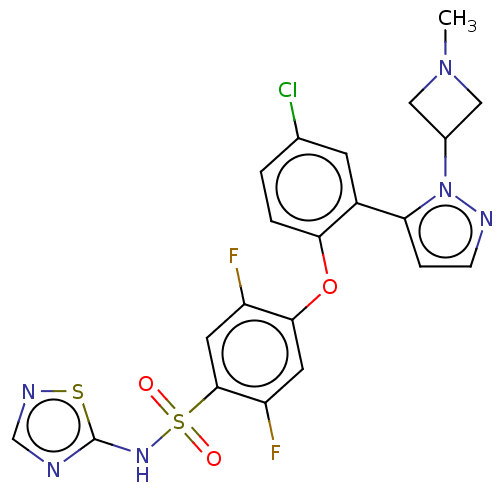
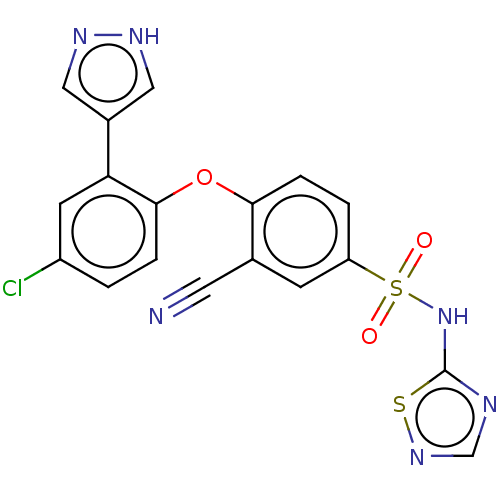
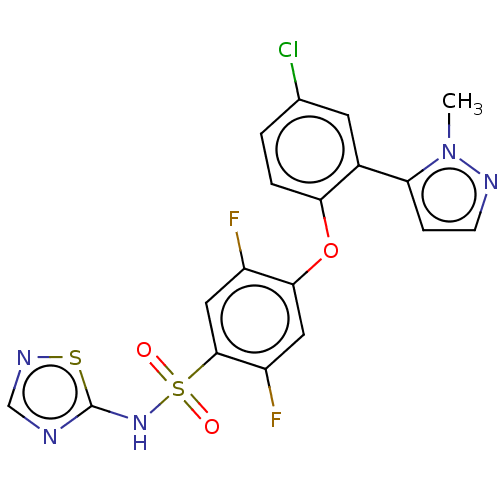
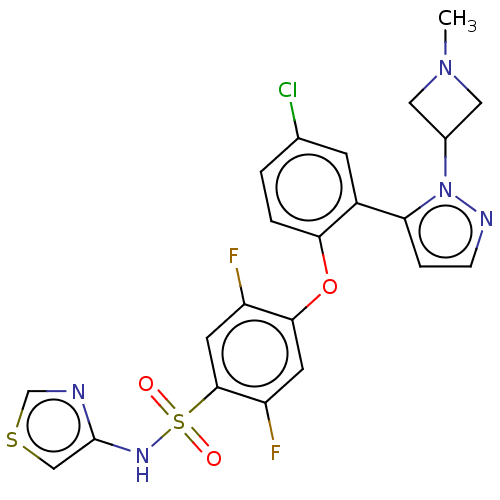
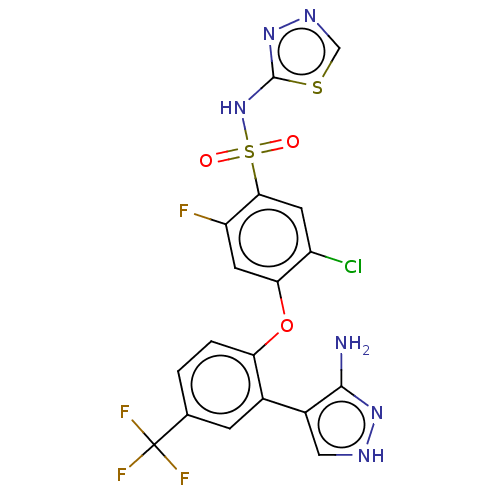
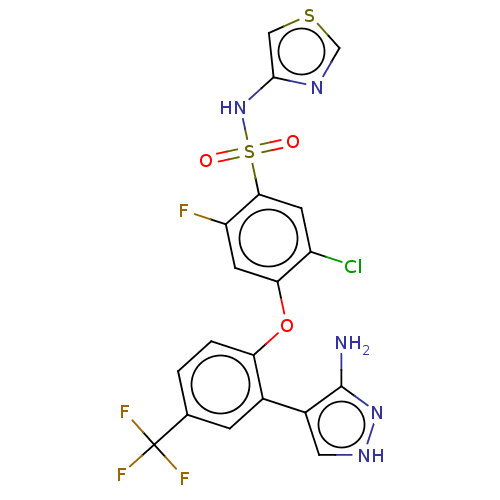
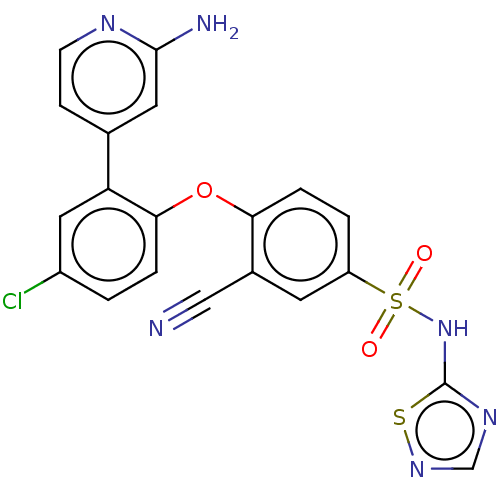
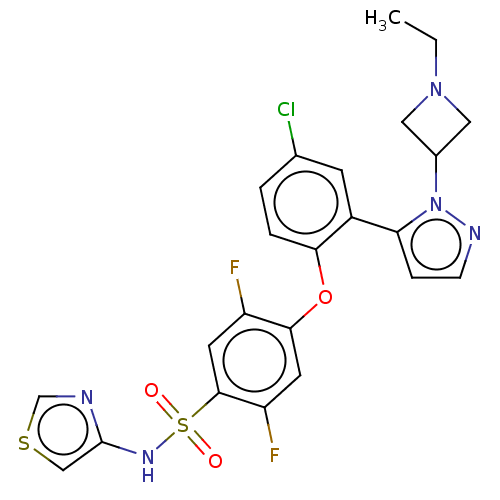
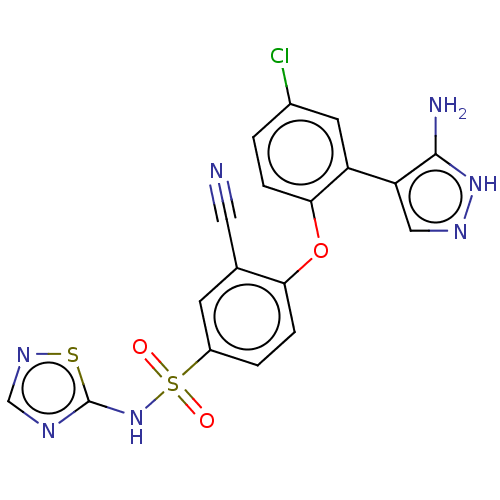

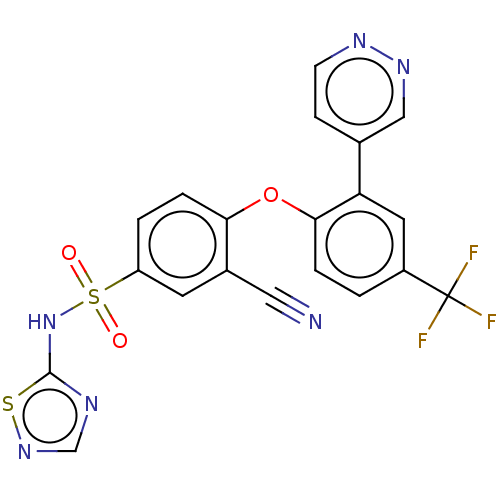
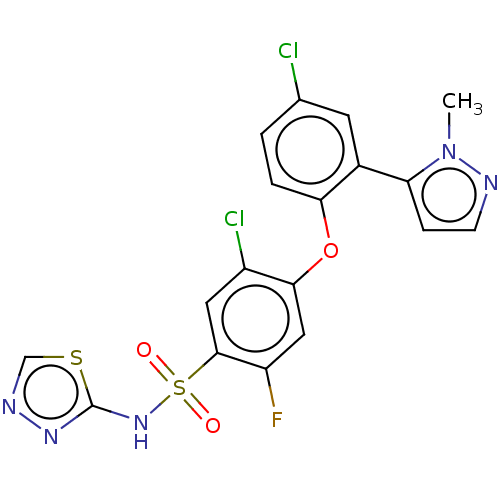
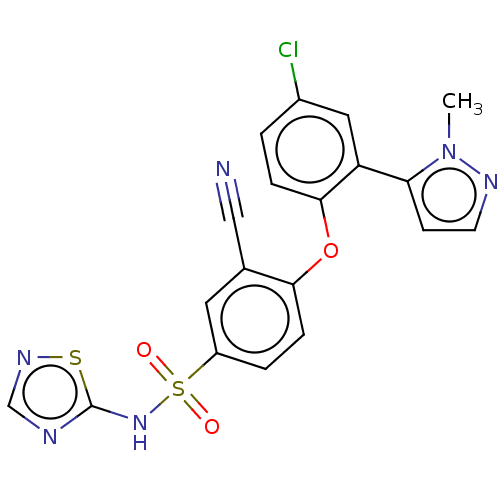
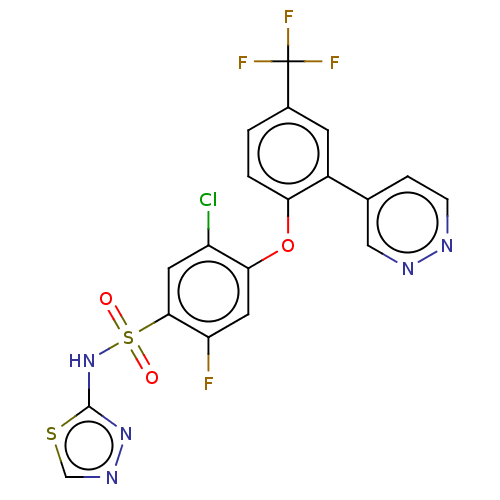
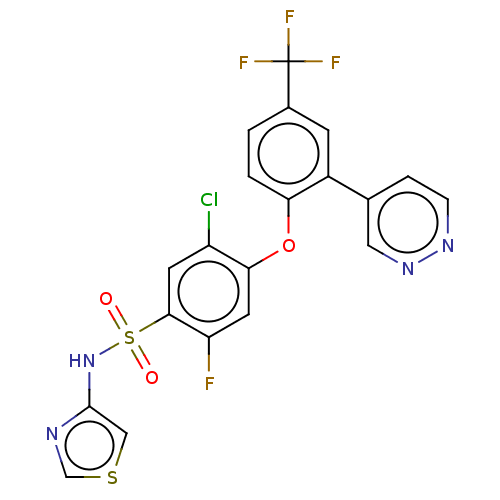
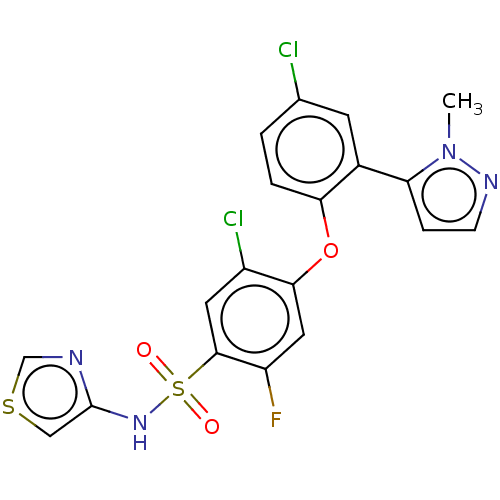
 3D Structure (crystal)
3D Structure (crystal)