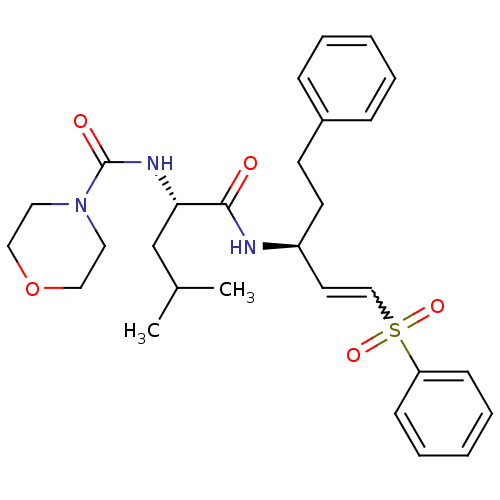
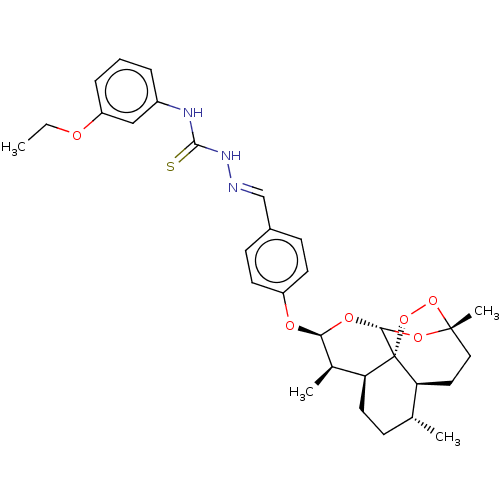
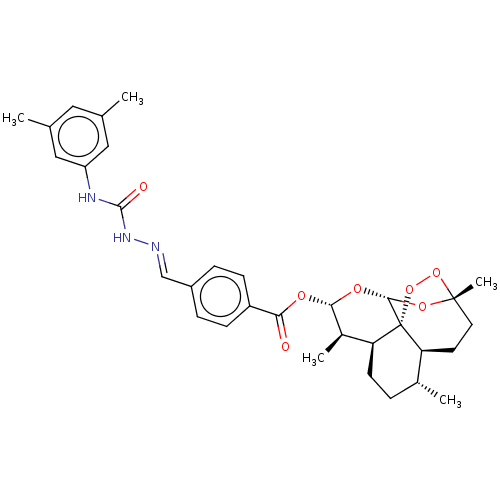
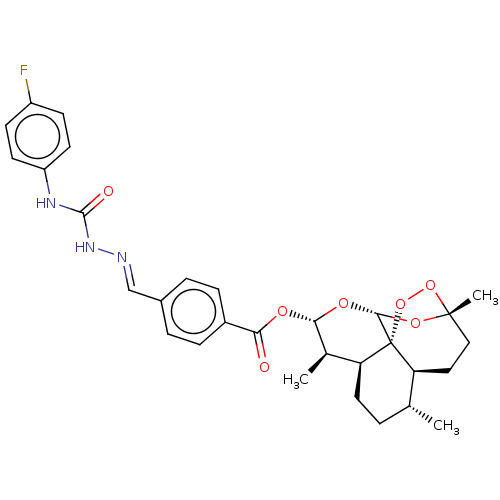
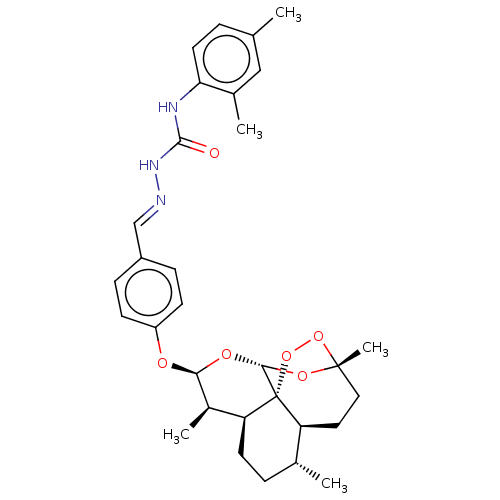
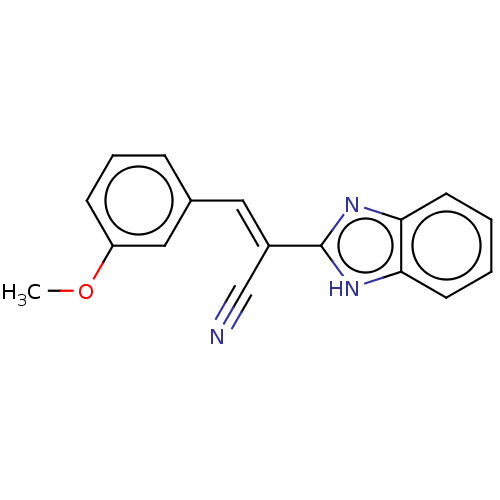
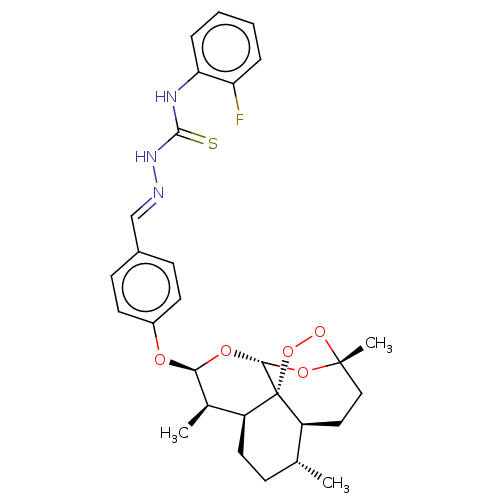
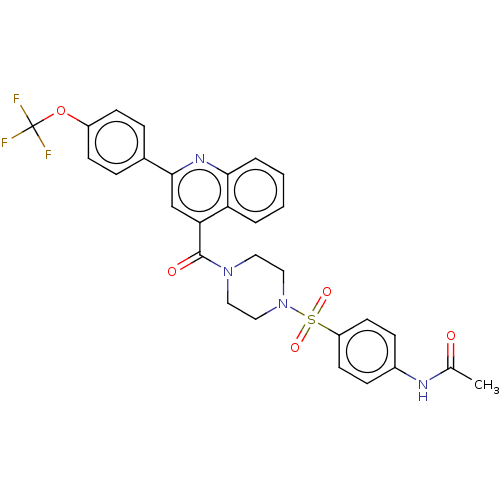
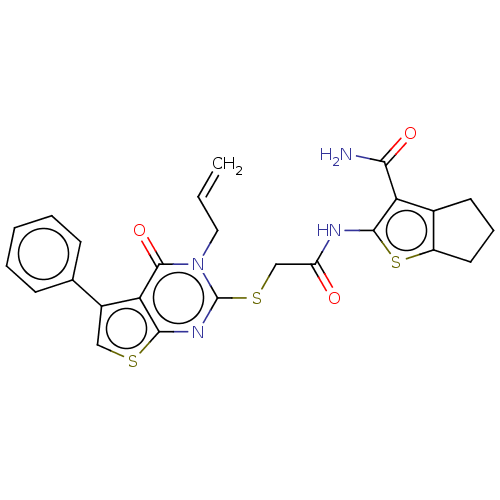
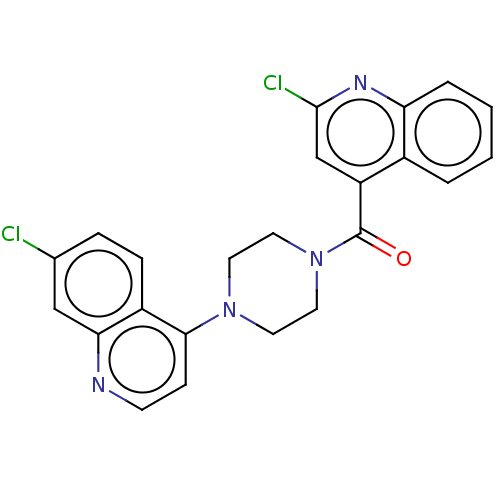
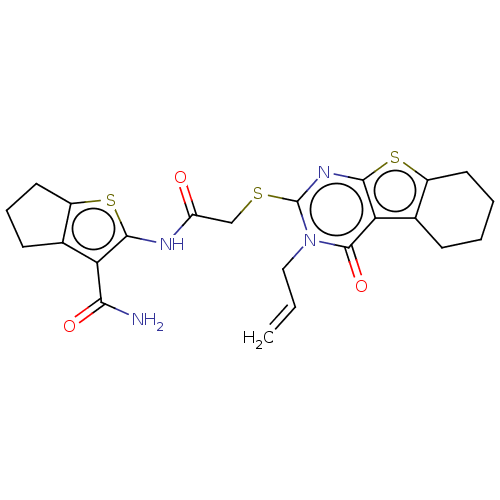
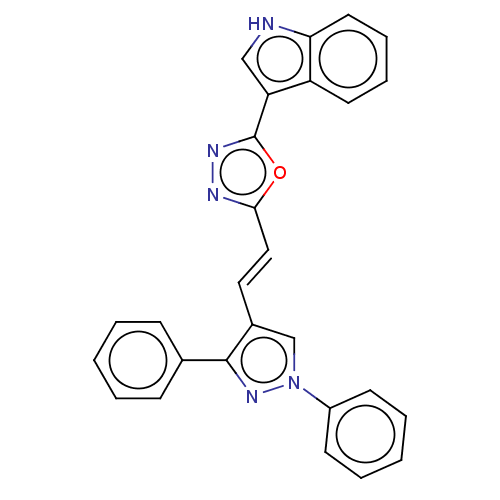
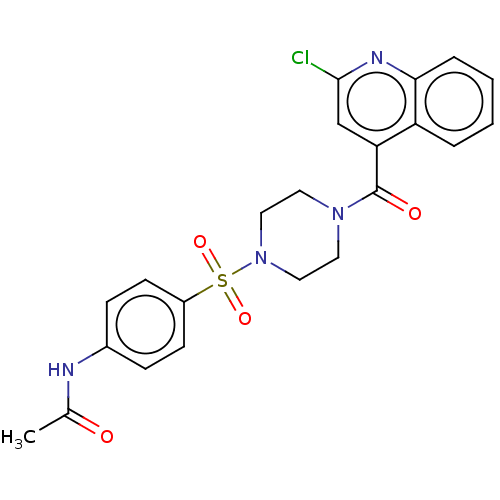
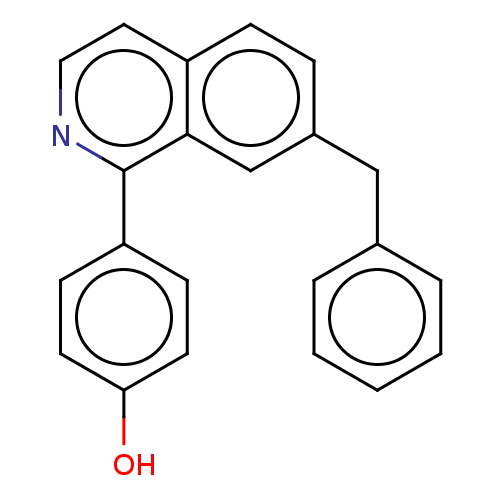
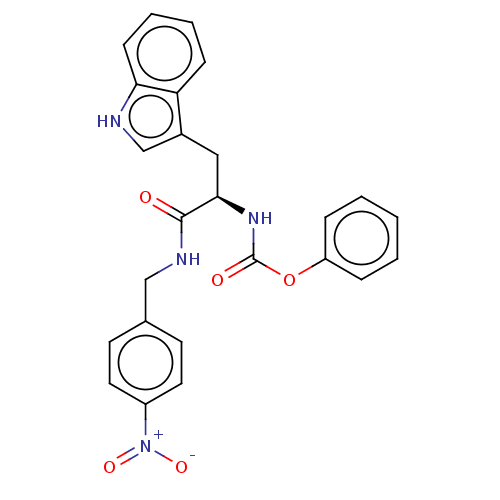
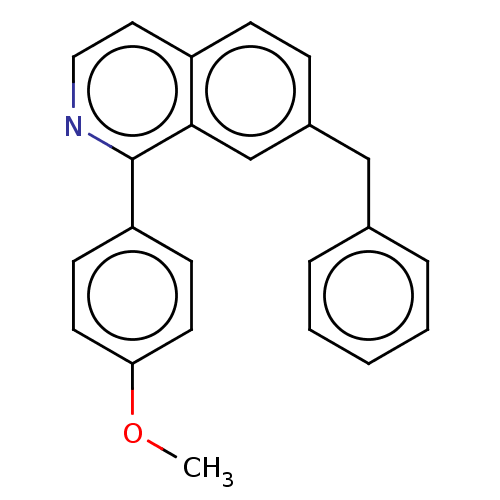
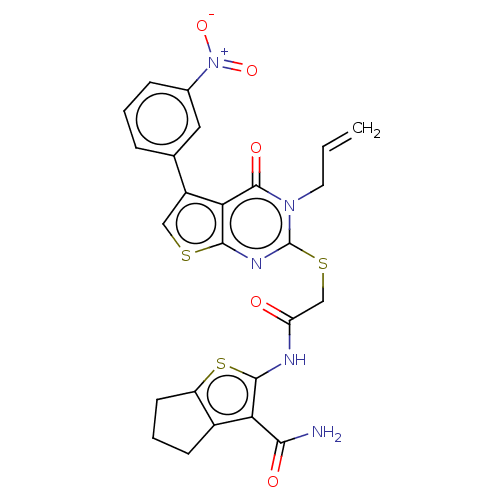
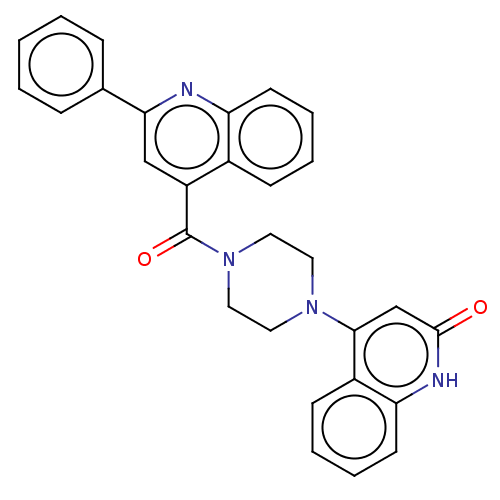
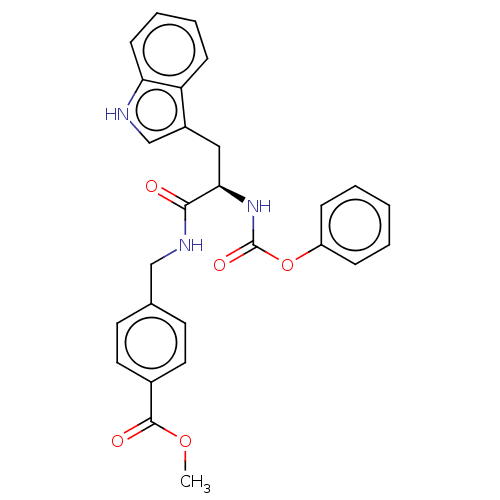
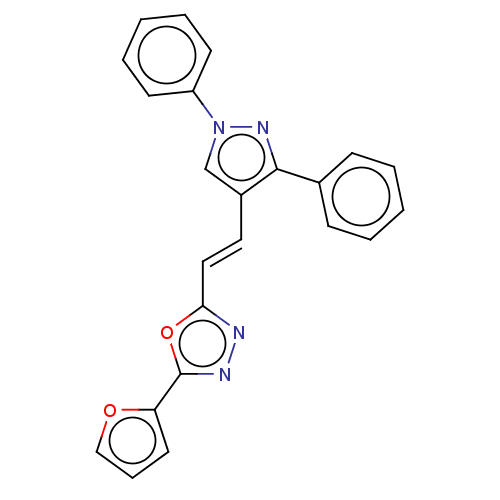
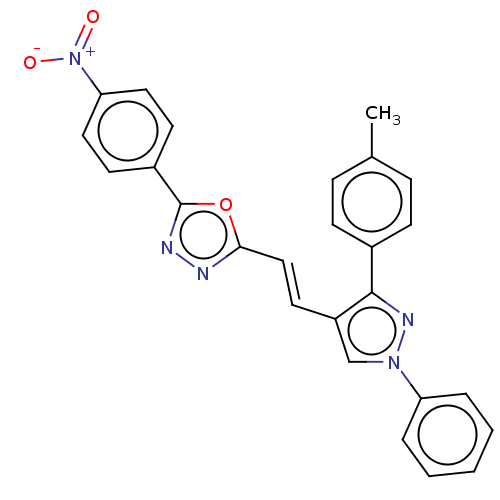
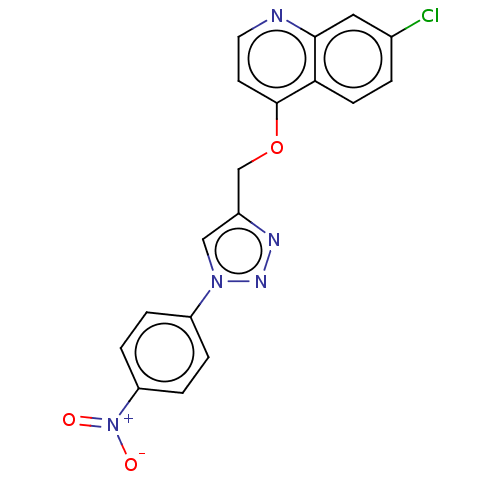
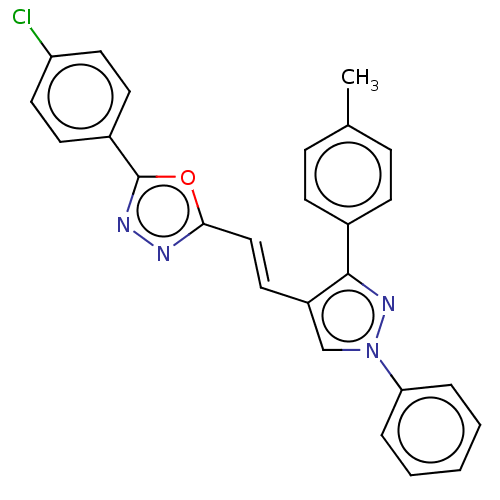
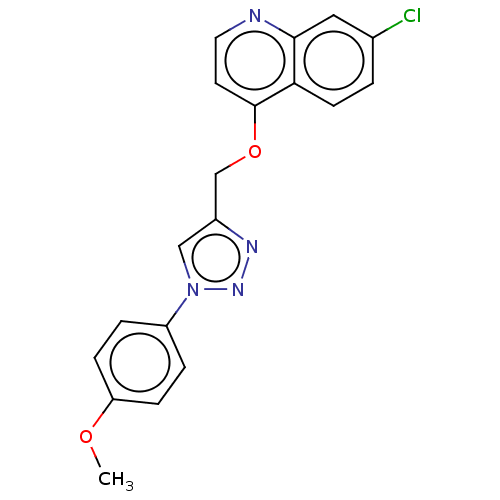
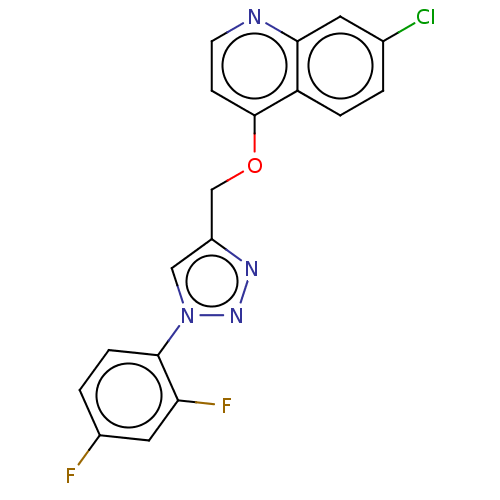
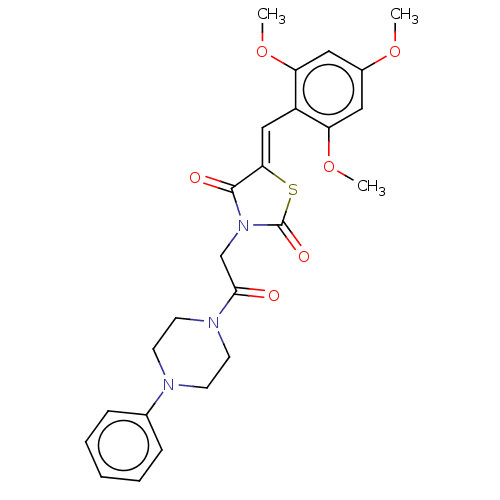
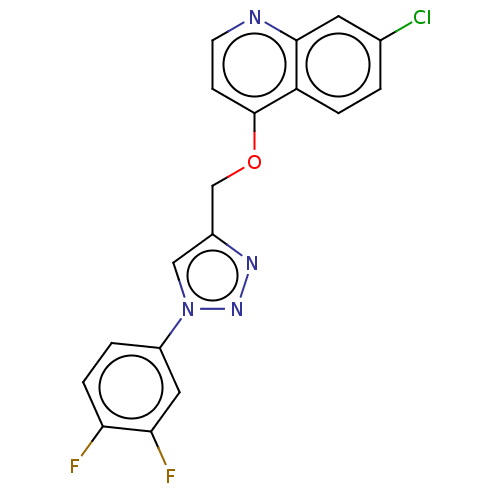
Report error Found 58 Enz. Inhib. hit(s) with all data for entry = 50019345
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
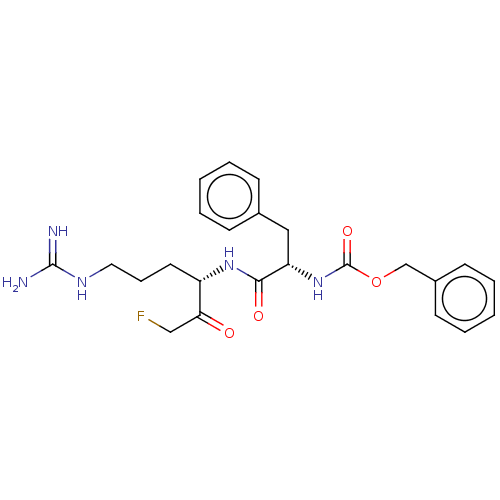
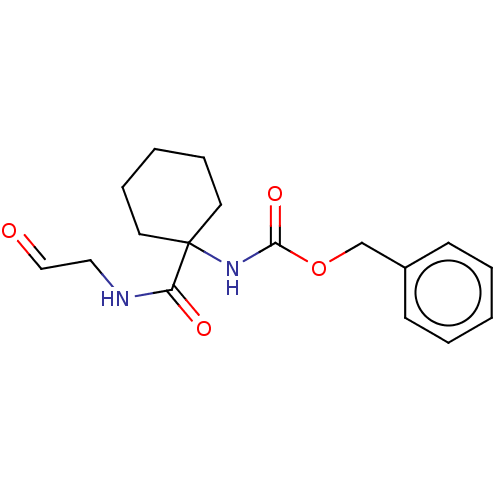
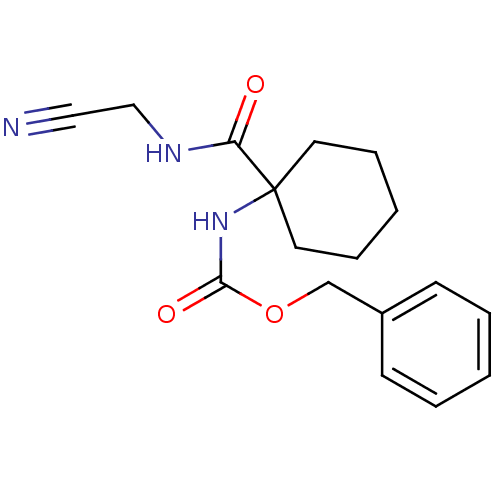
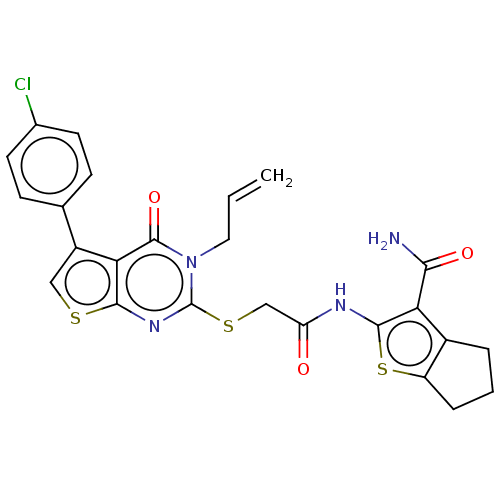
Affinity DataIC50: 0.360nMAssay Description:Inhibition of Plasmodium falciparum trophozite extracts FP-2More data for this Ligand-Target Pair
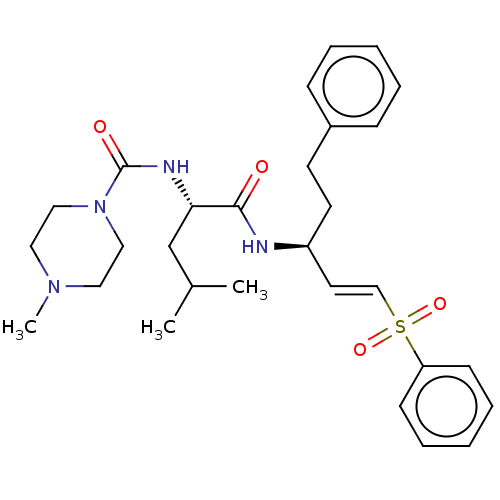
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
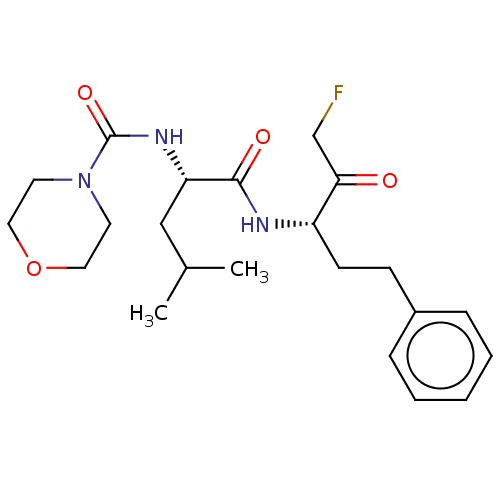
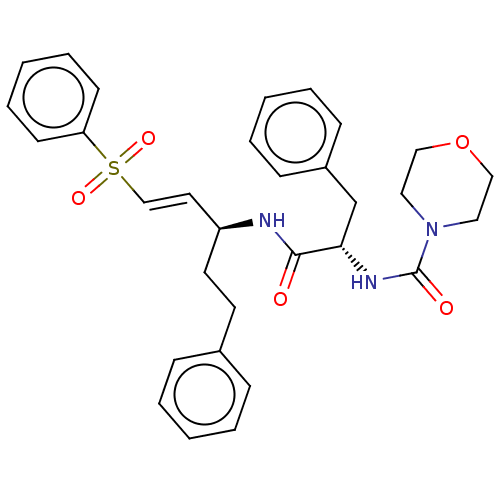
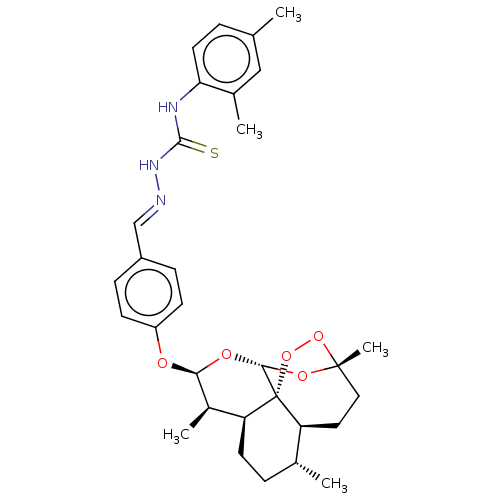
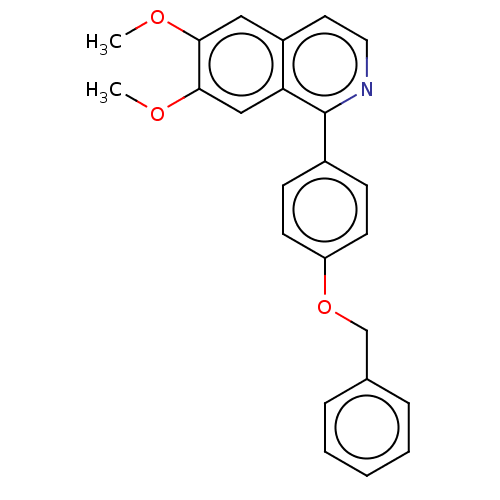
Affinity DataIC50: 0.420nMAssay Description:Inhibition of Plasmodium falciparum trophozite extracts FP-2More data for this Ligand-Target Pair
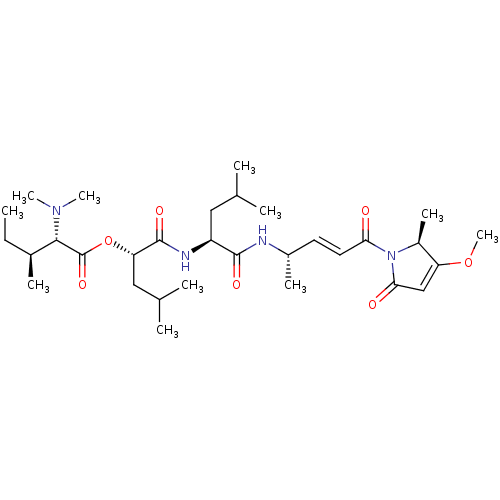
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
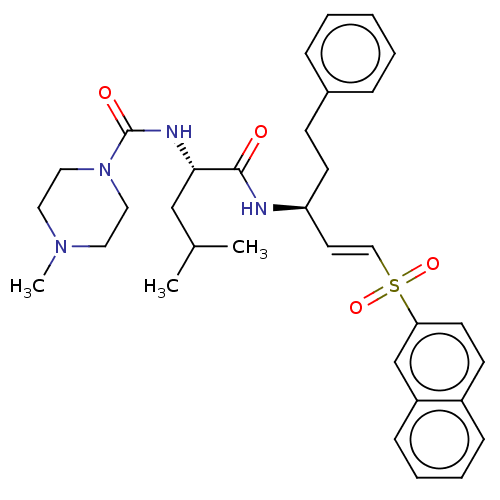
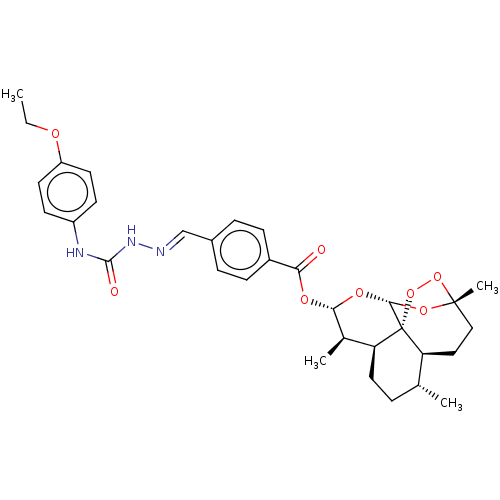
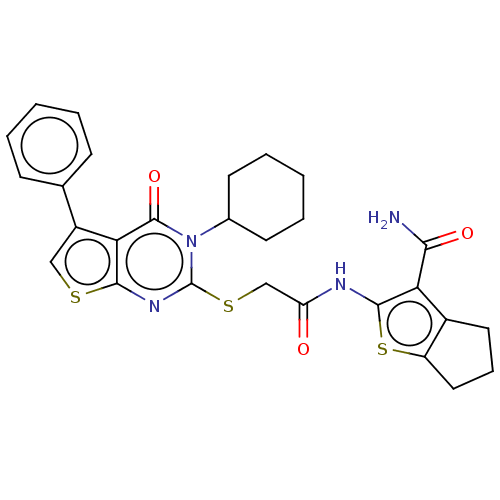
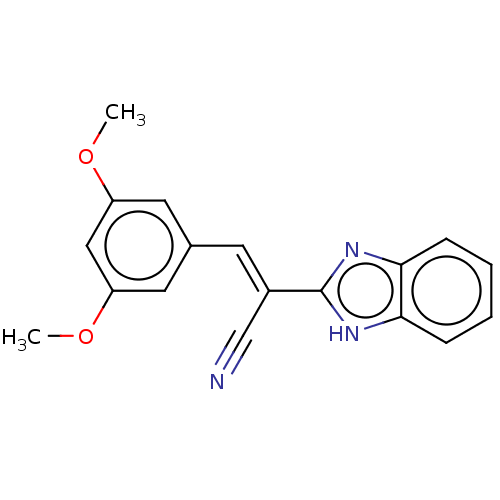
Affinity DataIC50: 2nMAssay Description:Inhibition of Plasmodium falciparum Itg2 trophozite extracts FP-2 using Z-phe-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by sub...More data for this Ligand-Target Pair
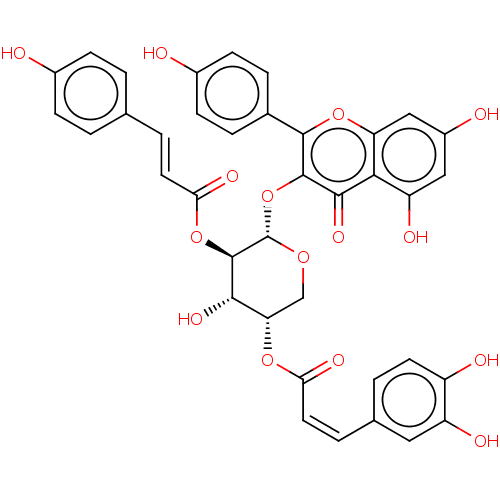
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
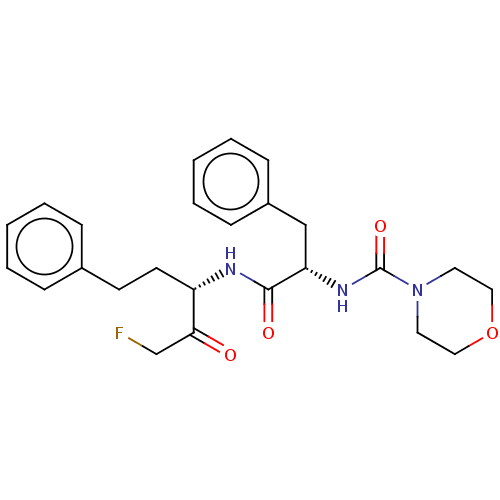
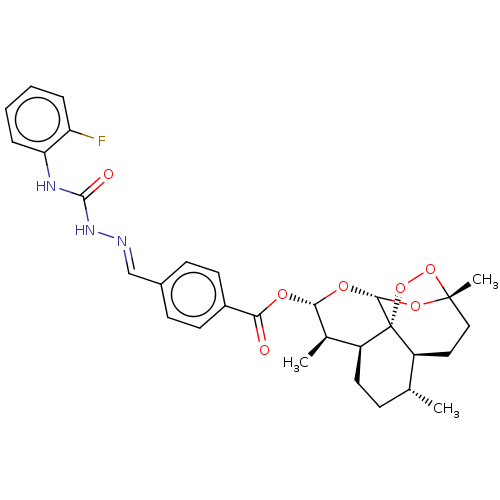
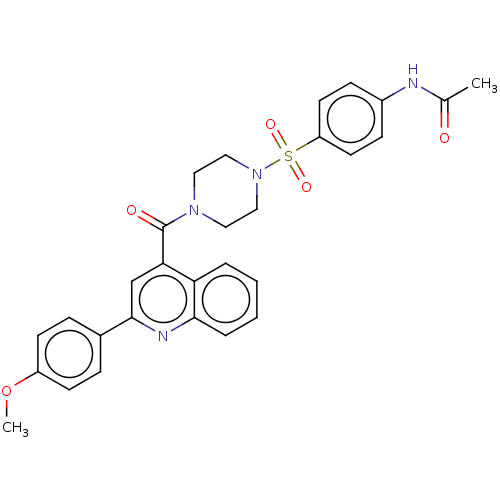
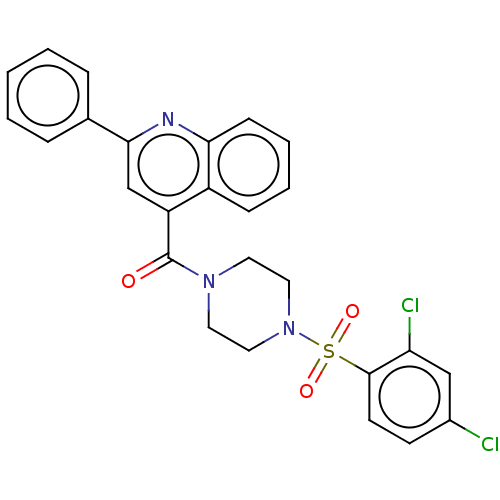
Affinity DataIC50: 3nMAssay Description:Inhibition of Plasmodium falciparum trophozite extracts FP-2More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 3nMAssay Description:Inhibition of Plasmodium falciparum Itg2 trophozite extracts FP-2 using Z-phe-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by sub...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 5nMAssay Description:Inhibition of Plasmodium falciparum Itg2 trophozite extracts FP-2 using Z-phe-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by sub...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 8.5nMAssay Description:Inhibition of Plasmodium falciparum D10 trophozite lysates FP-2 pretreated with compound for 1 hr followed by labeling with Cy5-DCG04 for 1 hr by flu...More data for this Ligand-Target Pair
TargetCysteine proteinase falcipain 3(Plasmodium falciparum (isolate 3D7))
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 22nMAssay Description:Inhibition of Plasmodium falciparum D10 trophozite lysates FP-3 pretreated with compound for 1 hr followed by labeling with Cy5-DCG04 for 1 hr by flu...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataKd: 34nMAssay Description:Binding affinity to Plasmodium falciparum FP-2 by surface plasmon resonance analysisMore data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataKi: 65nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Cbz-Leu-Arg-AMC as substrate and measured for 5 to 10 mins by fluorescence spectrophotometric analysisMore data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 80nMAssay Description:Inhibition of Plasmodium falciparum Itg2 trophozite extracts FP-2 using Z-phe-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by sub...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 290nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Z-Leu-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by substrate addition and measu...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 440nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Z-Leu-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by substrate addition and measu...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 470nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Z-Leu-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by substrate addition and measu...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 520nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Z-Leu-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by substrate addition and measu...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 570nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Z-Leu-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by substrate addition and measu...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 650nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Z-Leu-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by substrate addition and measu...More data for this Ligand-Target Pair
Affinity DataKi: 820nMAssay Description:Inhibition of human Cathepsin L using Cbz-Phe-Arg-AMC as substrate by fluorescence spectrophotometric analysisMore data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 1.03E+3nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Z-Leu-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by substrate addition and measu...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataKi: 1.20E+3nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Cbz-Leu-Arg-AMC as substrate and measured for 5 to 10 mins by fluorescence spectrophotometric analysisMore data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 1.35E+3nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Z-Leu-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by substrate addition and measu...More data for this Ligand-Target Pair
Affinity DataKi: 1.35E+3nMAssay Description:Inhibition of human Cathepsin B using Cbz-Phe-Arg-AMC as substrate by fluorescence spectrophotometric analysisMore data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 1.46E+3nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Z-Leu-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by substrate addition and measu...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 2.14E+3nMAssay Description:Inhibition of Plasmodium falciparum FP2 using ZFR-AMC as substrate preincubated with enzyme for 10 mins followed by substrate addition and measured f...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 2.21E+3nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Z-Phe-Arg-AMC as substrate pretreated with enzyme for 10 mins followed by susbtrate addition and measu...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 2.51E+3nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Z-Leu-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by substrate addition and measu...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 2.64E+3nMAssay Description:Inhibition of Plasmodium falciparum FP2 using ZFR-AMC as substrate preincubated with enzyme for 10 mins followed by substrate addition and measured f...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 2.81E+3nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Z-Leu-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by substrate addition and measu...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 2.95E+3nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Z-Leu-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by substrate addition and measu...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 3.00E+3nMAssay Description:Inhibition of Plasmodium falciparum Itg2 trophozite extracts FP-2 using Z-phe-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by sub...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 3.61E+3nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Z-Phe-Arg-AMC as substrate pretreated with enzyme for 10 mins followed by susbtrate addition and measu...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 4.20E+3nMAssay Description:Inhibition of Plasmodium falciparum FP2 using ZFR-AMC as substrate preincubated with enzyme for 10 mins followed by substrate addition and measured f...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 4.78E+3nMAssay Description:Inhibition of Plasmodium falciparum FP2 using ZFR-AMC as substrate preincubated with enzyme for 10 mins followed by substrate addition and measured f...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 5.77E+3nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Z-Leu-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by substrate addition and measu...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 7.00E+3nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Cbz-Phe-Arg-AMC as substrate pretreated with enzyme for 10 mins followed by substrate addition and mea...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 7.37E+3nMAssay Description:Inhibition of Plasmodium falciparum FP2 using ZFR-AMC as substrate preincubated with enzyme for 10 mins followed by substrate addition and measured f...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 8.00E+3nMAssay Description:Inhibition of Plasmodium falciparum Itg2 trophozite extracts FP-2 using Z-phe-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by sub...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Z-Leu-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by substrate addition and measu...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 1.00E+4nMAssay Description:Inhibition of Plasmodium falciparum Itg2 trophozite extracts FP-2 using Z-phe-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by sub...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 1.18E+4nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Z-Leu-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by substrate addition and measu...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 1.30E+4nMAssay Description:Inhibition of Plasmodium falciparum FP2 using ZFR-AMC as substrate preincubated with enzyme for 10 mins followed by substrate addition and measured f...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 1.33E+4nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Z-Leu-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by substrate addition and measu...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 1.40E+4nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Cbz-Phe-Arg-AMC as substrate pretreated with enzyme for 10 mins followed by substrate addition and mea...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 1.60E+4nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Cbz-Phe-Arg-AMC as substrate pretreated with enzyme for 10 mins followed by substrate addition and mea...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 1.62E+4nMAssay Description:Inhibition of Plasmodium falciparum FP2 using ZFR-AMC as substrate preincubated with enzyme for 10 mins followed by substrate addition and measured f...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 2.00E+4nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Cbz-Phe-Arg-AMC as substrate pretreated with enzyme for 10 mins followed by substrate addition and mea...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 2.54E+4nMAssay Description:Inhibition of Plasmodium falciparum FP2 using ZFR-AMC as substrate preincubated with enzyme for 10 mins followed by substrate addition and measured f...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 2.54E+4nMAssay Description:Inhibition of Plasmodium falciparum FP2 using ZFR-AMC as substrate preincubated with enzyme for 10 mins followed by substrate addition and measured f...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 2.55E+4nMAssay Description:Inhibition of Plasmodium falciparum FP-2 using Z-Leu-Arg-AMC as substrate pretreated with enzyme for 30 mins followed by substrate addition and measu...More data for this Ligand-Target Pair
TargetFalcipain 2(malaria parasite P. falciparum)
University of Petroleum and Energy Studies
Curated by ChEMBL
University of Petroleum and Energy Studies
Curated by ChEMBL
Affinity DataIC50: 2.56E+4nMAssay Description:Inhibition of Plasmodium falciparum FP2 using ZFR-AMC as substrate preincubated with enzyme for 10 mins followed by substrate addition and measured f...More data for this Ligand-Target Pair