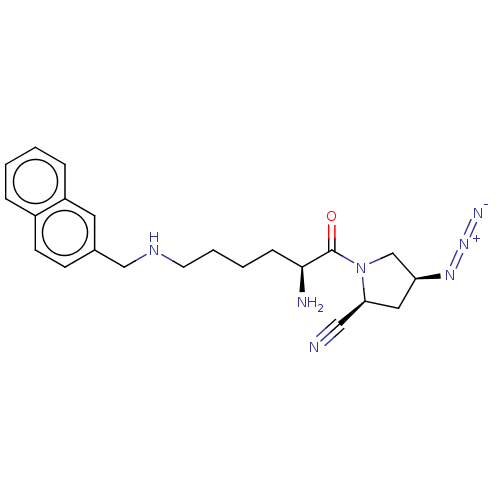
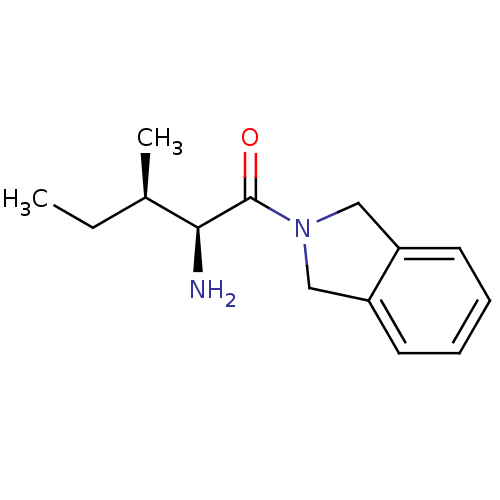
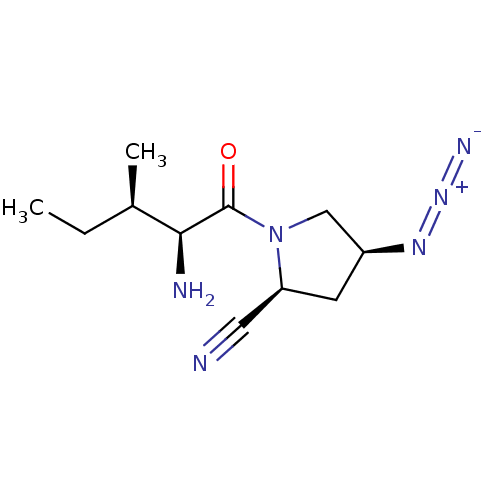
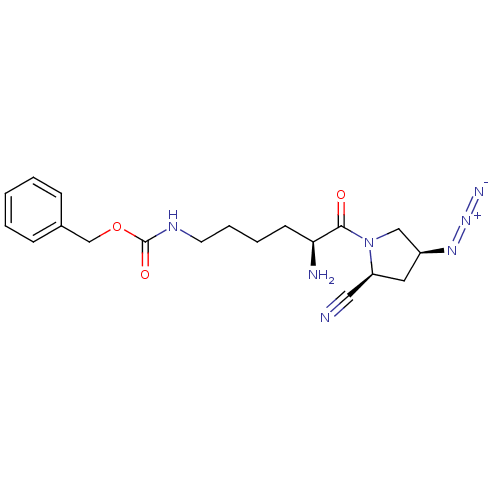
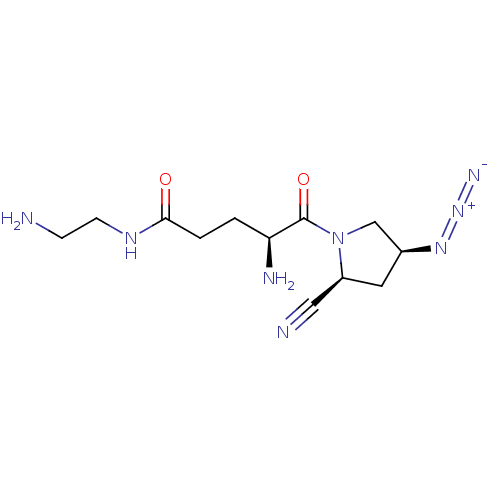
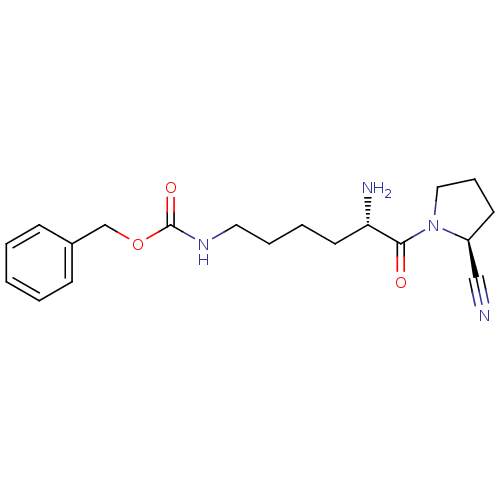
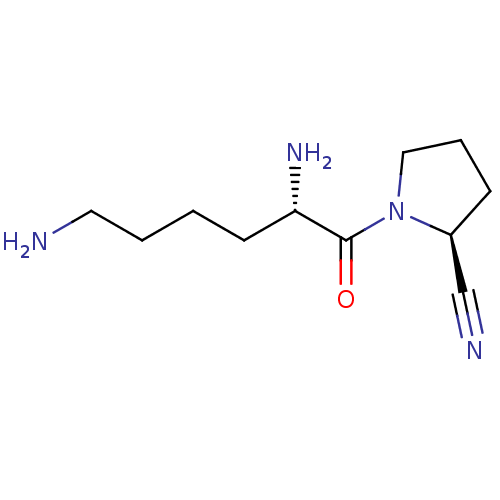
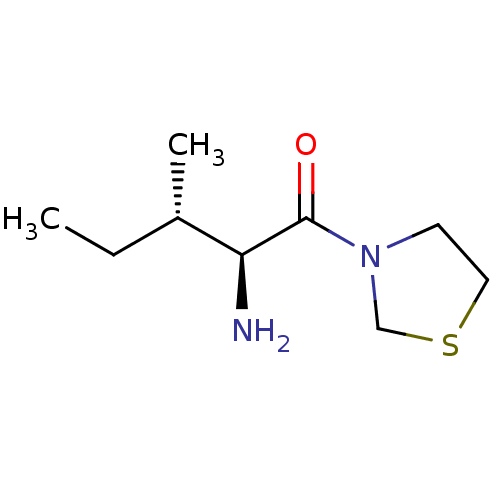
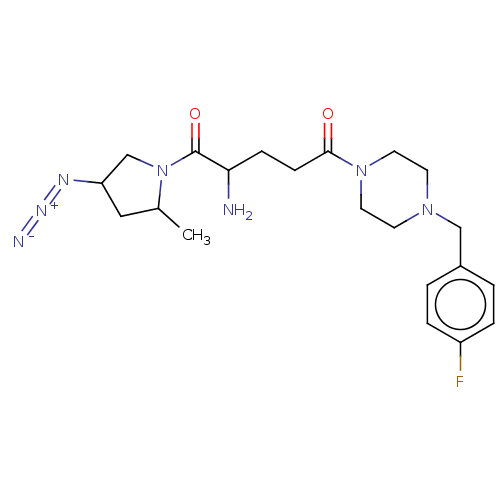
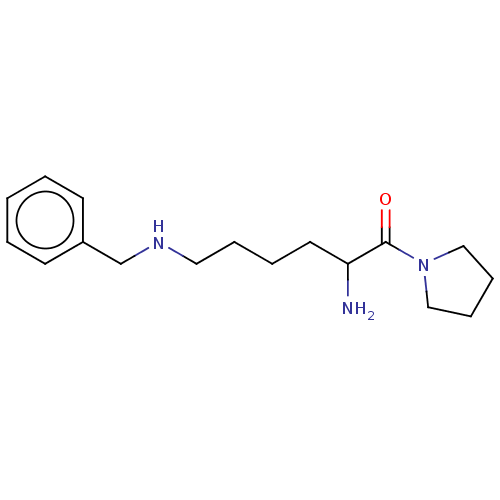
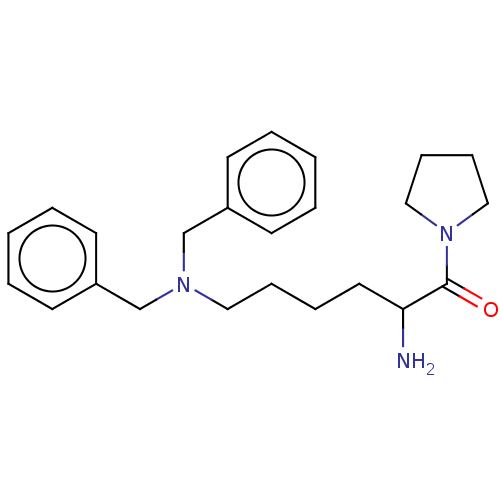
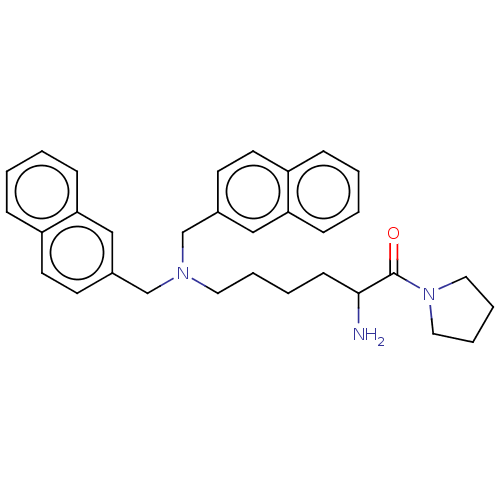
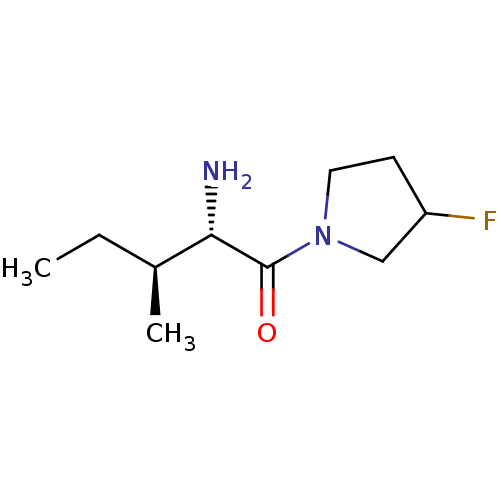
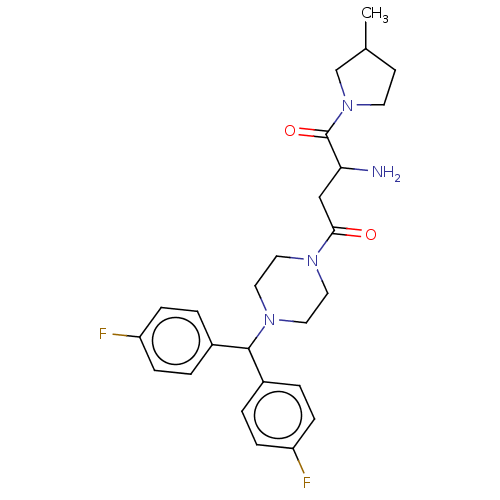
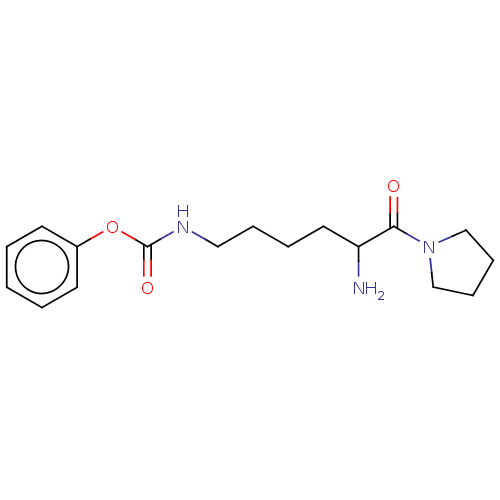
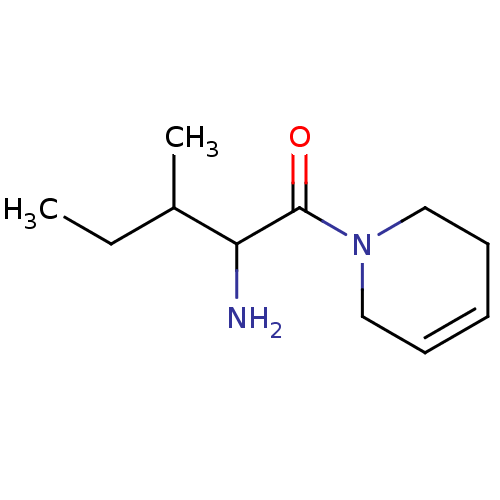
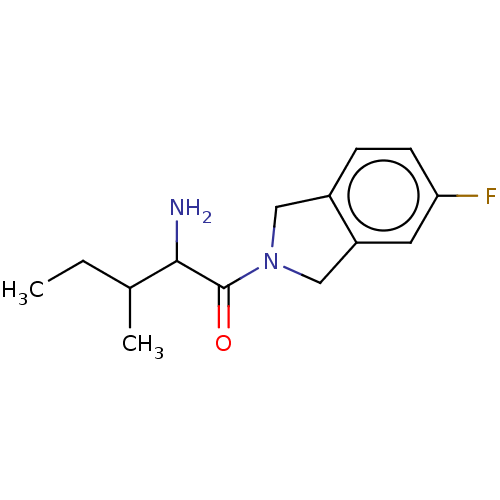
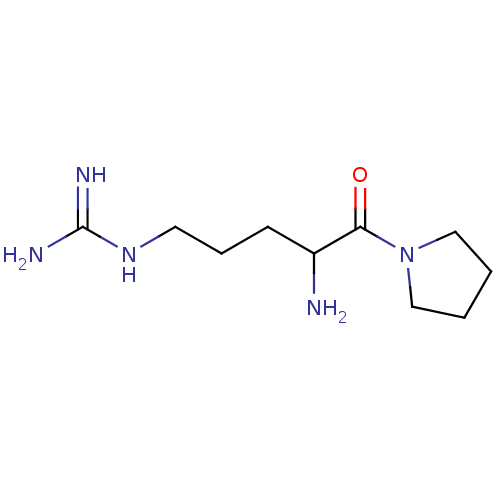
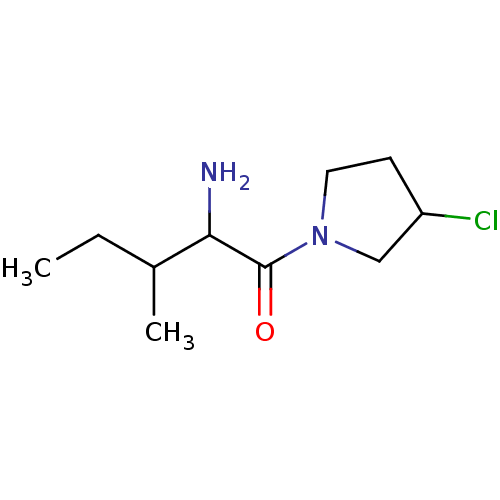
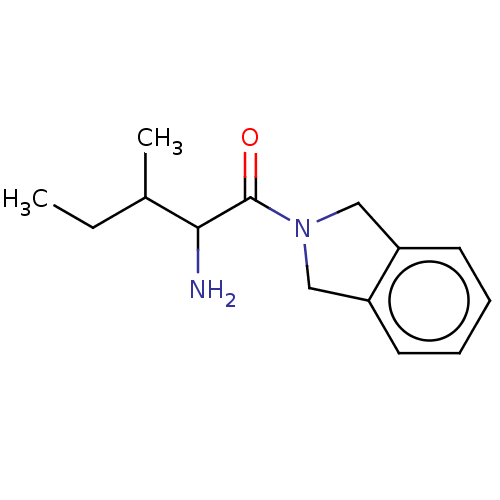
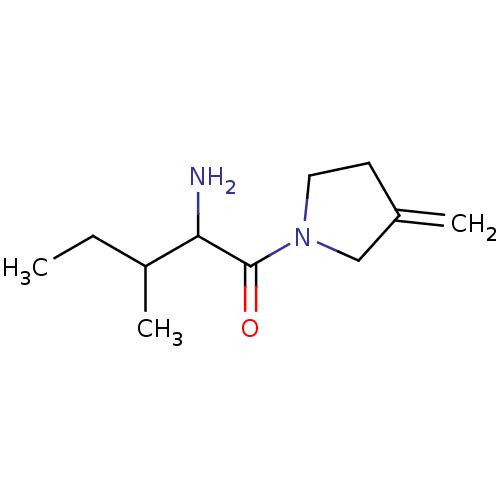
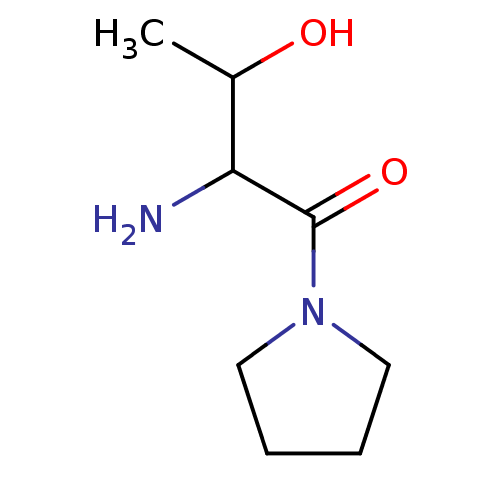
Report error Found 108 Enz. Inhib. hit(s) with all data for entry = 50001366
Affinity DataIC50: 0.5nMAssay Description:Inhibition of DPP2 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 2nMAssay Description:Inhibition of human DPP4More data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrate preincubate...More data for this Ligand-Target Pair
Affinity DataIC50: 22nMAssay Description:Inhibition of human DPP4More data for this Ligand-Target Pair
Affinity DataIC50: 40nMAssay Description:Inhibition of human DPP4More data for this Ligand-Target Pair
Affinity DataIC50: 51nMAssay Description:Inhibition of DPP8 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 79nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 86nMAssay Description:Inhibition of DPP2 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Inhibition of human DPP4More data for this Ligand-Target Pair
Affinity DataIC50: 120nMAssay Description:Inhibition of DPP8 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 140nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 230nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 290nMAssay Description:Inhibition of bovine DPP9More data for this Ligand-Target Pair
Affinity DataIC50: 300nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 310nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 330nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 480nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 580nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 680nMAssay Description:Inhibition of bovine DPP9More data for this Ligand-Target Pair
Affinity DataIC50: 800nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+3nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.50E+3nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.60E+3nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.70E+3nMAssay Description:Inhibition of human DPP4More data for this Ligand-Target Pair
Affinity DataIC50: 2.40E+3nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 3.10E+3nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 3.40E+3nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 3.50E+3nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 5.20E+3nMAssay Description:Inhibition of DPP8 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 5.40E+3nMAssay Description:Inhibition of human DPP4More data for this Ligand-Target Pair
Affinity DataIC50: 6.00E+3nMAssay Description:Inhibition of DPP8 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 6.60E+3nMAssay Description:Inhibition of bovine DPP9More data for this Ligand-Target Pair
Affinity DataIC50: 8.00E+3nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 9.00E+3nMAssay Description:Inhibition of DPP8 (unknown origin)More data for this Ligand-Target Pair
Affinity DataIC50: 9.40E+3nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Porphyromonas gingivalis N-terminal His-tagged DPP4 expressed in Escherichia coli using Gly-Pro-p-nitroanilide as substrateMore data for this Ligand-Target Pair