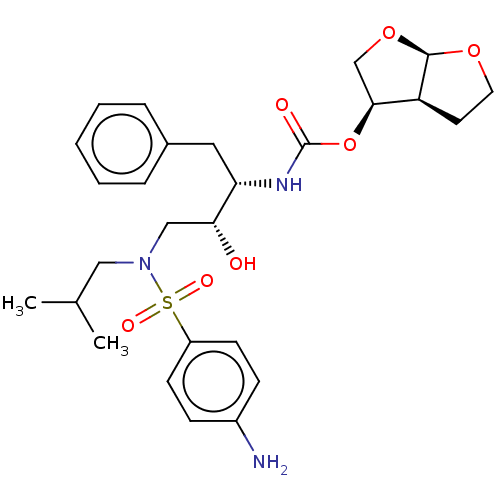
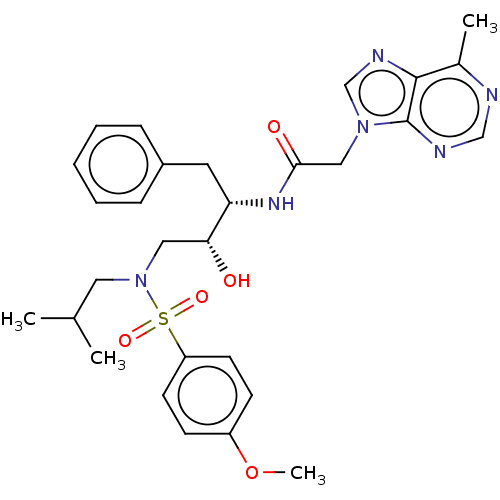
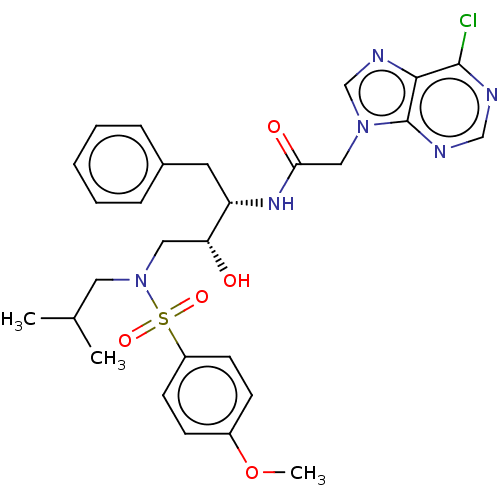
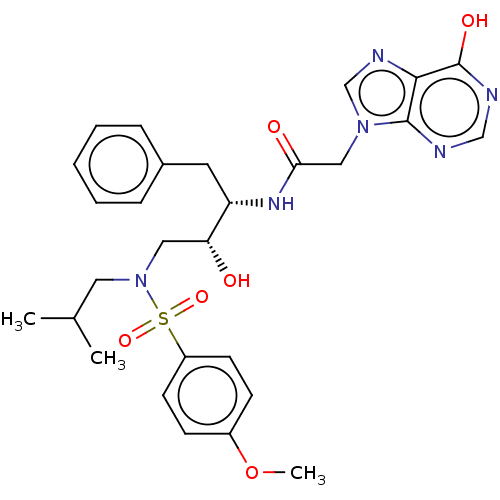
Report error Found 34 Enz. Inhib. hit(s) with all data for entry = 50007553
Affinity DataIC50: 4nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 42nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 43nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 68nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 80nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 150nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 180nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 190nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 240nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 310nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 360nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 460nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 570nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 570nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 570nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 640nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 790nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 960nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.24E+3nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.51E+3nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.73E+3nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.81E+3nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.98E+3nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 2.43E+3nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 2.53E+3nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 2.58E+3nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 2.60E+3nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 3.58E+3nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 3.68E+3nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 3.76E+3nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 4.73E+3nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 6.79E+3nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 7.02E+3nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair
Affinity DataIC50: 1.17E+4nMAssay Description:Inhibition of wild type HIV1 protease expressed in Escherichia coli using Arg-Glu (EDANS)-Ser-Gln-Asn-Tyr-Pro-Ile-Val-Gln-Lys(DABCYL)-Arg as substrat...More data for this Ligand-Target Pair