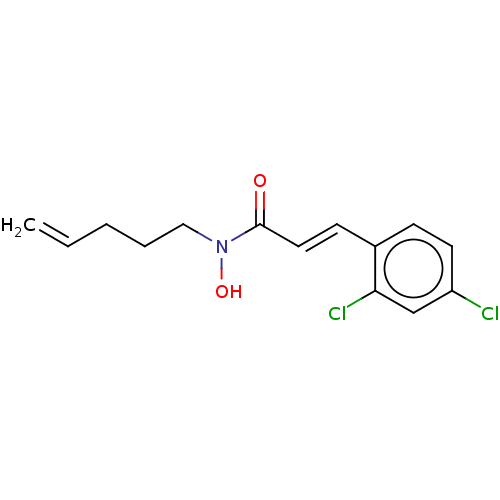
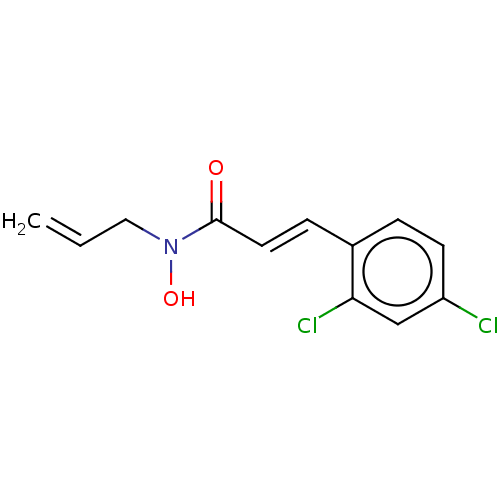
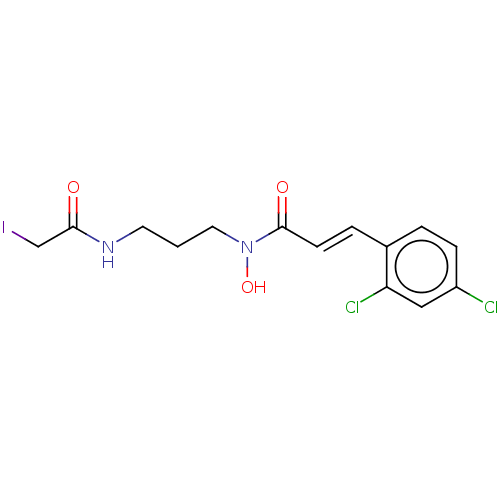
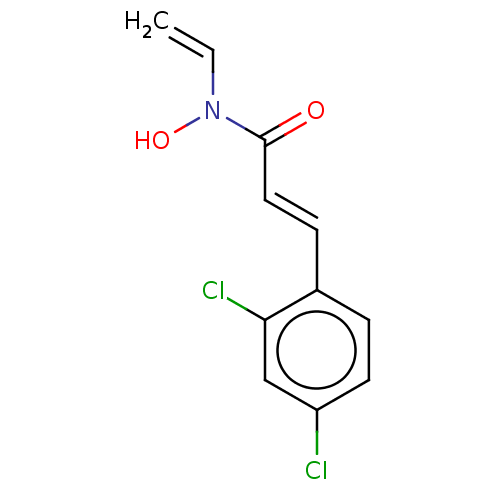
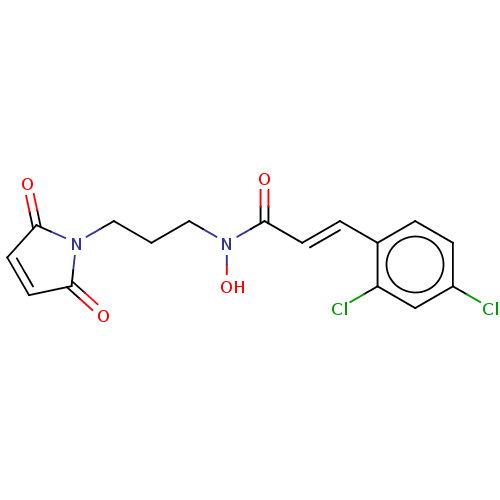
Report error Found 11 Enz. Inhib. hit(s) with all data for entry = 50015056
TargetBotulinum neurotoxin type A(Clostridium botulinum)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 300nMAssay Description:Inhibition of Clostridium botulinum BoNT/A light chain assessed as inhibition constant at 10 nM using SNAPtide flp6 as substrate measured after 30 mi...More data for this Ligand-Target Pair
TargetBotulinum neurotoxin type A(Clostridium botulinum)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 2.47E+3nMAssay Description:Inhibition of Clostridium botulinum BoNT/A light chain assessed as inhibition constant at 10 nM using SNAPtide flp6 as substrate measured after 30 mi...More data for this Ligand-Target Pair
TargetBotulinum neurotoxin type A(Clostridium botulinum)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 2.84E+3nMAssay Description:Inhibition of Clostridium botulinum BoNT/A light chain assessed as inhibition constant at 10 nM using SNAPtide flp6 as substrate measured after 30 mi...More data for this Ligand-Target Pair
TargetBotulinum neurotoxin type A(Clostridium botulinum)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 4.66E+3nMAssay Description:Inhibition of Clostridium botulinum BoNT/A light chain assessed as inhibition constant at 10 nM using SNAPtide flp6 as substrate measured after 30 mi...More data for this Ligand-Target Pair
TargetBotulinum neurotoxin type A(Clostridium botulinum)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 6.31E+3nMAssay Description:Inhibition of Clostridium botulinum BoNT/A light chain assessed as inhibition constant at 10 nM using SNAPtide flp6 as substrate measured after 30 mi...More data for this Ligand-Target Pair
TargetBotulinum neurotoxin type A(Clostridium botulinum)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: >8.00E+3nMAssay Description:Inhibition of Clostridium botulinum BoNT/A light chain assessed as inhibition constant at 10 nM using SNAPtide flp6 as substrate measured after 30 mi...More data for this Ligand-Target Pair
TargetBotulinum neurotoxin type A(Clostridium botulinum)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: >1.60E+4nMAssay Description:Inhibition of Clostridium botulinum BoNT/A light chain assessed as inhibition constant at 10 nM using SNAPtide flp6 as substrate measured after 30 mi...More data for this Ligand-Target Pair
TargetBotulinum neurotoxin type A(Clostridium botulinum)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 1.72E+4nMAssay Description:Inhibition of Clostridium botulinum BoNT/A light chain assessed as inhibition constant at 10 nM using SNAPtide flp6 as substrate measured after 30 mi...More data for this Ligand-Target Pair
TargetBotulinum neurotoxin type A(Clostridium botulinum)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 2.54E+4nMAssay Description:Inhibition of Clostridium botulinum BoNT/A light chain assessed as inhibition constant at 10 nM using SNAPtide flp6 as substrate measured after 30 mi...More data for this Ligand-Target Pair
TargetBotulinum neurotoxin type A(Clostridium botulinum)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: >3.20E+4nMAssay Description:Inhibition of Clostridium botulinum BoNT/A light chain assessed as inhibition constant at 10 nM using SNAPtide flp6 as substrate measured after 30 mi...More data for this Ligand-Target Pair
TargetBotulinum neurotoxin type A(Clostridium botulinum)
The Scripps Research Institute
Curated by ChEMBL
The Scripps Research Institute
Curated by ChEMBL
Affinity DataKi: 4.30E+4nMAssay Description:Inhibition of Clostridium botulinum BoNT/A light chain assessed as inhibition constant at 10 nM using SNAPtide flp6 as substrate measured after 30 mi...More data for this Ligand-Target Pair
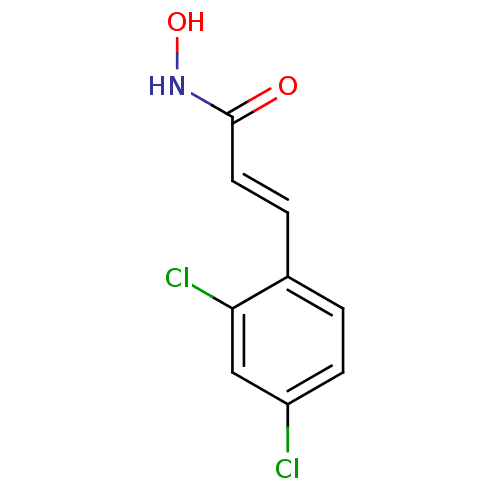
 3D Structure (crystal)
3D Structure (crystal)