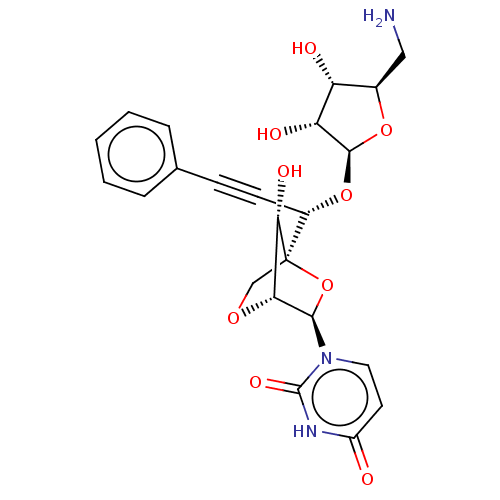
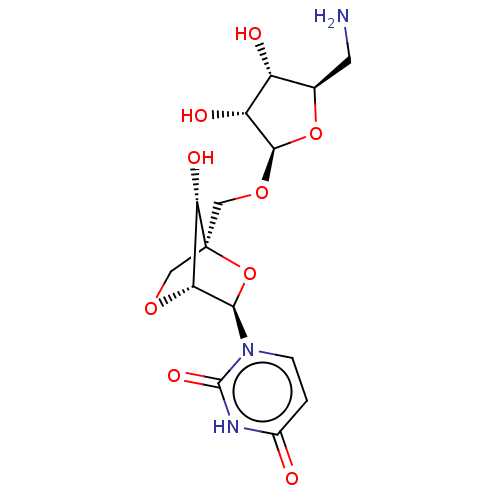
Report error Found 6 Enz. Inhib. hit(s) with all data for entry = 50017246
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Staphylococcus aureus)
Hokkaido University
Curated by ChEMBL
Hokkaido University
Curated by ChEMBL
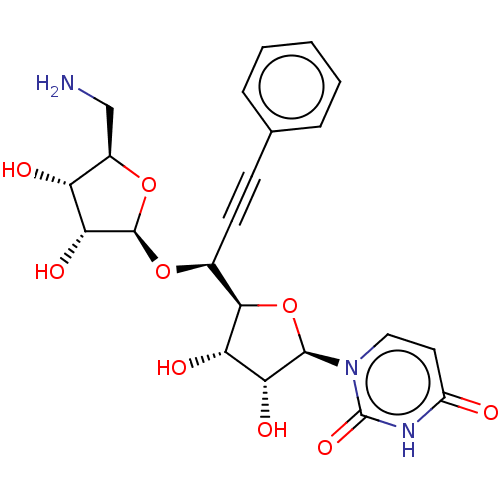
Affinity DataIC50: 2.30E+3nMAssay Description:Inhibition of Staphylococcus aureus MraY assessed as reduction of dansylated lipid I formation using UDP-MurNAcdansylpentapeptide as substrate incuba...More data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Staphylococcus aureus)
Hokkaido University
Curated by ChEMBL
Hokkaido University
Curated by ChEMBL
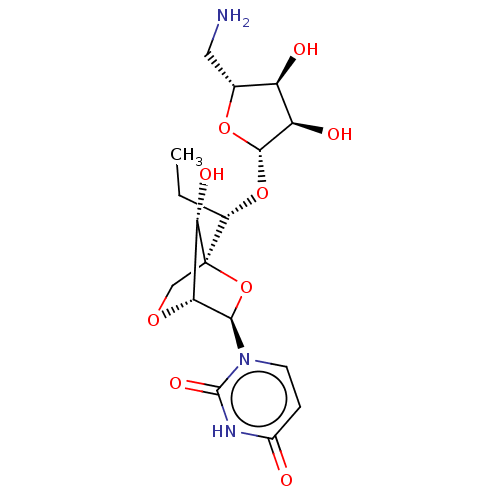
Affinity DataIC50: 4.50E+3nMAssay Description:Inhibition of Staphylococcus aureus MraY assessed as reduction of dansylated lipid I formation using UDP-MurNAcdansylpentapeptide as substrate incuba...More data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Staphylococcus aureus)
Hokkaido University
Curated by ChEMBL
Hokkaido University
Curated by ChEMBL
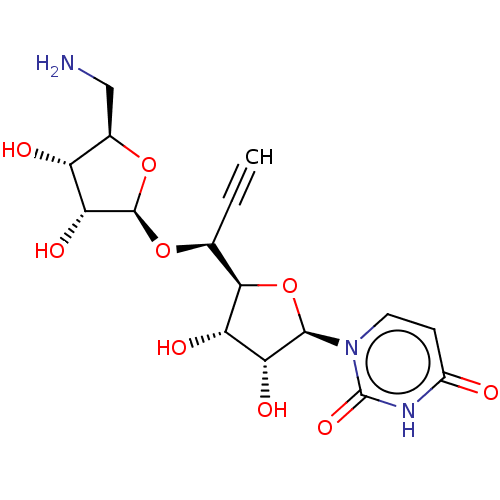
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of Staphylococcus aureus MraY assessed as reduction of dansylated lipid I formation using UDP-MurNAcdansylpentapeptide as substrate incuba...More data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Staphylococcus aureus)
Hokkaido University
Curated by ChEMBL
Hokkaido University
Curated by ChEMBL
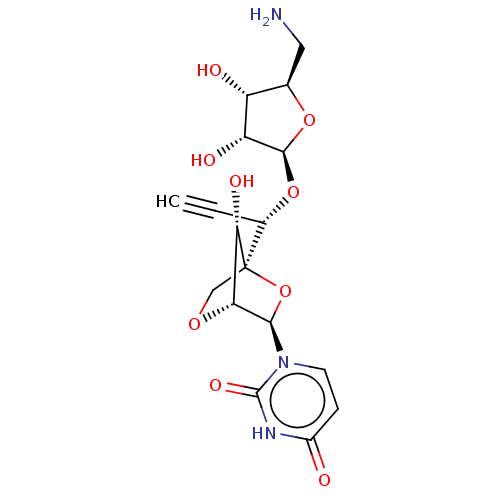
Affinity DataIC50: 2.40E+4nMAssay Description:Inhibition of Staphylococcus aureus MraY assessed as reduction of dansylated lipid I formation using UDP-MurNAcdansylpentapeptide as substrate incuba...More data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Staphylococcus aureus)
Hokkaido University
Curated by ChEMBL
Hokkaido University
Curated by ChEMBL
Affinity DataIC50: 3.70E+4nMAssay Description:Inhibition of Staphylococcus aureus MraY assessed as reduction of dansylated lipid I formation using UDP-MurNAcdansylpentapeptide as substrate incuba...More data for this Ligand-Target Pair
TargetPhospho-N-acetylmuramoyl-pentapeptide-transferase(Staphylococcus aureus)
Hokkaido University
Curated by ChEMBL
Hokkaido University
Curated by ChEMBL
Affinity DataIC50: 4.60E+4nMAssay Description:Inhibition of Staphylococcus aureus MraY assessed as reduction of dansylated lipid I formation using UDP-MurNAcdansylpentapeptide as substrate incuba...More data for this Ligand-Target Pair