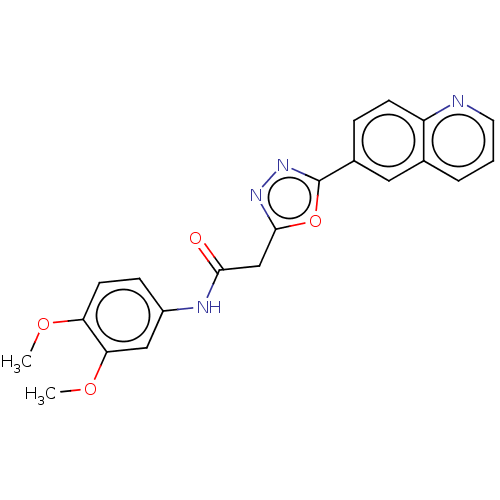
Report error Found 23 Enz. Inhib. hit(s) with all data for entry = 50012665
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 9.5nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 139nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 190nMAssay Description:Inhibition of PI3Kbeta (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 1.15E+3nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 2.60E+3nMAssay Description:Inhibition of PI3Kbeta (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit beta isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 4.10E+3nMAssay Description:Inhibition of PI3Kbeta (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 6.60E+3nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 7.60E+3nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 9.80E+3nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 1.42E+4nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 1.56E+4nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit delta isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 1.66E+4nMAssay Description:Inhibition of PI3Kdelta (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 1.70E+4nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 1.83E+4nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 2.92E+4nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit gamma isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 1.52E+5nMAssay Description:Inhibition of PI3Kgamma (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair
TargetPhosphatidylinositol 4,5-bisphosphate 3-kinase catalytic subunit alpha isoform(Human)
Zhejiang University
Curated by ChEMBL
Zhejiang University
Curated by ChEMBL
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of PI3Kalpha (unknown origin) using PIP2 as substrate measured after 1 hr by ADP-glo plus luminescence assayMore data for this Ligand-Target Pair

 3D Structure (crystal)
3D Structure (crystal)