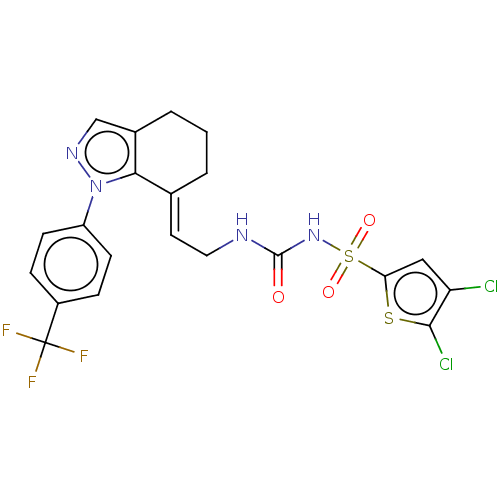
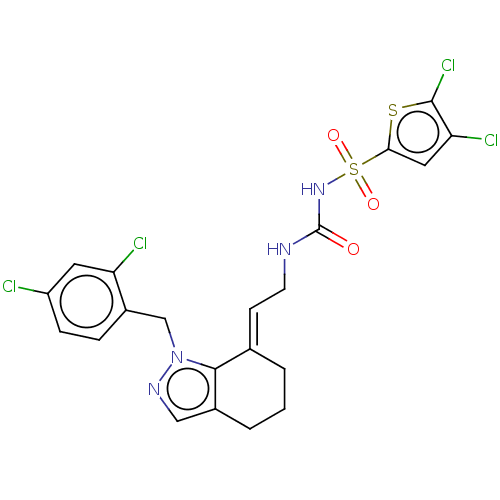
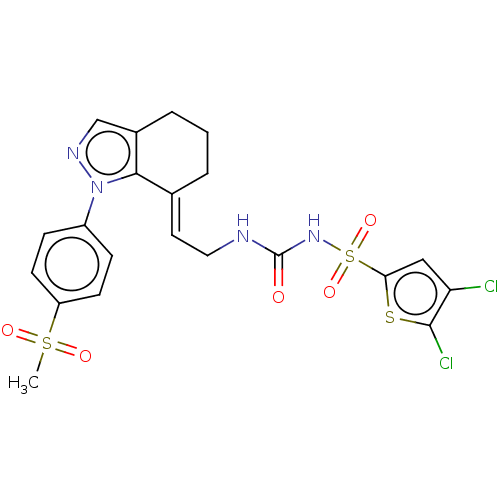
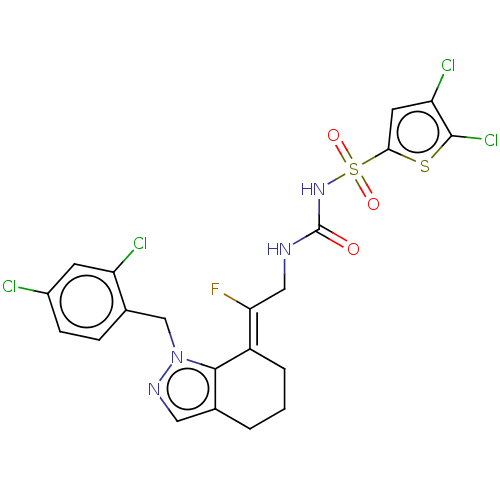
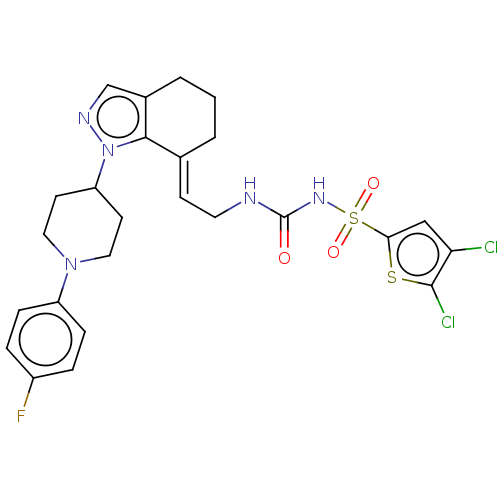
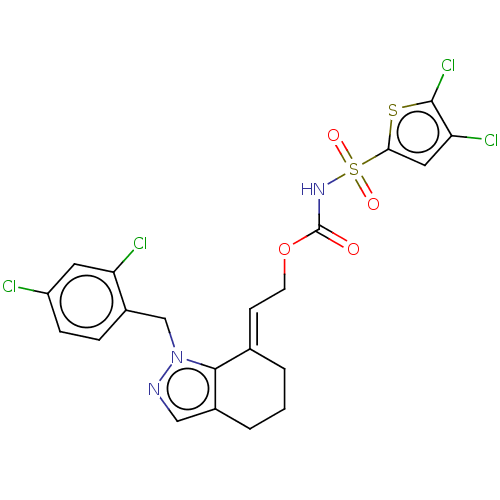
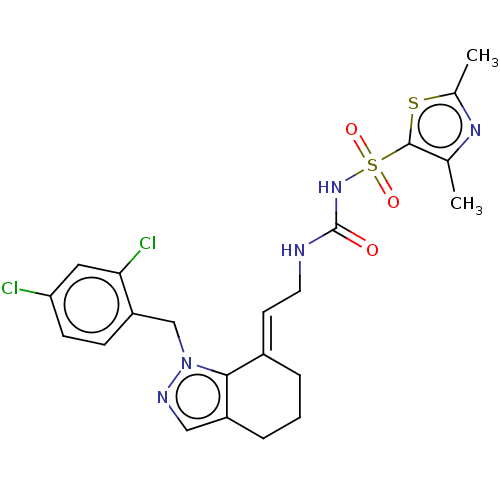
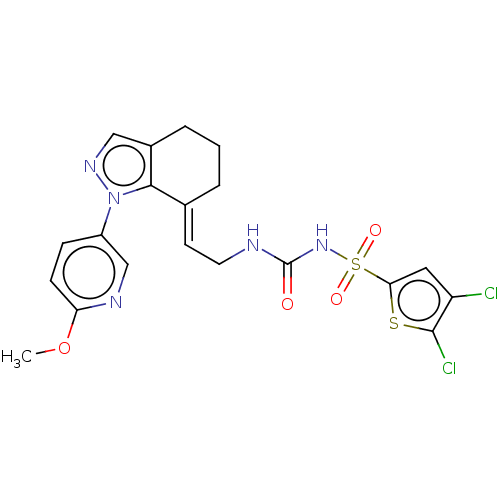
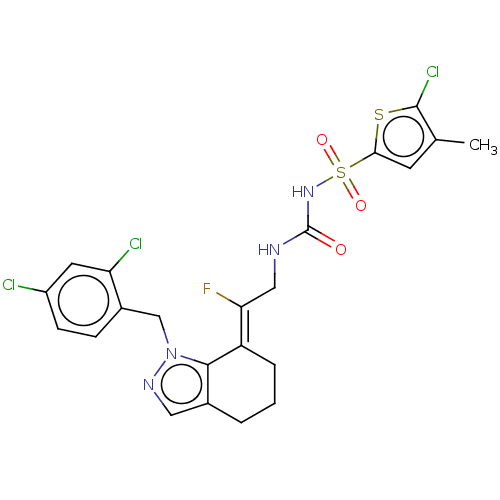
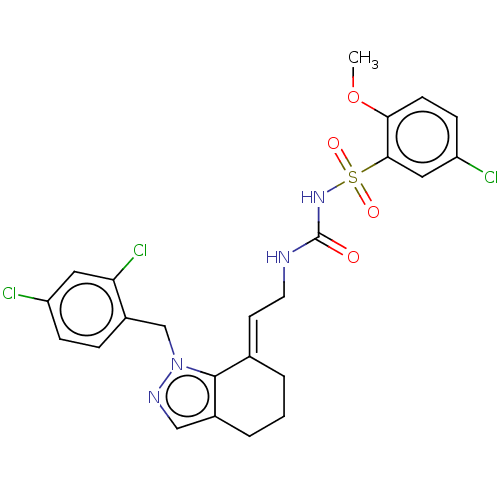
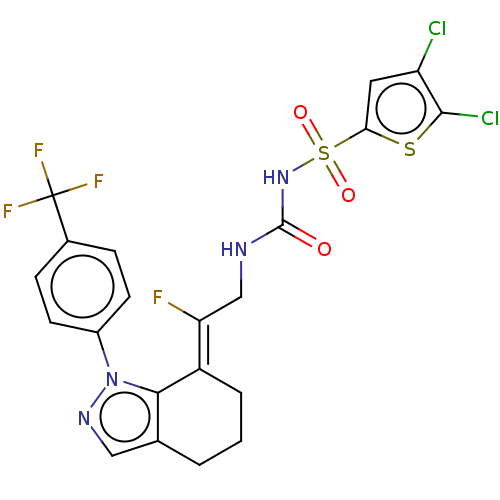
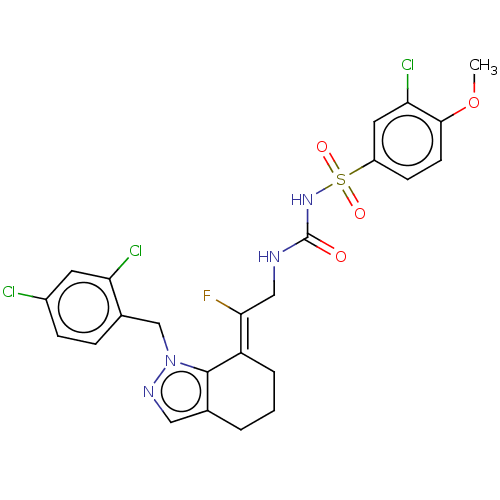
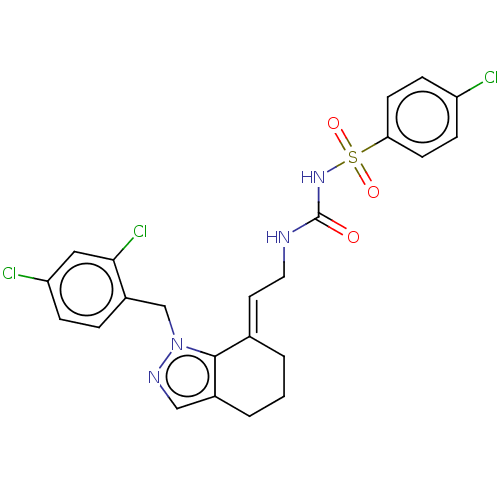
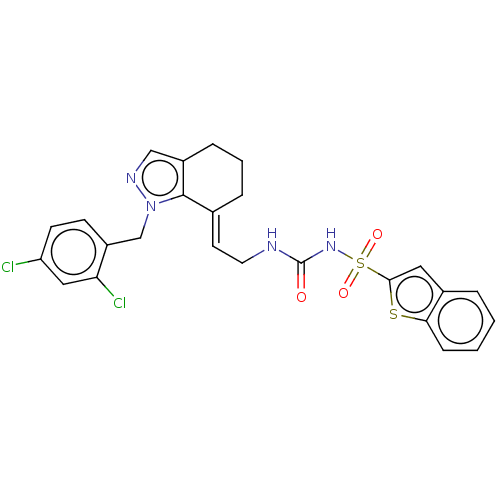
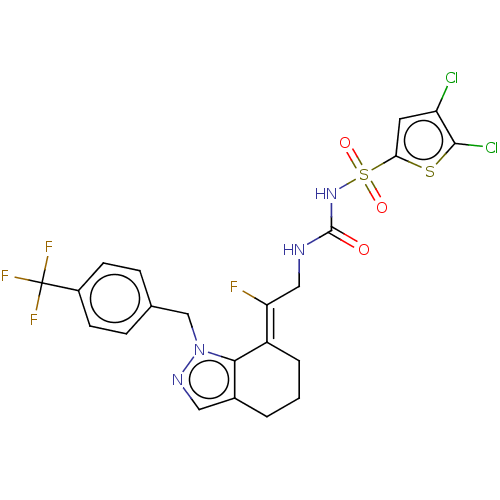
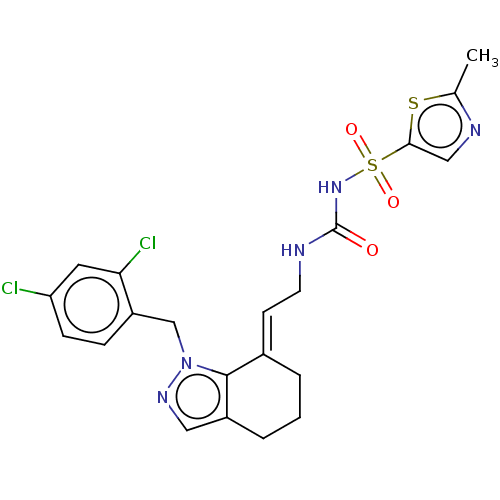
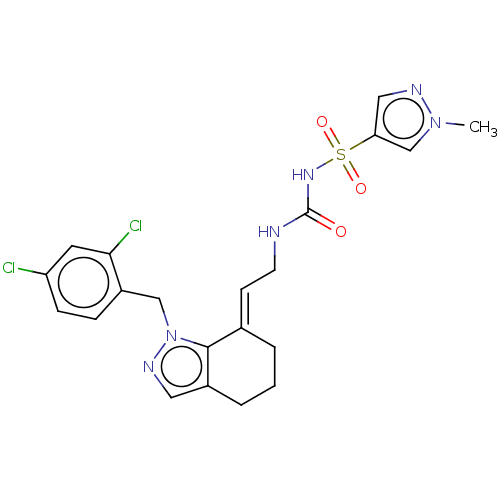
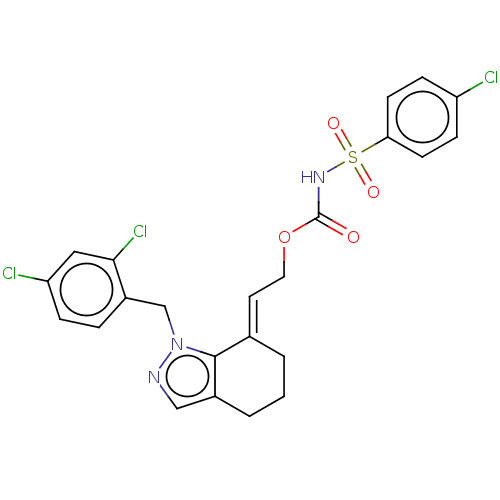
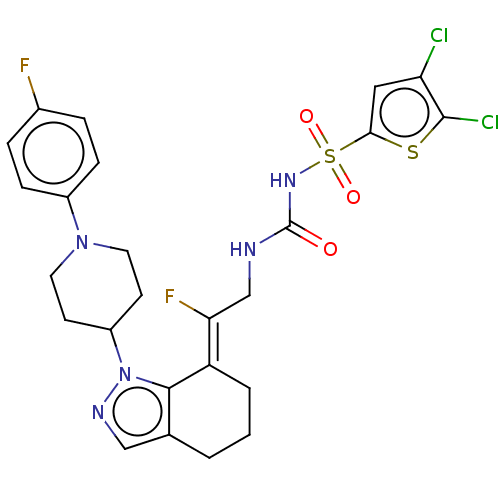
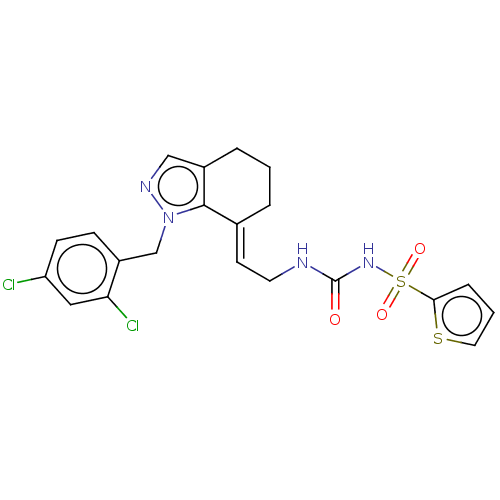
Report error Found 56 Enz. Inhib. hit(s) with all data for entry = 50014779
Affinity DataKi: 0.400nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 0.5nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataIC50: 1nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 1nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 2nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 3nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 3.60nMAssay Description:Binding affinity to rat EP3 receptor assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataKi: 4nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataIC50: 4nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataKi: 5nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataIC50: 5nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataIC50: 6nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 7nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataIC50: 8nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataKi: 9nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataIC50: 9nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataKi: 10nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataIC50: 11nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataKi: 11nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataKi: 13nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataIC50: 13nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataKi: 15nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataKi: 17nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataIC50: 18nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataKi: 18nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataIC50: 20nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataKi: 26nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataIC50: 27nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataKi: 28nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataIC50: 28nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataIC50: 29nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataIC50: 30nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataKi: 32nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataIC50: 34nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataIC50: 52nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataKi: 62nMAssay Description:Displacement of [3H]-PGE2 from human EP3 receptor assessed as inhibition constant incubated for 2 hrs by TopCount scintillation counting methodMore data for this Ligand-Target Pair
Affinity DataIC50: 69nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair
Affinity DataIC50: 92nMAssay Description:Antagonist activity at human EP3 receptor expressed in CHO-K1 cells assessed as reduction in sulprostone induced inhibition of forskolin stimulated c...More data for this Ligand-Target Pair