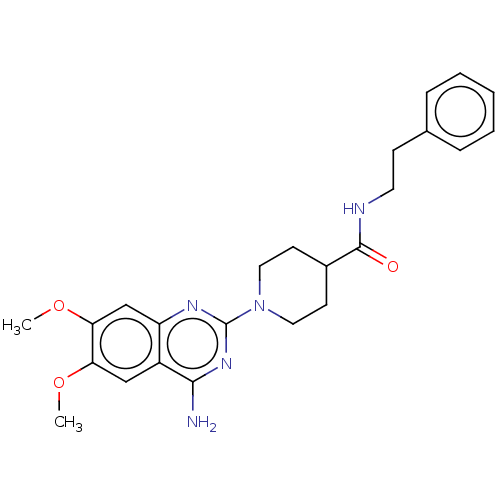
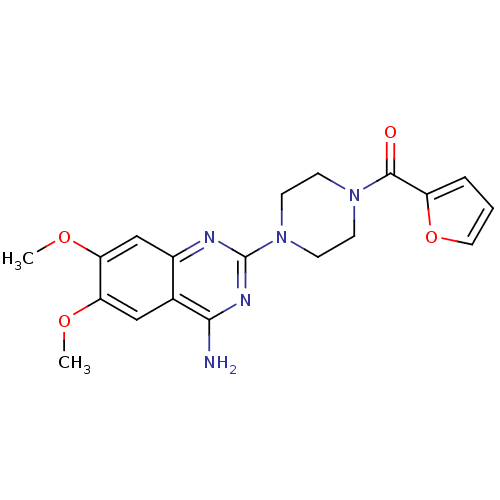
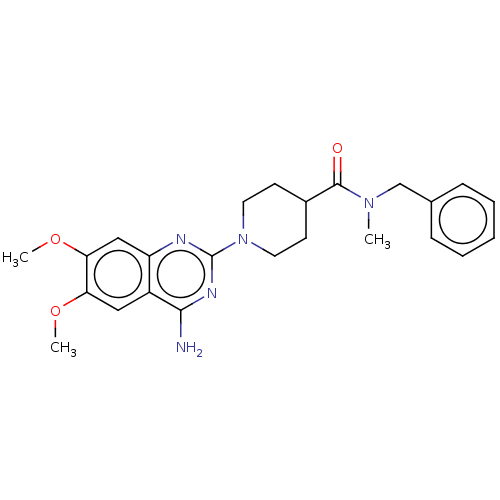
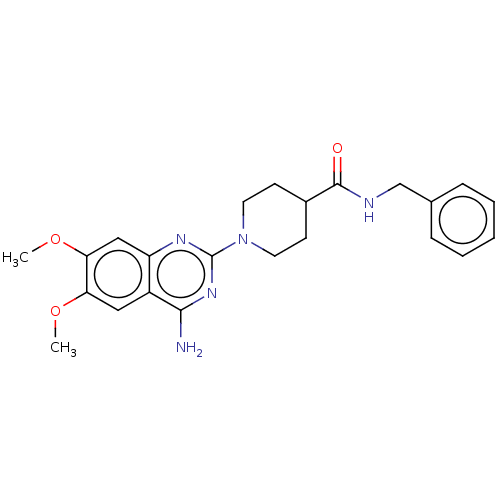
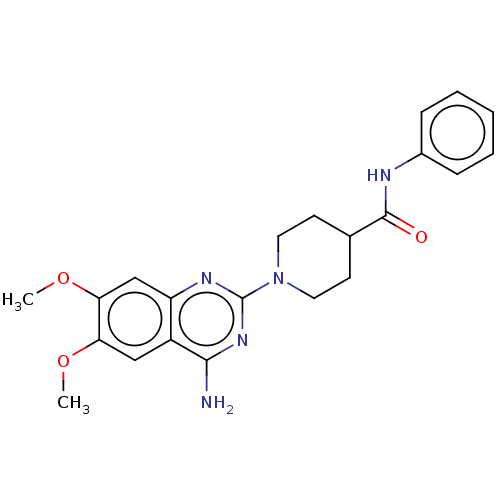
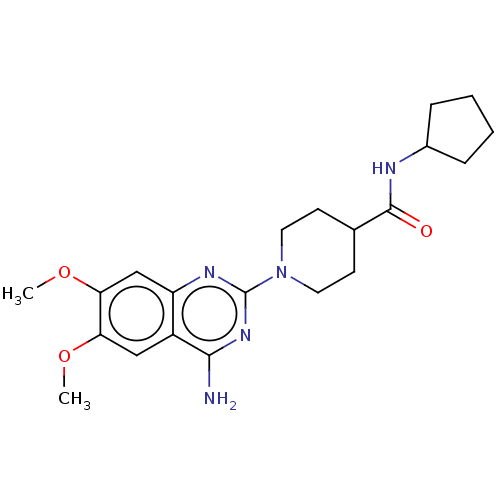
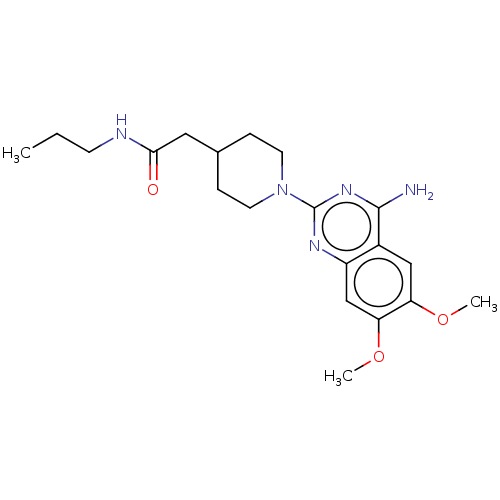
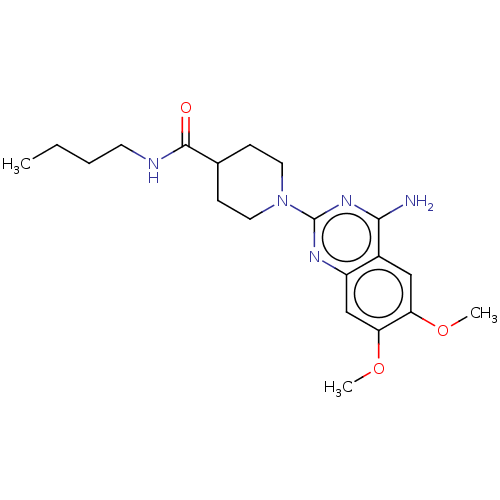
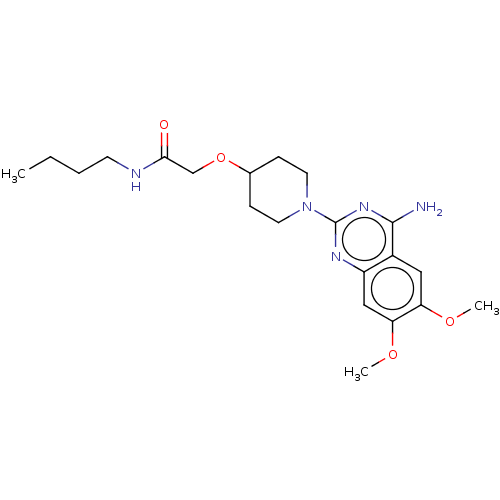
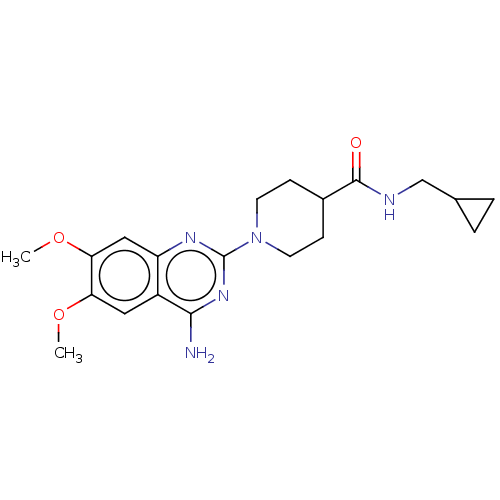
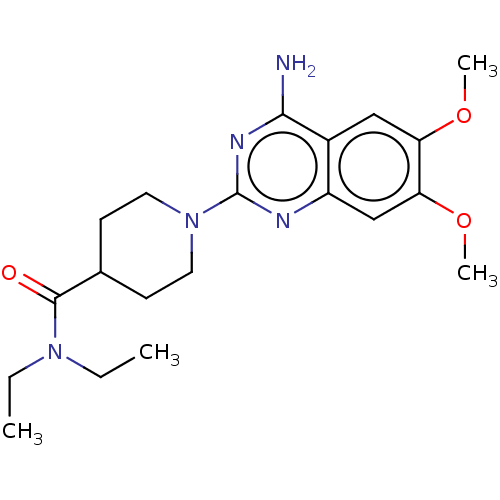
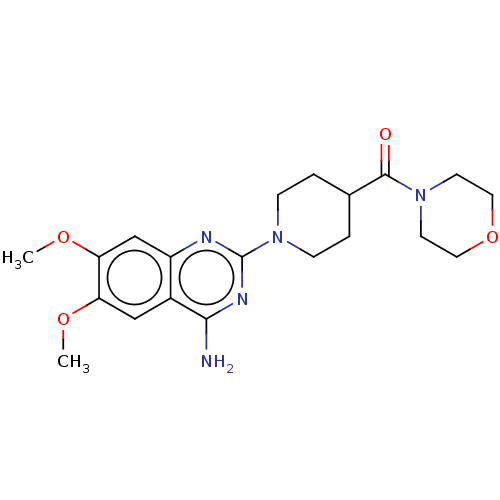
Report error Found 34 Enz. Inhib. hit(s) with all data for entry = 50048753
Affinity DataKi: 0.100nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 0.190nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 0.200nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 0.220nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 0.420nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 0.470nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 0.470nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 0.630nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 0.650nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 0.670nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 0.790nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 0.900nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 1.10nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 1.40nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 1.5nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 1.60nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 1.70nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
Affinity DataKi: 8.30nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataKi: 53nMAssay Description:In vitro binding affinity determined for alpha-2 adrenergic receptor by the percentage displacement of [3H]- clonidine at 10e-6 MMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataKi: 54nMAssay Description:In vitro binding affinity determined for alpha-2 adrenergic receptor by the percentage displacement of [3H]- clonidine at 10e-6 MMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataKi: 55nMAssay Description:In vitro binding affinity determined for alpha-2 adrenergic receptor by the percentage displacement of [3H]- clonidine at 10e-6 MMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataKi: 56nMAssay Description:In vitro binding affinity determined for alpha-2 adrenergic receptor by the percentage displacement of [3H]- clonidine at 10e-6 MMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataKi: 56nMAssay Description:In vitro binding affinity determined for alpha-2 adrenergic receptor by the percentage displacement of [3H]- clonidine at 10e-6 MMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataKi: 61nMAssay Description:In vitro binding affinity determined for alpha-2 adrenergic receptor by the percentage displacement of [3H]- clonidine at 10e-6 MMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataKi: 62nMAssay Description:In vitro binding affinity determined for alpha-2 adrenergic receptor by the percentage displacement of [3H]- clonidine at 10e-6 MMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataKi: 63nMAssay Description:In vitro binding affinity determined for alpha-2 adrenergic receptor by the percentage displacement of [3H]- clonidine at 10e-6 MMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataKi: 65nMAssay Description:In vitro binding affinity determined for alpha-2 adrenergic receptor by the percentage displacement of [3H]- clonidine at 10e-6 MMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataKi: 68nMAssay Description:In vitro binding affinity determined for alpha-2 adrenergic receptor by the percentage displacement of [3H]- clonidine at 10e-6 MMore data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataKi: 71nMAssay Description:In vitro binding affinity determined for alpha-2 adrenergic receptor by the percentage displacement of [3H]- clonidine at 10e-6 MMore data for this Ligand-Target Pair
Affinity DataKi: 117nMAssay Description:In vitro binding affinity determined for alpha-1 adrenergic receptor by the displacement of [3H]- prazosin.More data for this Ligand-Target Pair
TargetAlpha-2A adrenergic receptor [16-465]/Alpha-2B adrenergic receptor/Alpha-2C adrenergic receptor(Rat)
TBA
Curated by ChEMBL
TBA
Curated by ChEMBL
Affinity DataKi: 4.83E+3nMAssay Description:In vitro binding affinity determined for alpha-2 adrenergic receptor by the percentage displacement of [3H]- clonidine at 10e-6 MMore data for this Ligand-Target Pair