Report error Found 15 Enz. Inhib. hit(s) with all data for entry = 50012234
TargetBroad substrate specificity ATP-binding cassette transporter ABCG2(Human)
Cnrs
Curated by ChEMBL
Cnrs
Curated by ChEMBL
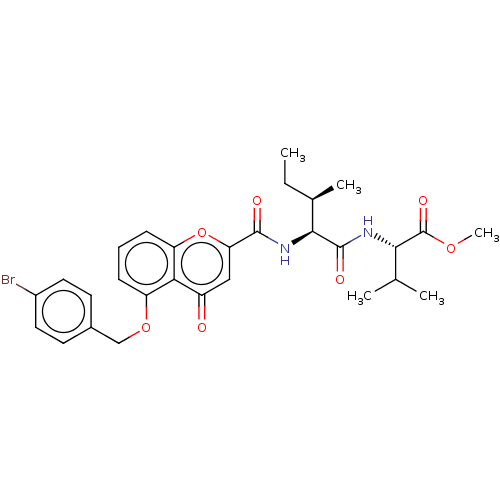
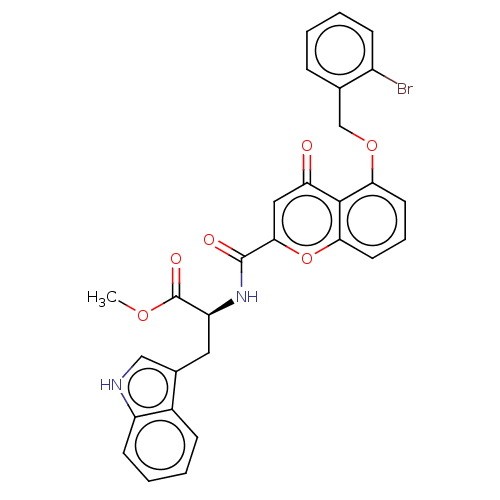
Affinity DataEC50: 50nMAssay Description:Inhibition of ABCG2 (unknown origin) in HEK293 cells assessed as maximal inhibition of mitoxantrone efflux by fluorescence based flow cytometryMore data for this Ligand-Target Pair
TargetBroad substrate specificity ATP-binding cassette transporter ABCG2(Human)
Cnrs
Curated by ChEMBL
Cnrs
Curated by ChEMBL
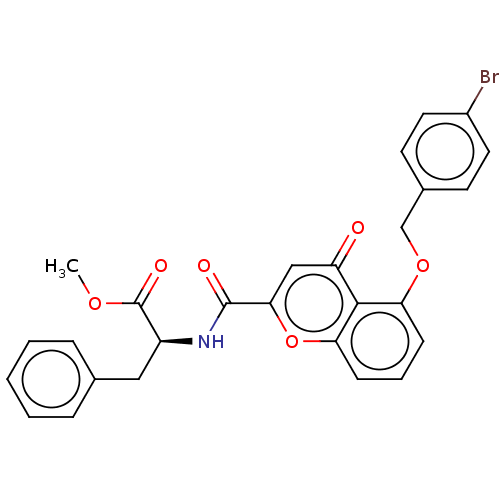
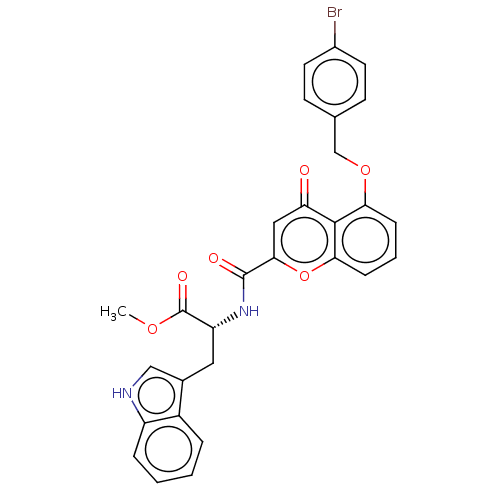
Affinity DataEC50: 70nMAssay Description:Inhibition of ABCG2 (unknown origin) in HEK293 cells assessed as maximal inhibition of mitoxantrone efflux by fluorescence based flow cytometryMore data for this Ligand-Target Pair
TargetBroad substrate specificity ATP-binding cassette transporter ABCG2(Human)
Cnrs
Curated by ChEMBL
Cnrs
Curated by ChEMBL
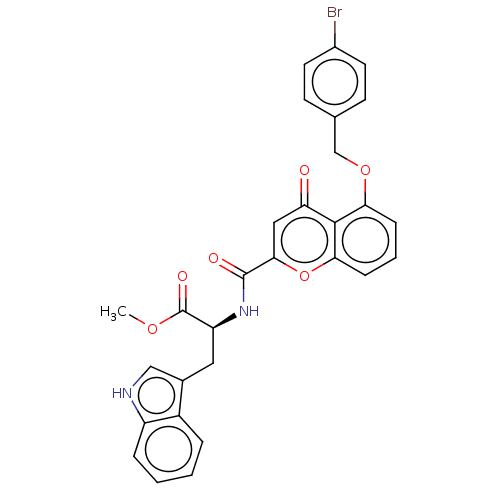
Affinity DataEC50: 70nMAssay Description:Inhibition of ABCG2 (unknown origin) in HEK293 cells assessed as maximal inhibition of mitoxantrone efflux by fluorescence based flow cytometryMore data for this Ligand-Target Pair
TargetBroad substrate specificity ATP-binding cassette transporter ABCG2(Human)
Cnrs
Curated by ChEMBL
Cnrs
Curated by ChEMBL
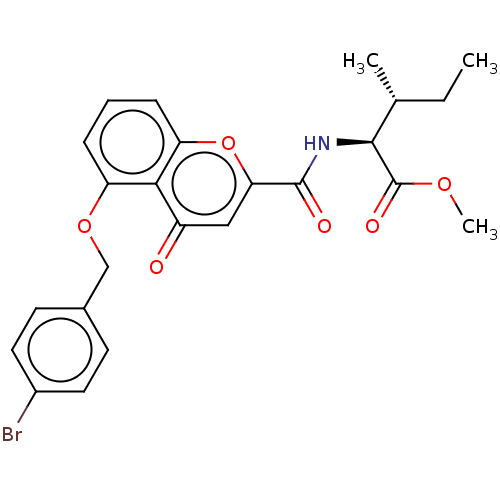
Affinity DataEC50: 90nMAssay Description:Inhibition of ABCG2 (unknown origin) in HEK293 cells assessed as maximal inhibition of mitoxantrone efflux by fluorescence based flow cytometryMore data for this Ligand-Target Pair
TargetBroad substrate specificity ATP-binding cassette transporter ABCG2(Human)
Cnrs
Curated by ChEMBL
Cnrs
Curated by ChEMBL
Affinity DataEC50: 100nMAssay Description:Inhibition of ABCG2 (unknown origin) in HEK293 cells assessed as maximal inhibition of mitoxantrone efflux by fluorescence based flow cytometryMore data for this Ligand-Target Pair
TargetBroad substrate specificity ATP-binding cassette transporter ABCG2(Human)
Cnrs
Curated by ChEMBL
Cnrs
Curated by ChEMBL
Affinity DataEC50: 100nMAssay Description:Inhibition of ABCG2 (unknown origin) in HEK293 cells assessed as maximal inhibition of mitoxantrone efflux by fluorescence based flow cytometryMore data for this Ligand-Target Pair
TargetBroad substrate specificity ATP-binding cassette transporter ABCG2(Human)
Cnrs
Curated by ChEMBL
Cnrs
Curated by ChEMBL
Affinity DataEC50: 110nMAssay Description:Inhibition of ABCG2 (unknown origin) in HEK293 cells assessed as maximal inhibition of mitoxantrone efflux by fluorescence based flow cytometryMore data for this Ligand-Target Pair
TargetBroad substrate specificity ATP-binding cassette transporter ABCG2(Human)
Cnrs
Curated by ChEMBL
Cnrs
Curated by ChEMBL
Affinity DataEC50: 130nMAssay Description:Inhibition of ABCG2 (unknown origin) in HEK293 cells assessed as maximal inhibition of mitoxantrone efflux by fluorescence based flow cytometryMore data for this Ligand-Target Pair
TargetBroad substrate specificity ATP-binding cassette transporter ABCG2(Human)
Cnrs
Curated by ChEMBL
Cnrs
Curated by ChEMBL
Affinity DataEC50: 140nMAssay Description:Inhibition of ABCG2 (unknown origin) in HEK293 cells assessed as maximal inhibition of mitoxantrone efflux by fluorescence based flow cytometryMore data for this Ligand-Target Pair
TargetBroad substrate specificity ATP-binding cassette transporter ABCG2(Human)
Cnrs
Curated by ChEMBL
Cnrs
Curated by ChEMBL
Affinity DataEC50: 250nMAssay Description:Inhibition of ABCG2 (unknown origin) in HEK293 cells assessed as maximal inhibition of mitoxantrone efflux by fluorescence based flow cytometryMore data for this Ligand-Target Pair
TargetBroad substrate specificity ATP-binding cassette transporter ABCG2(Human)
Cnrs
Curated by ChEMBL
Cnrs
Curated by ChEMBL
Affinity DataEC50: 270nMAssay Description:Inhibition of ABCG2 (unknown origin) in HEK293 cells assessed as maximal inhibition of mitoxantrone efflux by fluorescence based flow cytometryMore data for this Ligand-Target Pair
TargetBroad substrate specificity ATP-binding cassette transporter ABCG2(Human)
Cnrs
Curated by ChEMBL
Cnrs
Curated by ChEMBL
Affinity DataEC50: 480nMAssay Description:Inhibition of ABCG2 (unknown origin) in HEK293 cells assessed as maximal inhibition of mitoxantrone efflux by fluorescence based flow cytometryMore data for this Ligand-Target Pair
TargetBroad substrate specificity ATP-binding cassette transporter ABCG2(Human)
Cnrs
Curated by ChEMBL
Cnrs
Curated by ChEMBL
Affinity DataEC50: 590nMAssay Description:Inhibition of ABCG2 (unknown origin) in HEK293 cells assessed as maximal inhibition of mitoxantrone efflux by fluorescence based flow cytometryMore data for this Ligand-Target Pair
TargetBroad substrate specificity ATP-binding cassette transporter ABCG2(Human)
Cnrs
Curated by ChEMBL
Cnrs
Curated by ChEMBL
Affinity DataEC50: 600nMAssay Description:Inhibition of ABCG2 (unknown origin) in HEK293 cells assessed as maximal inhibition of mitoxantrone efflux by fluorescence based flow cytometryMore data for this Ligand-Target Pair
TargetBroad substrate specificity ATP-binding cassette transporter ABCG2(Human)
Cnrs
Curated by ChEMBL
Cnrs
Curated by ChEMBL
Affinity DataEC50: 2.90E+4nMAssay Description:Inhibition of ABCG2 (unknown origin) in HEK293 cells assessed as maximal inhibition of mitoxantrone efflux by fluorescence based flow cytometryMore data for this Ligand-Target Pair
 3D Structure (crystal)
3D Structure (crystal)