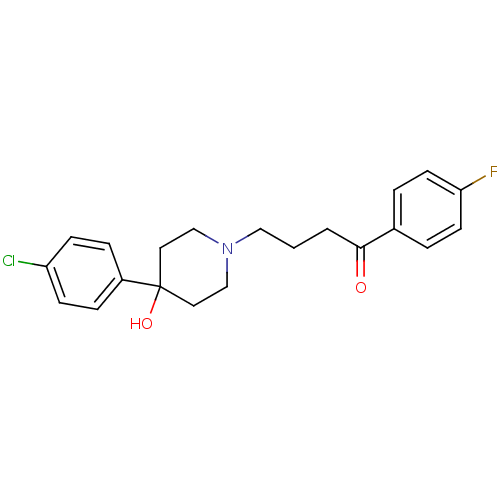
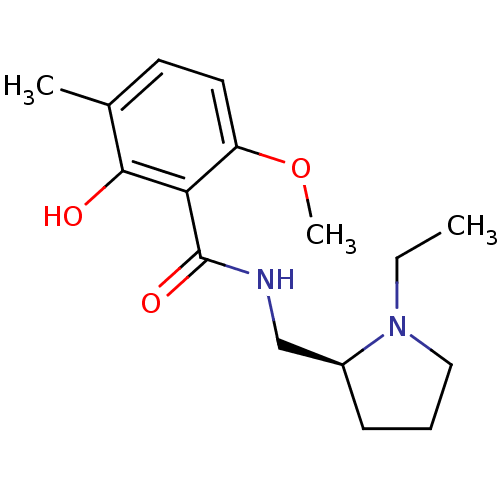
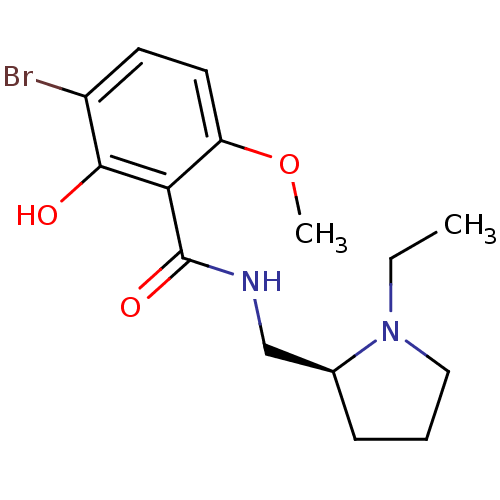
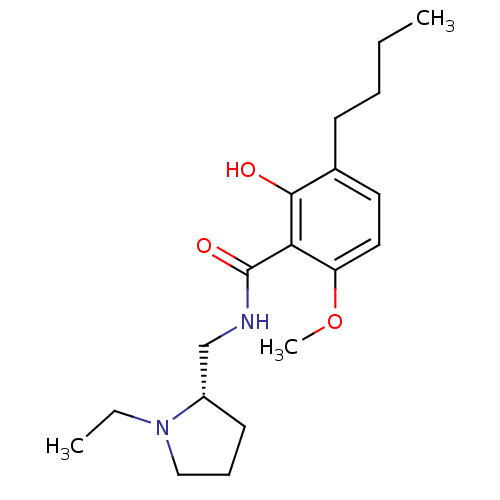
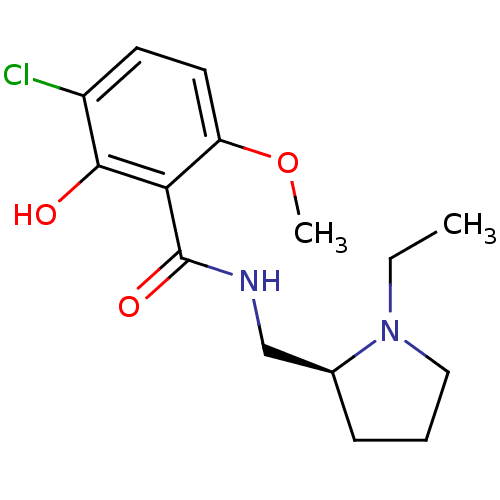
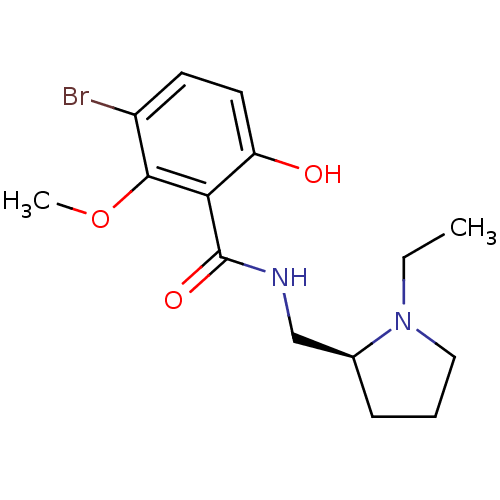
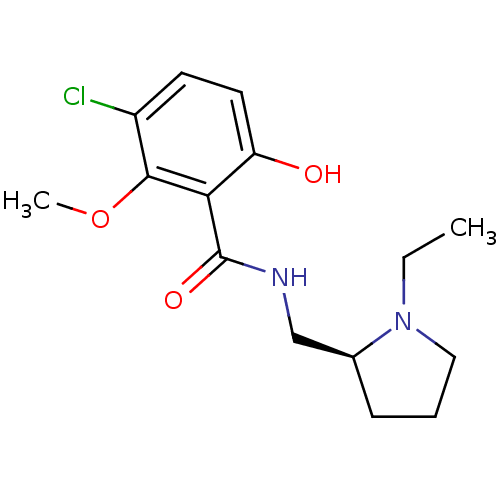
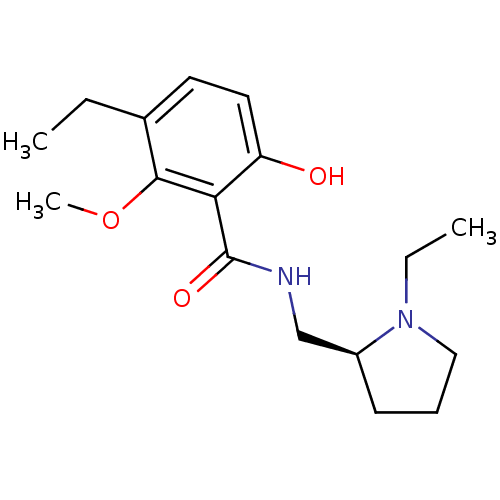
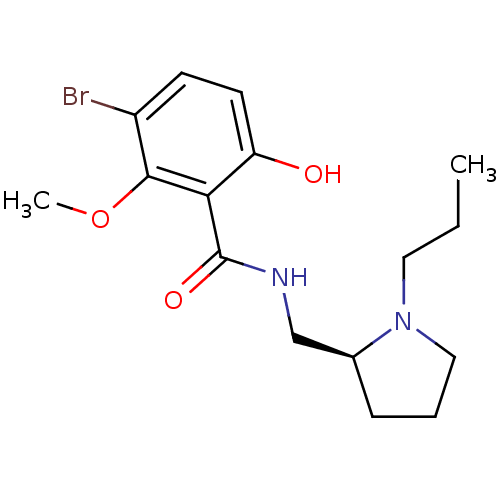
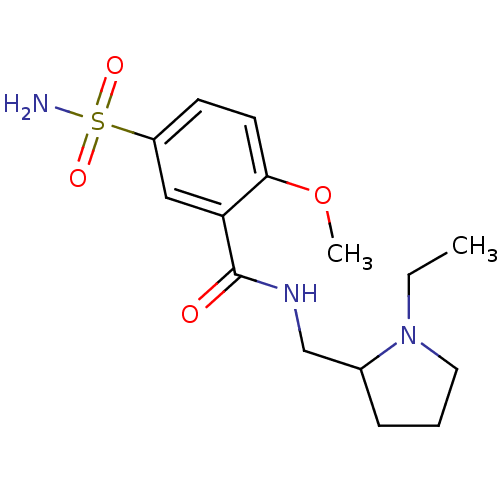
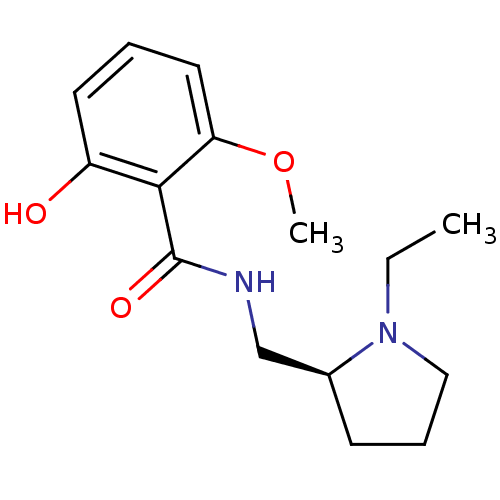
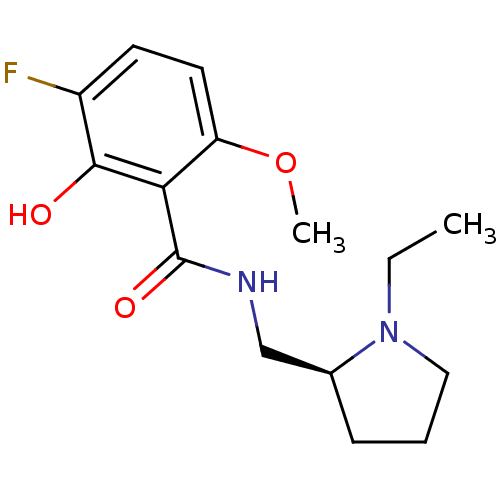
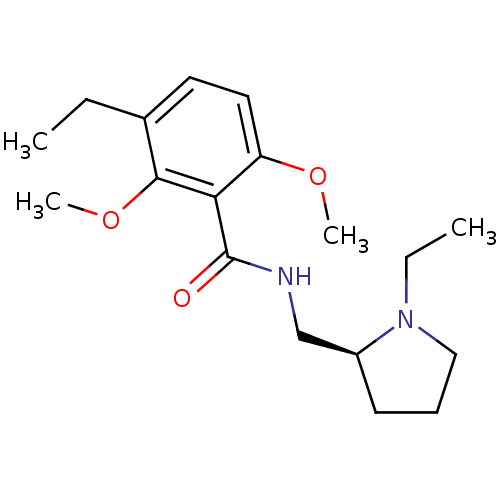
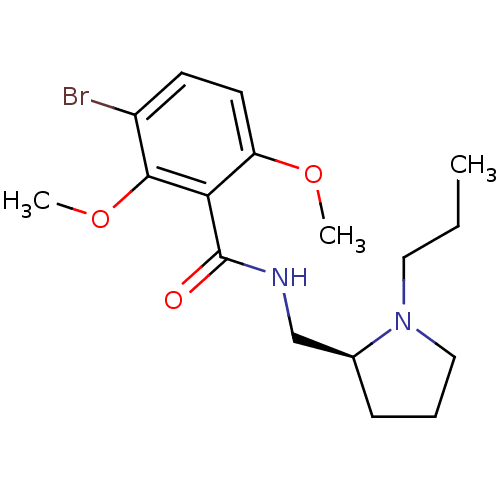
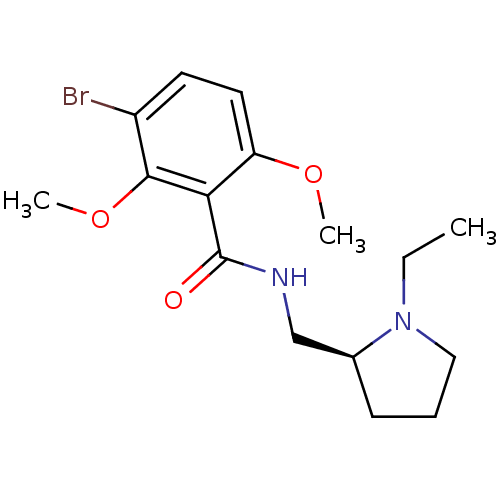
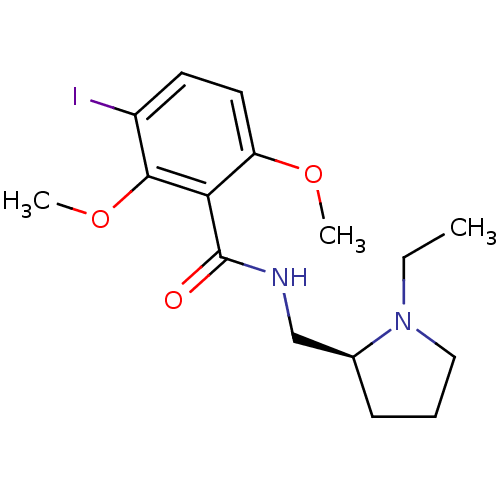
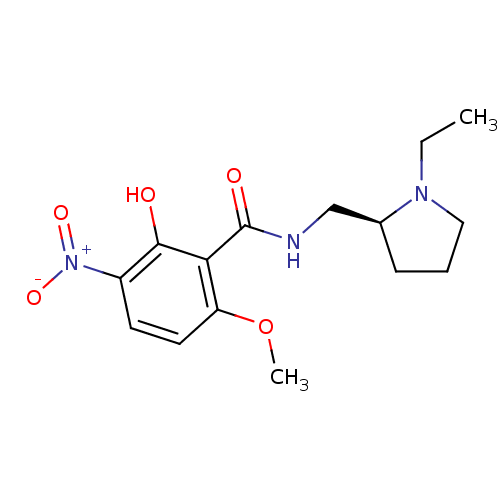
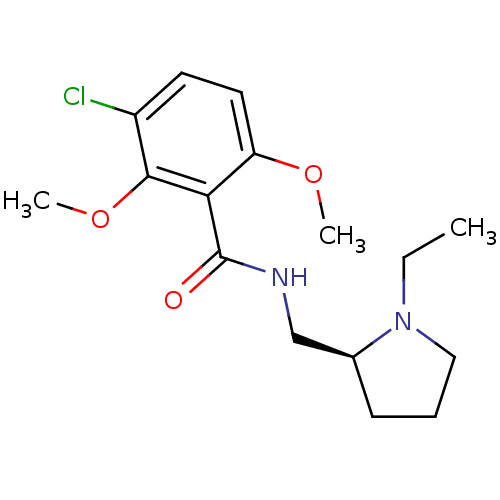
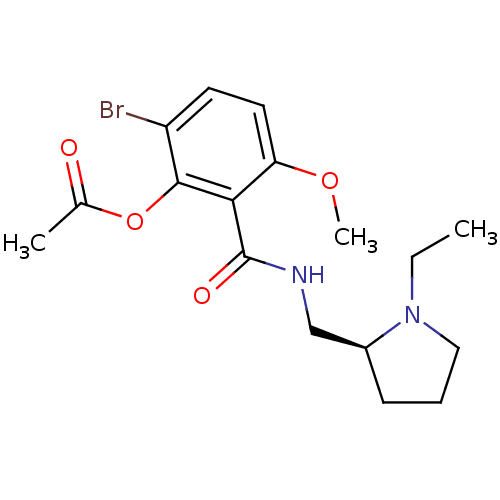
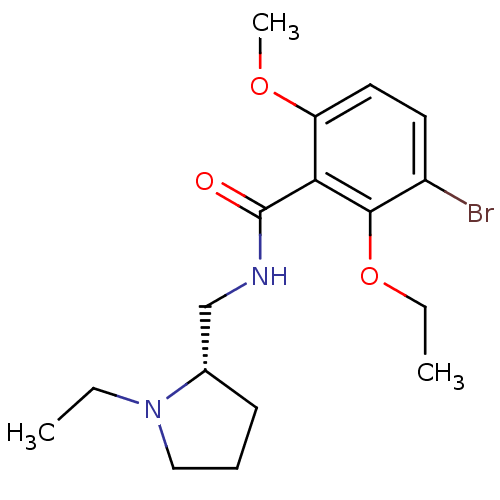
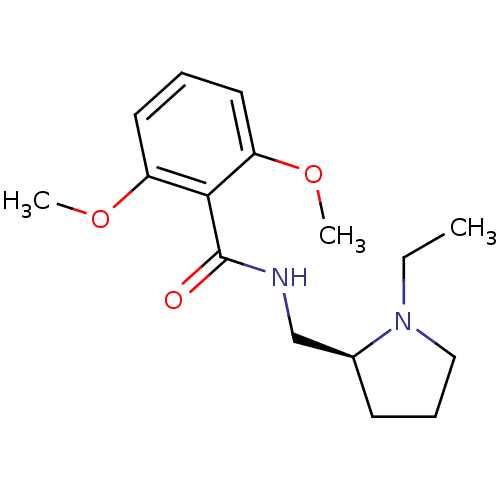
Report error Found 26 Enz. Inhib. hit(s) with all data for entry = 50035583
Affinity DataIC50: 3.40nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 4.70nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 4.80nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 6.30nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 12nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 19nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 21nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 21nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 26nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 39nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 56nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 64nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 123nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 233nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 315nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 362nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 921nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 987nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 1.57E+3nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 2.79E+3nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 3.02E+3nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 3.19E+3nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 3.50E+3nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 1.30E+4nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair
Affinity DataIC50: 1.53E+4nMAssay Description:In vitro binding affinity at Dopamine receptor D2 in rat by displacing [3H]- spiperone from rat striatal membraneMore data for this Ligand-Target Pair