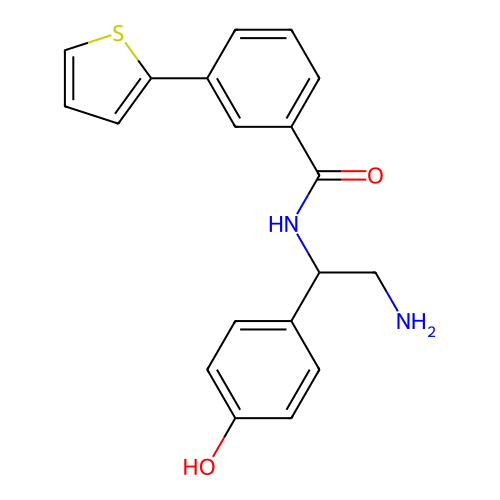
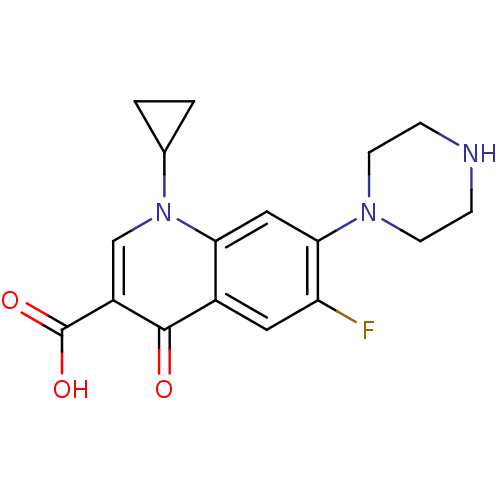
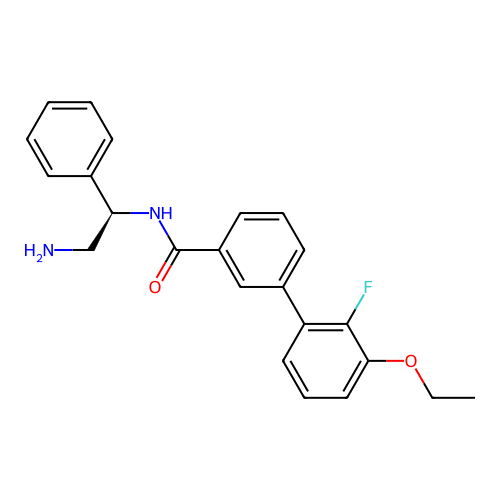
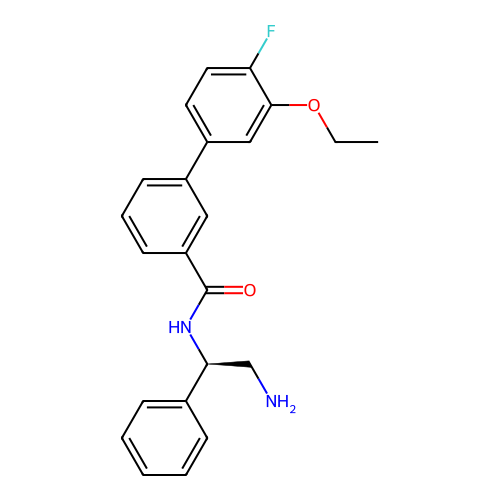
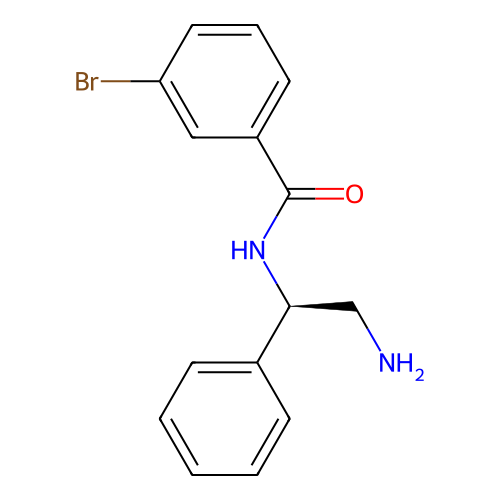
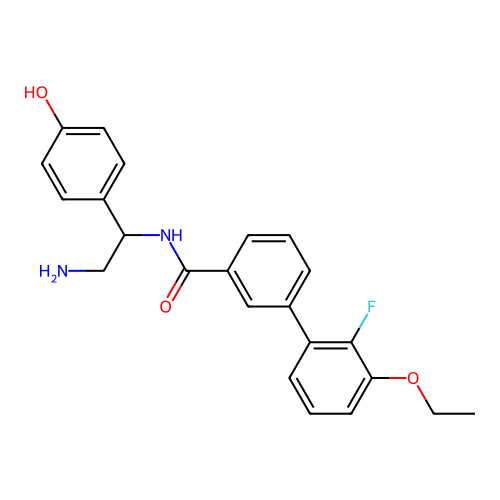
Report error Found 31 Enz. Inhib. hit(s) with all data for entry = 50015783
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
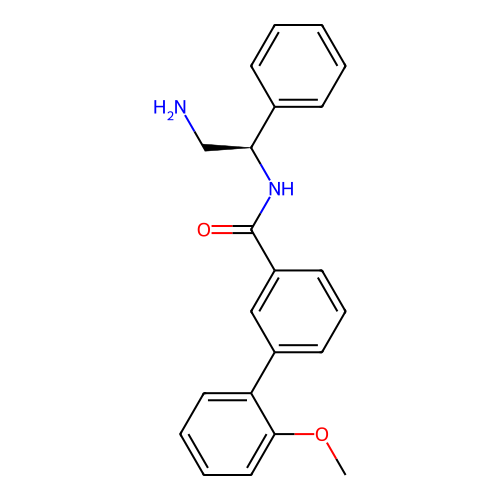
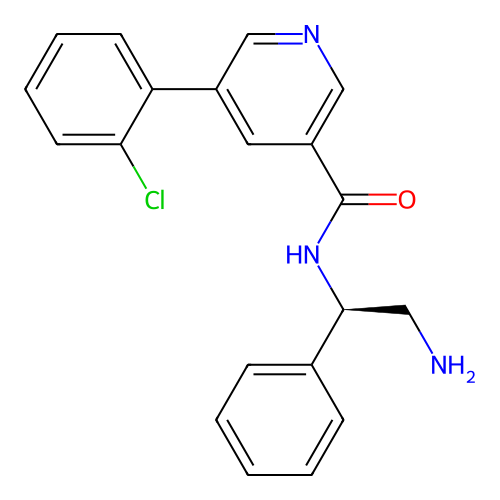
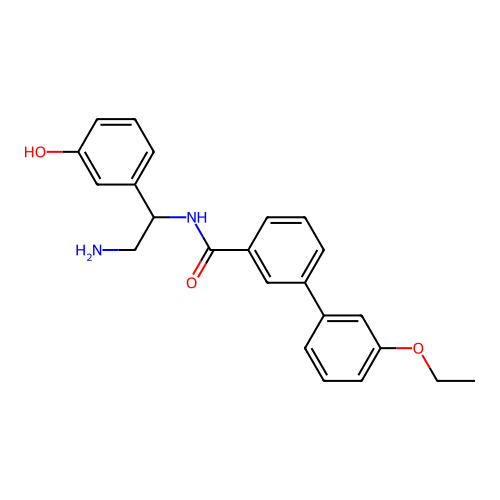
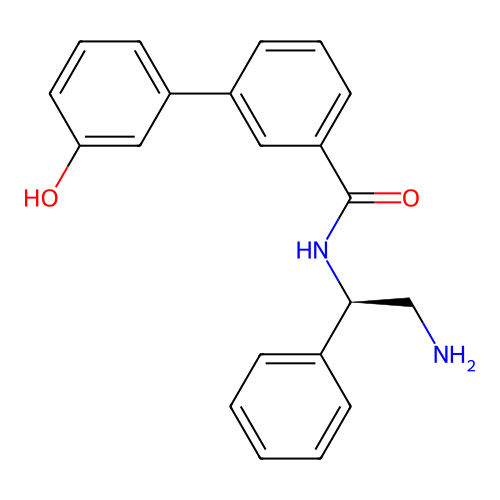
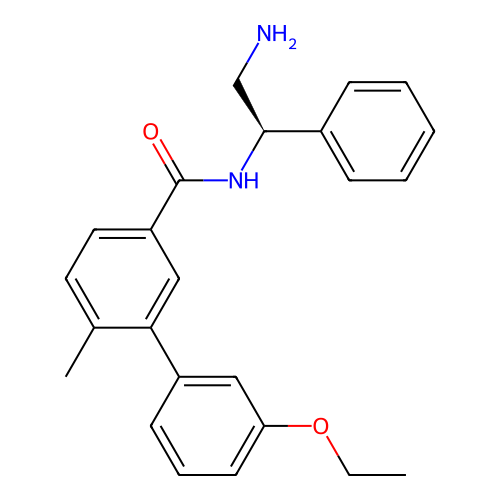
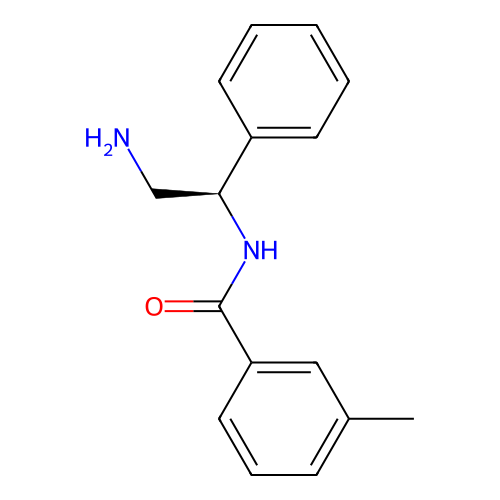
Affinity DataIC50: 300nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
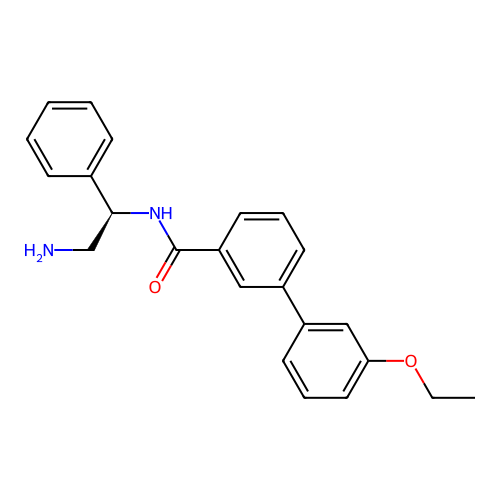
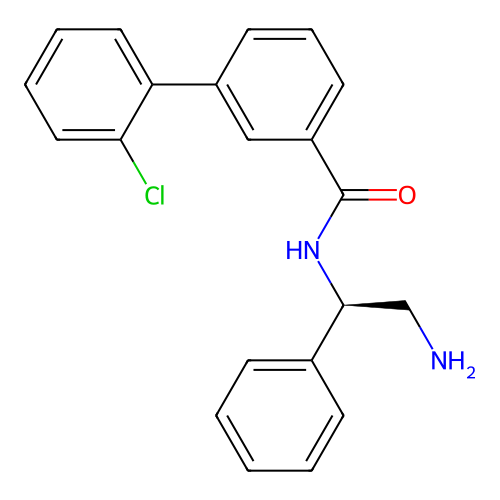
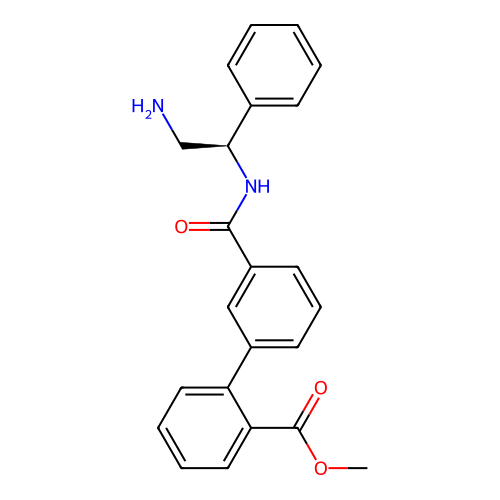
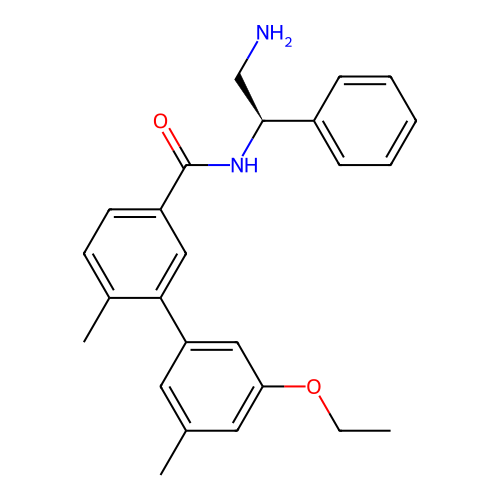
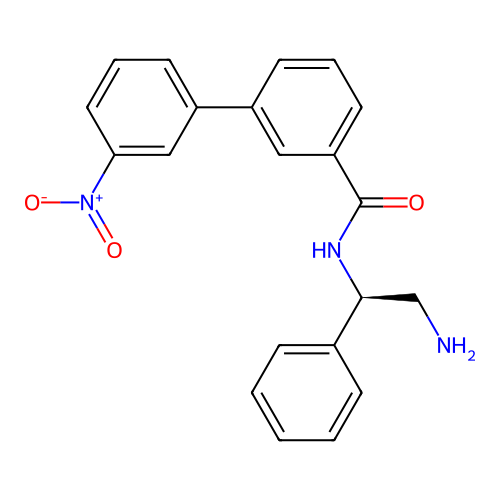
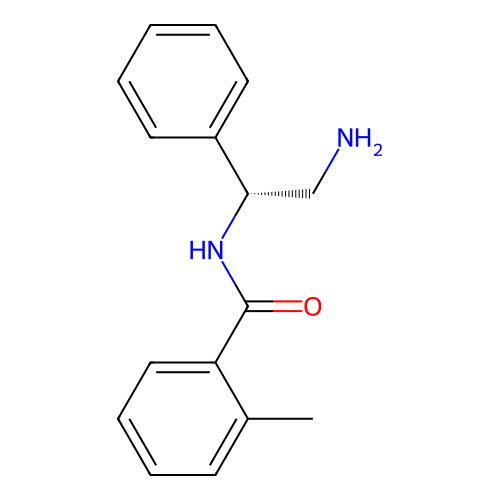
Affinity DataIC50: 600nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
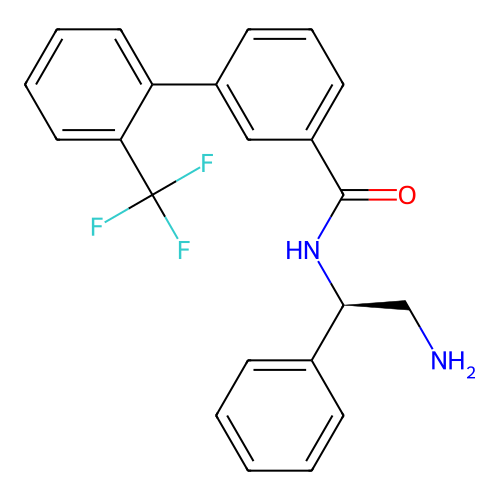
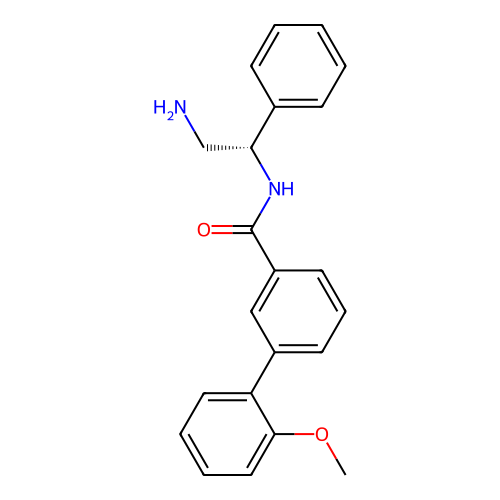
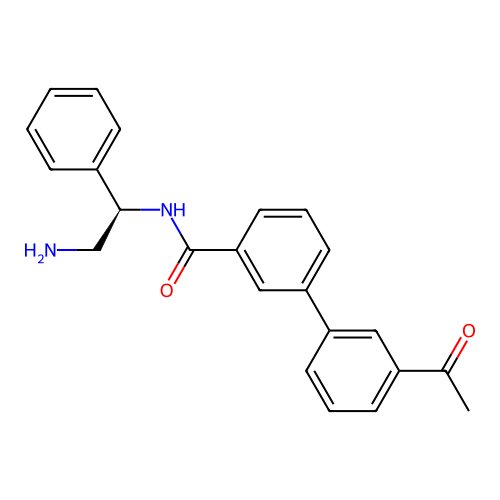
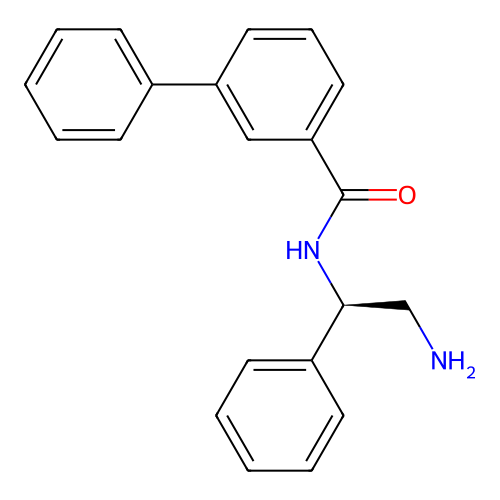
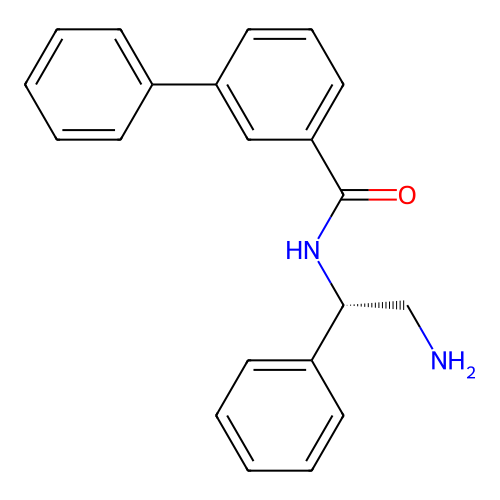
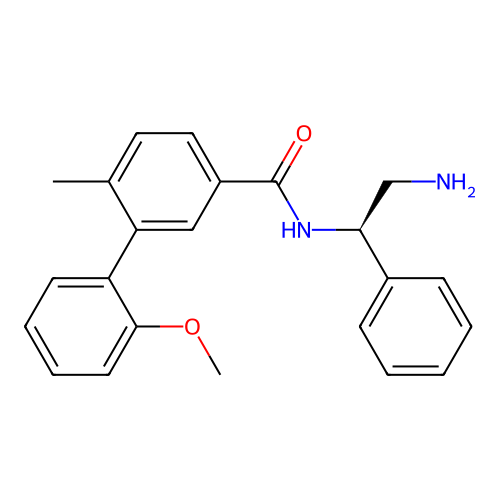
Affinity DataIC50: 1.20E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
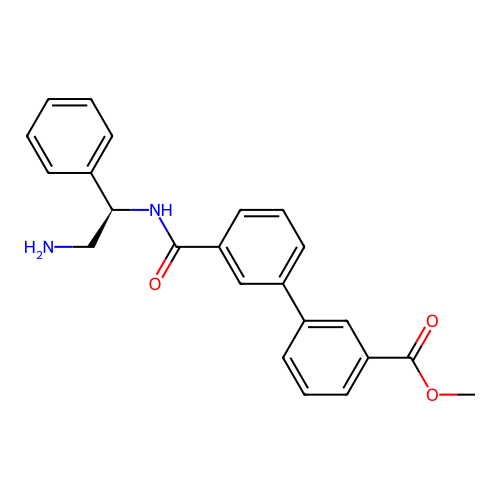
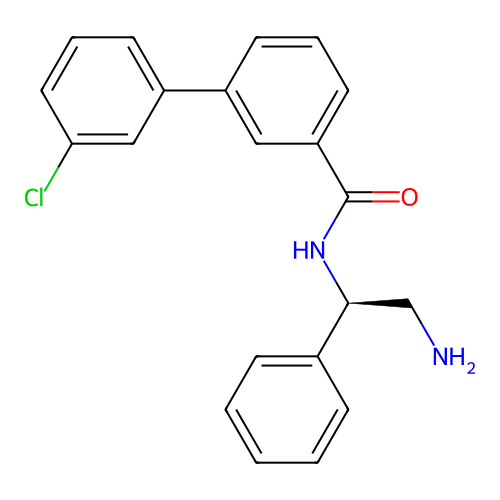
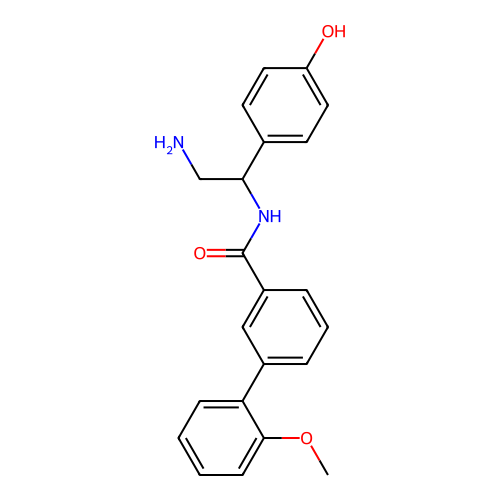
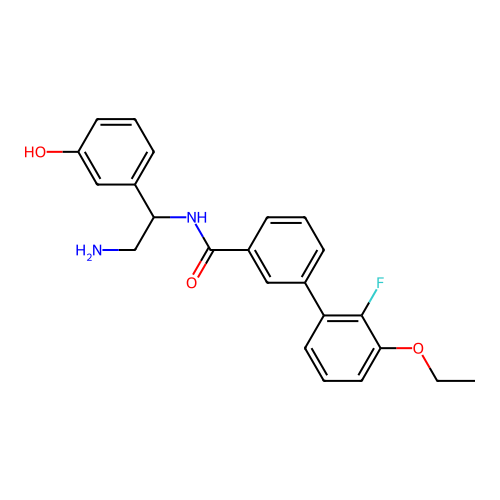
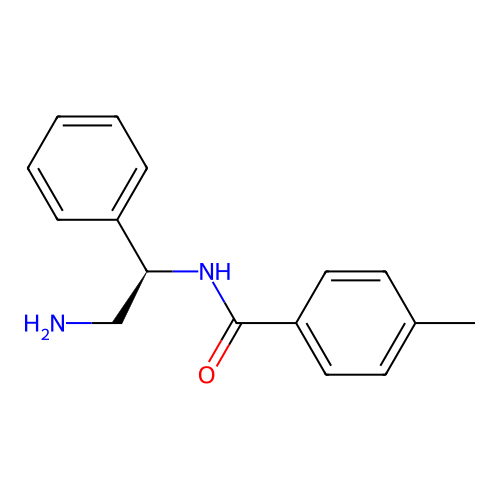
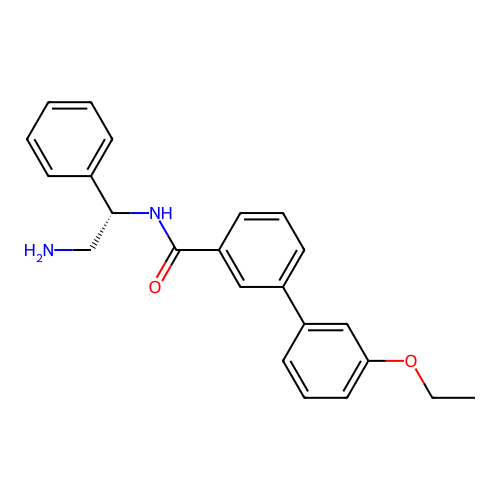
Affinity DataIC50: 1.70E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 2.10E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 2.40E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 3.50E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 3.50E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 3.60E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 3.90E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 4.10E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 4.10E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 4.10E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 4.20E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 4.50E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 4.60E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 5.60E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 6.00E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 6.20E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 6.30E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 7.30E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 7.60E+4nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 1.00E+5nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 1.53E+5nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 1.80E+5nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair
TargetDNA gyrase A/A/B/B tetramer(Escherichia coli (strain K12))
University of Leeds
Curated by ChEMBL
University of Leeds
Curated by ChEMBL
Affinity DataIC50: 2.00E+5nMAssay Description:Inhibition of Escherichia coli DNA gyrase supercoiling activity using pBR322 as substrate incubated for 30 mins measured by gel-based assayMore data for this Ligand-Target Pair