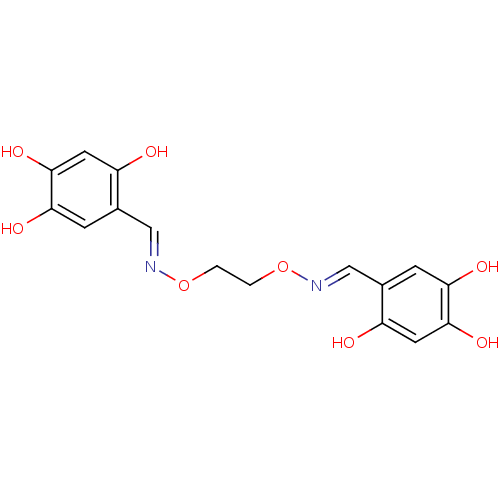
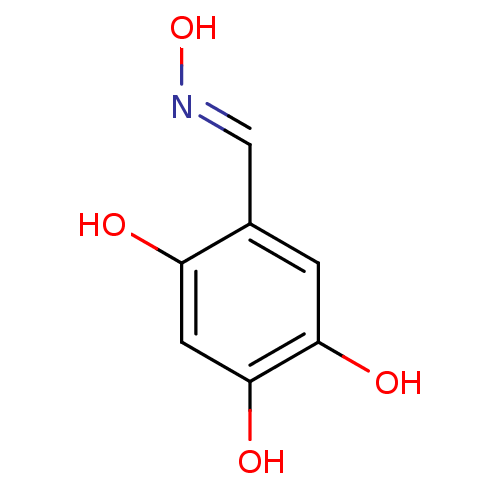
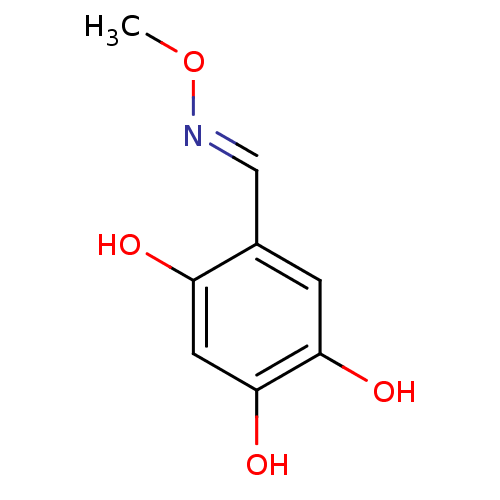
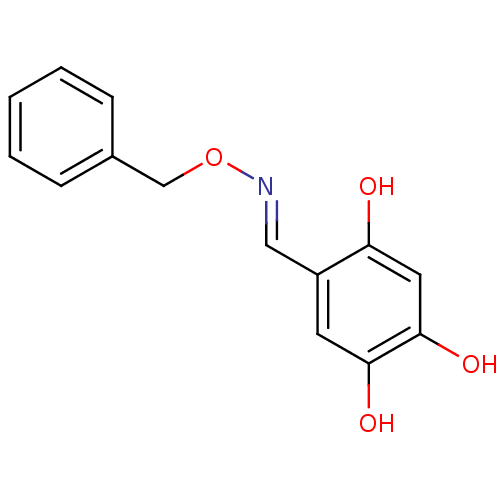
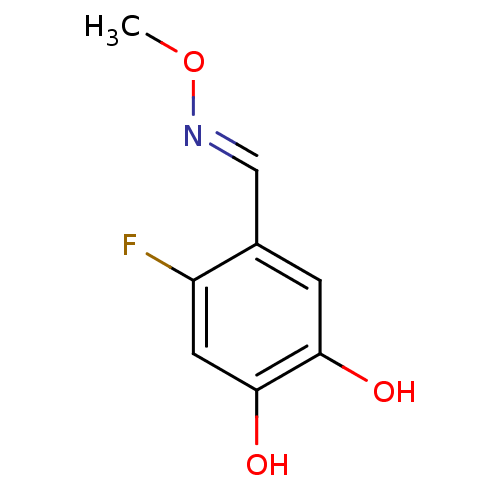
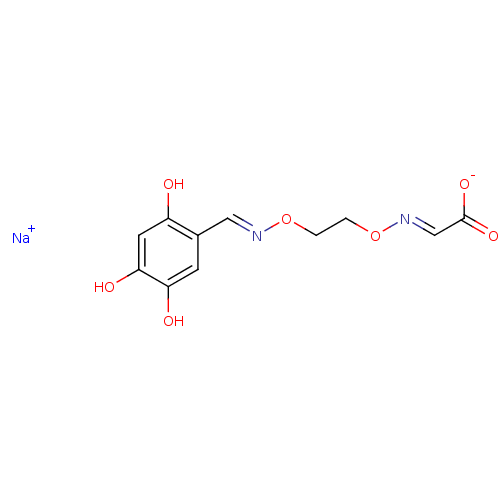
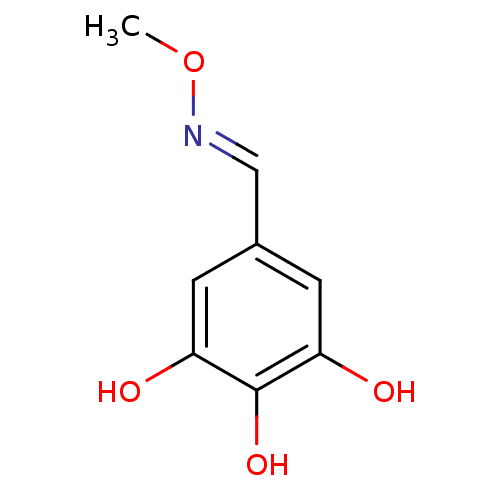
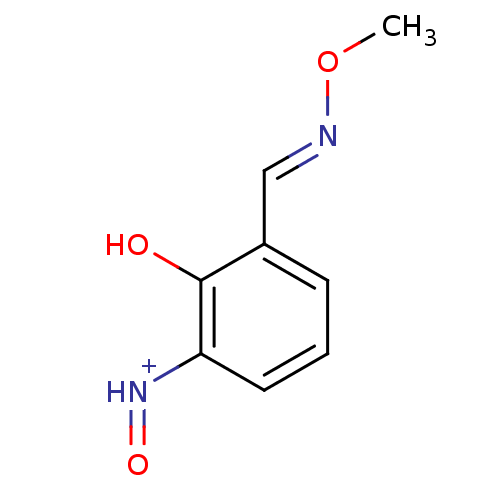
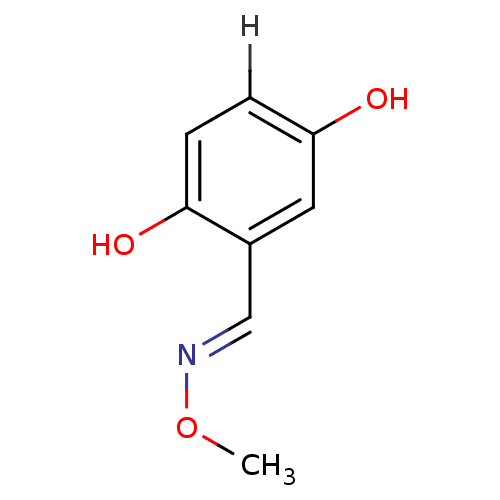
Report error Found 21 Enz. Inhib. hit(s) with all data for entry = 6773
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: 1.00E+3nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: 2.60E+3nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: 3.90E+3nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: 9.10E+3nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: 1.51E+4nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: 1.84E+4nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: 2.94E+4nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: 4.78E+4nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: >7.50E+4nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: >7.50E+4nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: >7.50E+4nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: >1.00E+5nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: >2.00E+5nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: >2.00E+5nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: >2.00E+5nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: >2.00E+5nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: >2.00E+5nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: >2.00E+5nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: >2.00E+5nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: >2.00E+5nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair
Target1-deoxy-D-xylulose 5-phosphate reductoisomerase(Escherichia coli (strain K12))
The Johns Hopkins University School of Medicine
The Johns Hopkins University School of Medicine
Affinity DataKi: >2.00E+5nMpH: 8.0Assay Description:DXP synthase reaction mixtures containing HEPES (100 mM, pH 8.0), MgCl2 (2 mM), NaCl (5 mM), ThDP (1 mM), BSA (1 mg/mL), pyruvate (12.5-250 µM), D-GA...More data for this Ligand-Target Pair